Volume 18, Number 12—December 2012
Dispatch
Arctic-like Rabies Virus, Bangladesh
Figure 1
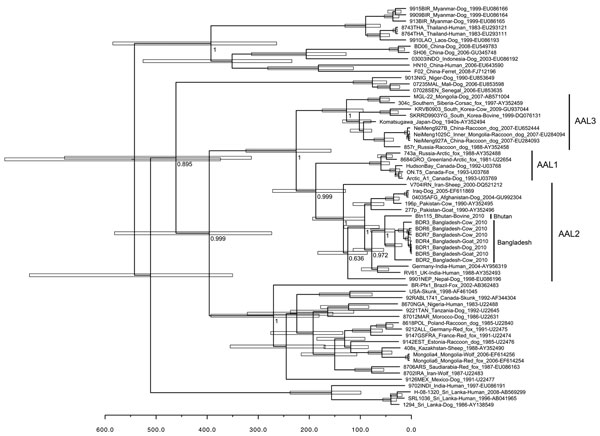
Figure 1. . . Bayesian maximum credibility tree showing genealogy of rabies virus obtained by analyzing nucleotide sequences of full nucleoprotein (N) gene sequences (1,350 nt), Bangladesh. Nodes indicate the mean age at which they are separated from the most recent common ancestor, and white horizontal bars at nodes indicate 95% highest posterior density values of the most recent common ancestor. Numbers at the main nodes indicate posterior values. Scale bar indicates time scale in years starting from 2010. Each strain name is followed by country of origin, host, year of detection, and GenBank accession number. Nucleotide sequence data of the N gene of rabies viruses from Bangladesh appear in the DDBJ/EMBL/GenBank nucleotide sequence databases: accession nos.: AB699214 (rabies virus strain BDR1), AB699215 (strain BDR2), AB699216 (strain BDR3), AB699217 (strain BDR4), AB699218 (strain BDR6), AB699219 (strain BDR7), and AB699220 (whole genome of strain BDR5). AAL, Arctic/Arctic-like.