Volume 19, Number 2—February 2013
Letter
Toscana Virus Isolated from Sandflies, Tunisia
Figure
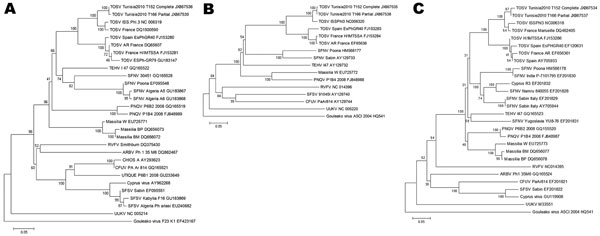
Figure. . Phylogenetic analysis of 3 segments of Toscana virus (TOSV) isolates from pools of sandflies collected in Tunisia and homologous sequences of other selected phleboviruses. A) Large segments; B) medium segments; C) small segments. Sequences are identified by virus name or acronym, strain name, and GenBank accession number. Scale bars indicate nucleotide substitutions per site. TEHV, Tehran virus; SFNV, sandfly fever Naples virus; PNQV, Punique virus; RVFV, Rift Valley fever virus; ARBV, Arbia virus; CHIOS, phlebovirus Chios-A; CFUV, Corfou virus; SFSV, sandfly fever Sicilian virus; UUKV, Uukuniemi virus.
1These authors contributed equally to this manuscript.
2These authors contributed equally to this manuscript.
Page created: January 23, 2013
Page updated: January 23, 2013
Page reviewed: January 23, 2013
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.