Volume 19, Number 12—December 2013
Dispatch
Novel Reassortant Influenza A(H1N2) Virus Derived from A(H1N1)pdm09 Virus Isolated from Swine, Japan, 2012
Figure
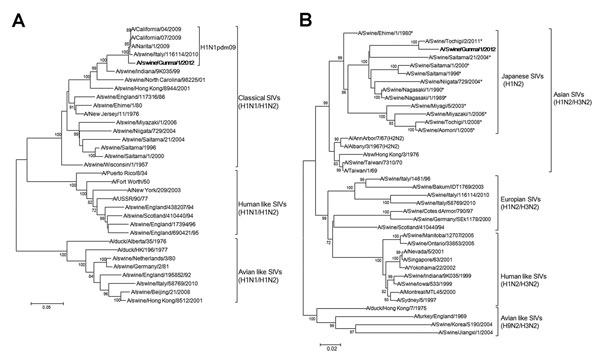
Figure. . Phylogenetic tree based on the nucleotide sequences of hemagglutinin (A) and neuraminidase (B) genes of A/swine/Gunma/1/2012, a novel H1N2 swine influenza virus (SIV) strain. Distance was calculated according to the Kimura 2-parameter method; the trees were constructed by using the neighbor-joining method with labeling of the branches showing at least 70% bootstrap support. Boldface text indicates the novel strain reassorted from strains of the SIV H1N2 subtype. Asterisks indicate reference strains compared with A/Swine/Gunma/1/2012 used to calculate the identity of neuraminidase gene. Scale bars indicate nucleotide substitutions per site.
Page created: November 19, 2013
Page updated: November 19, 2013
Page reviewed: November 19, 2013
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.