Volume 11, Number 12—December 2005
Research
Antimicrobial-drug Susceptibility of Human and Animal Salmonella Typhimurium, Minnesota, 1997–2003
Abstract
We compared antimicrobial resistance phenotypes and pulsed-field gel electrophoresis (PFGE) subtypes of 1,028 human and 716 animal Salmonella enterica serotype Typhimurium isolates from Minnesota from 1997 to 2003. Overall, 29% of human isolates were multidrug resistant. Predominant phenotypes included resistance to ampicillin, chloramphenicol or kanamycin, streptomycin, sulfisoxazole, and tetracycline (ACSSuT or AKSSuT). Most human multidrug-resistant isolates belonged to PFGE clonal group A, characterized by ACSSuT resistance (64%), or clonal group B, characterized by AKSSuT resistance (19%). Most animal isolates were from cattle (n = 358) or swine (n = 251). Eighty-one percent were multidrug resistant; of these, 54% were at least resistance phenotype ACSSuT, and 43% were at least AKSSuT. More than 80% of multidrug-resistant isolates had a clonal group A or B subtype. Resistance to ceftriaxone and nalidixic acid increased, primarily among clonal group A/ACSSuT isolates. Clonal group B/AKSSuT isolates decreased over time. These data support the hypothesis that food animals are the primary reservoir of multidrug-resistant S. Typhimurium.
Nontyphoidal salmonellae are a leading cause of acute gastroenteritis in the United States (1). Salmonella enterica serotype Typhimurium is the most common serotype isolated from humans (2). In the 1990s, multidrug-resistant (MDR) S. Typhimurium definitive phage type 104 (DT104) emerged in the United States; most isolates were resistant to ampicillin, chloramphenicol, streptomycin, sulfisoxazole, and tetracycline (resistance phenotype [R-type] ACSSuT) (3). S. Typhimurium R-type AKSSuT (with resistance to kanamycin) has also recently emerged in the United States (4). Several studies have documented adverse health effects due to the increasing resistance observed in S. Typhimurium (5–9). These effects include an increased risk for infection with S. Typhimurium (5), increased risk for bloodstream infection (6), increased risk for hospitalization (6,7), treatment failures (8), and increased risk for death (9).
MDR S. Typhimurium strains have been well documented in food animals, as have MDR S. Typhimurium outbreaks in humans from animal contact or foods of animal origin (8,10–17). However, contemporaneous parallel data on resistance in human and animal S. Typhimurium isolates in the United States are limited (18), and an advisory panel has called for linking surveillance for bacterial resistance in animals and humans to further evaluate the human health effects of antimicrobial drug use in agriculture (19). The objectives of our study were to evaluate antimicrobial resistance and molecular subtyping data from all human clinical S. Typhimurium isolates received through statewide, population-based, active laboratory surveillance in Minnesota and to compare the human isolates to isolates from clinically ill animals in Minnesota identified by the Minnesota Veterinary Diagnostic Laboratory (MVDL).
Human and Animal Isolates
The Minnesota Department of Health (MDH) requires clinical laboratories to submit all Salmonella isolates to its public health laboratory as part of active, laboratory-based surveillance. MDH audits clinical laboratories to ensure complete reporting. Human S. Typhimurium isolates submitted to MDH from 1997 to 2003 were eligible for this study. Isolates that were part of an identified outbreak were excluded, except for the index case-isolate. Isolates from secondary cases in household clusters and duplicate submissions from the same case also were excluded.
MVDL is a regional laboratory for veterinarians; pertinent diagnostic samples are cultured for Salmonella spp. Isolates are sent to the National Veterinary Services Laboratories (Ames, Iowa) for serotyping. Confirmed S. Typhimurium isolates are forwarded to MDH. S. Typhimurium isolates obtained from diagnostic specimens from sick animals cultured at MVDL from 1997 to 2003 were eligible for this study. Isolates from the same farm with the same pulsed-field gel electrophoresis (PFGE) subtype discovered within 1 year of the initial isolate collection date were excluded. Research animal submission, environmental sample, and non-Minnesota animal isolates were excluded.
Study Populations
From 1997 to 2003, a total of 4,333 culture-confirmed cases of human salmonellosis were reported in Minnesota. S. Typhimurium was the most common serotype; it accounted for 1,193 (28%) cases overall (median 172 cases/year, range 124–201). Of the 1,193 human S. Typhimurium case-isolates, 1,028 (86%) were included in this study (Table 1).
