Volume 5, Number 1—February 1999
Research
Genetic Diversity and Distribution of Peromyscus-Borne Hantaviruses in North America
Figure 2
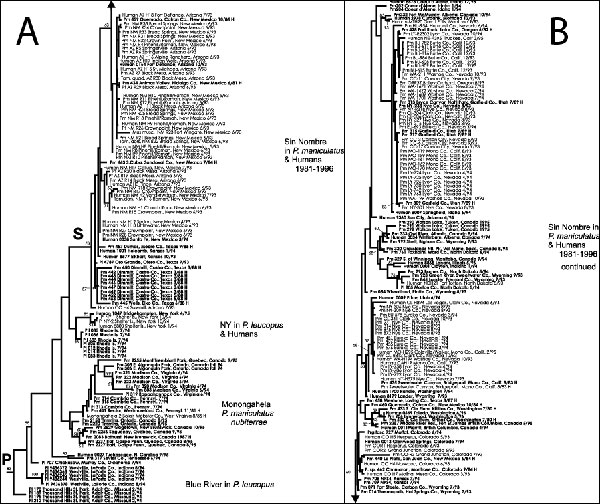
Figure 2. Phylogenetic tree of hantaviruses associated with Peromyscus species rodents. Figure provides a detailed view of clade P in Figure 1. S indicates clade containing classical SN virus samples detected in humans or P. maniculatus. See Figure 1 legend for overall tree description. Additional species source of material abbreviations include: Pm, Peromyscus maniculatus; Pl, Peromyscus leucopus, Prg.fasc, Perognathus fasciatus; Tam.quad, Tamias quadrimaculatus; Pt, Peromyscus truei; Mus musc., Mus musculus, and Tam.dors., Tamias dorsalis. Samples from historic materials are followed by an H.
Page created: December 10, 2010
Page updated: December 10, 2010
Page reviewed: December 10, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.