Volume 18, Number 2—February 2012
Dispatch
Shuni Virus as Cause of Neurologic Disease in Horses
Figure 2
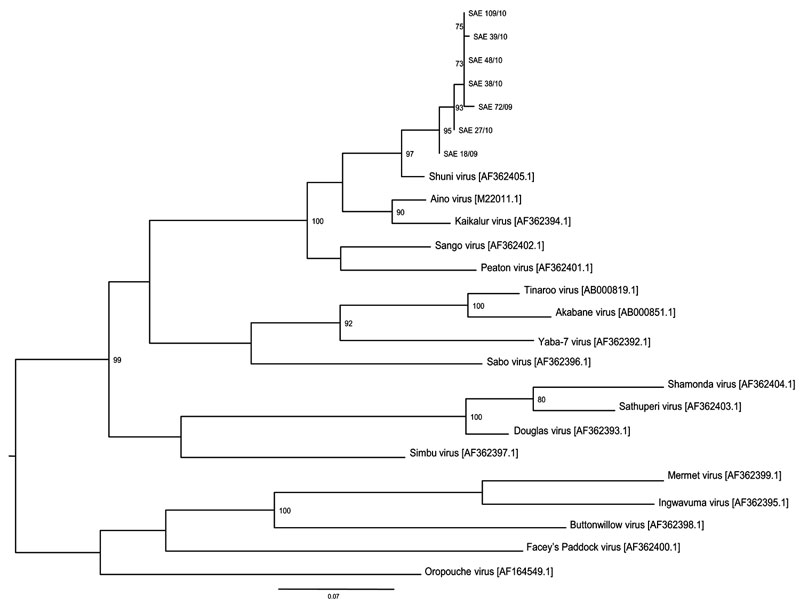
Figure 2. Maximum-likelihood tree constructed under the HKY codon position substitution model using PhyML (http://code.google.com/p/phyml/) of a 330-bp fragment of small segment RNA of Shuni virus (SHUV) identified in horses in South Africa, with representative sequences of selected other orthobunyaviruses. Scale bar = 0.07 nt substitutions. Estimates were based on bootstrap resampling conducted with 100 replicates. Only values >70 are shown. All SHUV amplicons were sequenced, and the data were deposited in GenBank, accession nos. SAE 18/09–HQ610137, SAE 72/09–HQ610138, SAE 27/10–HQ 610139, SAE 38/10–HQ 610140, SAE 39/10–HQ 610141, SAE 48/10–HQ 610142, and SAE 109/10–HQ 610143. Reference strains and GenBank accession numbers are indicated.
Page created: January 19, 2012
Page updated: January 19, 2012
Page reviewed: January 19, 2012
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.