A total of 716 animal isolates were included in this study (median 91/year, range, 67–150) (Table 1). Isolates represented 644 farms and animal owners and 72 of 87 Minnesota counties. Most isolates were of bovine (n = 358, 50%) or porcine (n = 251, 35%) origin. Cattle isolates decreased markedly over time: 106 isolates in 1997, 100 isolates in 1998, 49 isolates in 1999, 31 isolates in 2000, 29 isolates in 2001, 18 isolates in 2002, and 25 isolates in 2003. Conversely, swine isolates increased over time: 32 isolates in 1997, 27 isolates in 1998, 33 isolates in 1999, 22 isolates in 2000, 44 isolates in 2001, 39 isolates in 2002, and 54 isolates in 2003. The remaining isolates included 38 (5%) avian (5 turkey, 1 chicken, 7 unknown, and 25 miscellaneous species), 29 (4%) equine, 21 (3%) feline, 7 (1%) canine, and 12 (2%) other species.
Isolate Testing
All S. Typhimurium isolates (including variant Copenhagen) submitted to MDH were confirmed as S. Typhimurium and subtyped by PFGE. PFGE patterns were compared by using BioNumerics software (Applied Maths, Sint-Martens-Latem, Belgium) with the Dice coefficient and a 1% band matching criterion (20). Patterns with no visible differences were considered indistinguishable. Subtypes for S. Typhimurium at MDH are designated with the prefix "TM" followed by a number (e.g., TM123). PFGE patterns are also submitted to the PulseNet national database. Antimicrobial susceptibility testing was performed with the disc diffusion method and interpretive standards of the National Committee for Clinical and Laboratory Standards (NCCLS) (21). Antimicrobial susceptibility was determined for ampicillin (A), chloramphenicol (C), kanamycin (K), streptomycin (S), sulfisoxazole (Su), tetracycline (T), cephalothin (Ct), ceftriaxone (Cr), ciprofloxacin (Cp), gentamicin (G), nalidixic acid (Na), and trimethoprim/sulfamethoxazole (Sxt). The Etest for MIC was performed on isolates with intermediate susceptibility to ceftriaxone by disc diffusion; MICs were interpreted according to NCCLS criteria (21). An MIC of 48 μg/mL was considered resistant. Multidrug resistance was defined as resistance to >5 antimicrobial drugs.
PFGE data were analyzed by the first 3 tiers of criteria described by Tenover et al. (0, 1- to 3-, and 4- to 6-band differences) (22). Two primary PFGE subtype clusters that accounted for a large proportion of MDR isolates were identified on the basis of a <3-band difference: 1) clonal group A (CGA), composed of subtypes <3 bands different from PFGE subtype TM5b, and 2) clonal group B (CGB), composed of subtypes <3 bands different from PFGE subtype TM54.
Statistical Analysis
Resistance was analyzed in terms of R-types ACSSuT, AKSSuT, and ACKSSuT. R-type ACKSSuT isolates were included in analyses of "at least R-type ACSSuT" isolates, but not "at least R-type AKSSuT" isolates. Where indicated, ACKSSuT isolates were evaluated independently of ACSSuT. R-types were analyzed in terms of clonal group. The χ2 test for trend was used to evaluate resistance trends (EpiInfo 6.04d, Centers for Disease Control and Prevention, Atlanta, GA, USA). Proportions were compared by using the χ2 test. Uncorrected p value and exact 95% mid-p limits for the maximum likelihood estimate of the odds ratio (OR) were used. A p value <0.05 was considered significant.
Human Isolates
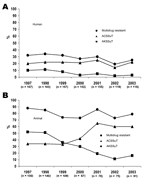
Of the 1,028 S. Typhimurium isolates, 455 (44%) were resistant to >1 antimicrobial drug, and 296 (29%) were MDR (Table 1). Among MDR isolates, 217 (73%) were at least R-type ACSSuT, and 64 (22%) were at least AKSSuT (Table 2). The proportion of MDR isolates decreased from 32% in 1997 to 25% in 2003 (χ2 for linear trend 6.3, p = 0.01) (Figure 1). The proportion that were at least AKSSuT also decreased, from 10% in 1997 to 3% in 2003 (χ2 for linear trend 17.7, p<0.001).
