Volume 17, Number 12—December 2011
Dispatch
Genogroup I and II Picobirnaviruses in Respiratory Tracts of Pigs
Figure
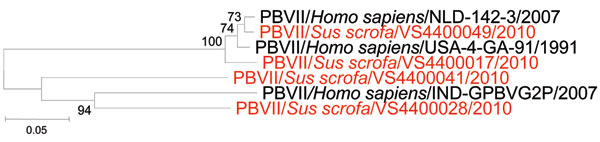
Figure. Phylogenetic analysis of genogroup II picobirnaviruses. Neighbor-joining (Jukes-Cantor model) phylogenetic tree of an ≈339-bp fragment (reference strain 4-GA-91 (9) of the picobirnavirus genogroup II RNA-dependent RNA polymerase gene from known human picobirnaviruses and newly characterized porcine picobirnaviruses in this study (JN176312–315, shown in red). Significant bootstrap values are shown. Nomenclature of depicted viruses is according to recent proposals (7). PBVII/Homo sapiens/USA-4-GA-91/1991 (AF246940) (9); PBVII/Homo sapiens/NLD-VS142–3/2007 (GU968925) (6); and PBVII/Homo sapiens/IND-GPBV6G2P/2007 (AB526257). Scale bar indicates nucleotide substitutions per site.
References
- Field HE, Mackenzie JS, Daszak P. Henipaviruses: emerging paramyxoviruses associated with fruit bats. Curr Top Microbiol Immunol. 2007;315:133–59. DOIPubMedGoogle Scholar
- Garten RJ, Davis CT, Russell CA, Shu B, Lindstrom S, Balish A, Antigenic and genetic characteristics of swine-origin 2009 A(H1N1) influenza viruses circulating in humans. Science. 2009;325:197–201. DOIPubMedGoogle Scholar
- Shinde V, Bridges CB, Uyeki TM, Shu B, Balish A, Xu X, Triple-reassortant swine influenza A (H1) in humans in the United States, 2005–2009. N Engl J Med. 2009;360:2616–25. DOIPubMedGoogle Scholar
- Allander T, Tammi MT, Eriksson M, Bjerkner A, Tiveljung-Lindell A, Andersson B. Cloning of a human parvovirus by molecular screening of respiratory tract samples. Proc Natl Acad Sci U S A. 2005;102:12891–6. DOIPubMedGoogle Scholar
- Smits SL, van Leeuwen M, Kuiken T, Hammer AS, Simon JH, Osterhaus AD. Identification and characterization of deer astroviruses. J Gen Virol. 2010;91:2719–22. DOIPubMedGoogle Scholar
- van Leeuwen M, Williams MM, Koraka P, Simon JH, Smits SL, Osterhaus AD. Human picobirnaviruses identified by molecular screening of diarrhea samples. J Clin Microbiol. 2010;48:1787–94. DOIPubMedGoogle Scholar
- Fregolente MC, Gatti MS. Nomenclature proposal for picobirnavirus. Arch Virol. 2009;154:1953–4. DOIPubMedGoogle Scholar
- Bányai K, Jakab F, Reuter G, Bene J, Uj M, Melegh B, Sequence heterogeneity among human picobirnaviruses detected in a gastroenteritis outbreak. Arch Virol. 2003;148:2281–91. DOIPubMedGoogle Scholar
- Rosen BI, Fang ZY, Glass RI, Monroe SS. Cloning of human picobirnavirus genomic segments and development of an RT-PCR detection assay. Virology. 2000;277:316–29. DOIPubMedGoogle Scholar
- Symonds EM, Griffin DW, Breitbart M. Eukaryotic viruses in wastewater samples from the United States. Appl Environ Microbiol. 2009;75:1402–9. DOIPubMedGoogle Scholar
- Giordano MO, Martinez LC, Rinaldi D, Guinard S, Naretto E, Casero R, Detection of picobirnavirus in HIV-infected patients with diarrhea in Argentina. J Acquir Immune Defic Syndr Hum Retrovirol. 1998;18:380–3. DOIPubMedGoogle Scholar
- Martínez LC, Giordano MO, Isa MB, Alvarado LF, Pavan JV, Rinaldi D, Molecular diversity of partial-length genomic segment 2 of human picobirnavirus. Intervirology. 2003;46:207–13. DOIPubMedGoogle Scholar
- Bányai K, Martella V, Bogdan A, Forgach P, Jakab F, Meleg E, Genogroup I picobirnaviruses in pigs: evidence for genetic diversity and relatedness to human strains. J Gen Virol. 2008;89:534–9. DOIPubMedGoogle Scholar
- Giordano MO, Martinez LC, Masachessi G, Barril PA, Ferreyra LJ, Isa MB, Evidence of closely related picobirnavirus strains circulating in humans and pigs in Argentina. J Infect. 2011;62:45–51. DOIPubMedGoogle Scholar
- Carruyo GM, Mateu G, Martinez LC, Pujol FH, Nates SV, Liprandi F, Molecular characterization of porcine picobirnaviruses and development of a specific reverse transcription–PCR assay. J Clin Microbiol. 2008;46:2402–5. DOIPubMedGoogle Scholar
Page created: December 01, 2011
Page updated: December 01, 2011
Page reviewed: December 01, 2011
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.