Volume 18, Number 10—October 2012
Dispatch
Schmallenberg Virus as Possible Ancestor of Shamonda Virus
Figure
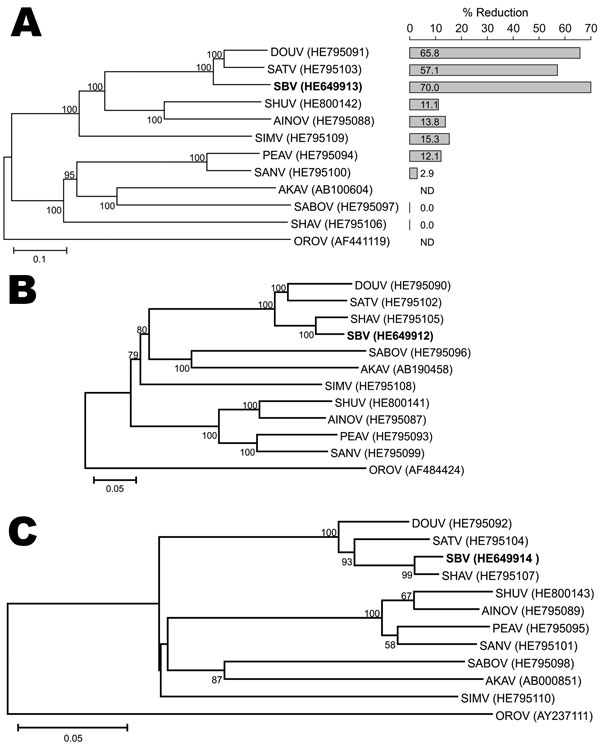
Figure. . . . Maximum-likelihood trees showing phylogenetic relationships of Simbu serogroup viruses for the M (A), L (B), and S (C) coding regions. The bar plot in panel A indicates the percentage of titer reduction of each virus by anti–Schmallenberg virus serum. GenBank accession numbers are shown. Numbers at nodes indicate percentage of 1,000 bootstrap replicates (values <50 are not shown). Scale bars indicate nucleotide substitutions per site. DOUV, Douglas virus; SATV, Sathuperi virus; SBV, Schmallenberg virus; SHUV, Shuni virus; AINOV, Aino virus; SIMV, Simbu virus; PEAV, Peaton virus; SANV, Sango virus; AKAV, Akabane virus; SABOV, Sabo virus; SHAV, Shamonda virus; OROV, Oropouche virus. ND, not determined.
1These authors contributed equally to this article.