Eighteen (1.8%) isolates were resistant to ceftriaxone; all were MDR (Table 1). Ceftriaxone resistance was more prevalent from 2000 to 2003 (2.8%) than from 1997 to 1999 (0.6%) (OR 4.6, 95% confidence interval [CI] 1.4–20.0, p = 0.008). Eleven (1.2%) isolates were resistant to nalidixic acid; all were MDR. Nalidixic acid resistance was more prevalent from 2000 to 2003 (1.8%) than from 1997 to 1999 (0.2%) (OR 9.2, 95% CI 1.5–200.8, p = 0.011). Fifty-one (5%) isolates were resistant to trimethoprim-sulfamethoxazole. Of these, 34 (67%) were MDR, including 20 (39%) that were at least R-type ACSSuT and 6 (12%) that were at least AKSSuT. Forty-three (4%) isolates were resistant to gentamicin; of these, 23 (53%) were MDR.
We identified 271 unique PFGE subtypes among the 1,028 human S. Typhimurium isolates (median 63 subtypes/year, range 52–72). The 10 most common subtypes accounted for 509 (50%) isolates. CGA was composed of 31 PFGE subtypes. These subtypes accounted for 217 (21%) of all 1,028 human isolates, 188 (64%) of 296 MDR isolates, and 181 (83%) of 217 isolates that were at least R-type ACSSuT, including 12 isolates that were at least R-type ACKSSuT (Table 2, Figures 2 and 3).
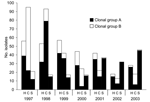
CGB was composed of 20 subtypes and accounted for 81 (8%) of all 1,028 human isolates, 55 (19%) of 296 MDR isolates, and 51 (80%) of 64 isolates that were at least R-type AKSSuT (Table 2, Figures 2 and 3). The number of isolates with CGB subtypes decreased substantially from 2001 to 2003 (Figure 2).
Animal Isolates
Overall, 640 (89%) of the 716 animal S. Typhimurium isolates were resistant to >1 antimicrobial drug, and 580 (81%) were MDR (Table 1). Of the 580 MDR isolates, 315 (54%) were at least ACSSuT, and 250 (43%) were at least AKSSuT (Table 2). The proportion of isolates that were at least ACSSuT increased over time (χ2 for linear trend 39.5, p<0.001). Conversely, the proportion that were at least AKSSuT decreased (χ2 for linear trend 71.7, p<0.001) (Figure 1).
Of the 358 cattle isolates, 205 (57%) were at least R-type AKSSuT, and 101 (28%) were at least ACSSuT. The decrease in cattle isolates over time reflected a decrease in the number that were at least AKSSuT (Figure 2). In addition, the proportion of cattle isolates that were at least AKSSuT decreased significantly over time (χ2 for linear trend 8.9, p = 0.003).
Of the 251 swine isolates, 180 (72%) were at least R-type ACSSuT, and 30 (12%) were at least AKSSuT. The increase in swine isolates over time reflected an increase in the number that were at least ACSSuT (Figure 2). In addition, the proportion of swine isolates that were at least ACSSuT increased significantly over time (χ2 for linear trend 25.4, p<0.001). Nine (24%) of 38 avian isolates, 19 (66%) of 29 equine isolates, and 15 (71%) of 21 feline isolates were MDR.
Twenty-five (3.5%) animal isolates were resistant to ceftriaxone. Ceftriaxone resistance was more prevalent from 2000 to 2003 (5.1%) than from 1997 to 1999 (2.2%) (OR 2.4, 95% CI 1.0–5.7, p = 0.035). Twelve ceftriaxone-resistant isolates were from cattle, and 10 were from swine. Four (0.6%) animal isolates were resistant to nalidixic acid, including 1 bovine isolate in 1997 and 3 turkey isolates in 2003. Eighty-one (11%) animal isolates were resistant to trimethoprim-sulfamethoxazole. Of these, 79 (98%) were MDR, and 62 (77%) were at least ACSSuT. Seventy-one (10%) animal isolates were resistant to gentamicin. Of these, 69 (97%) were MDR, and 44 (62%) were at least ACSSuT.
A total of 190 unique PFGE subtypes were identified among the 716 animal isolates (median 36 subtypes/year, range 31–47). Among animal isolates, CGA was composed of 48 PFGE subtypes. CGA accounted for 264 (37%) of all 716 animal isolates, 256 (44%) of 580 MDR isolates, and 249 (79%) of 315 isolates that were at least R-type ACSSuT, including 67 at least ACKSSuT isolates (Table 2, Figures 2 and 3). CGB was composed of 35 subtypes. CGB accounted for 278 (39%) of all 716 animal isolates, 250 (43%) of 580 MDR isolates, and 227 (91%) of 250 isolates that were at least R-type AKSSuT.
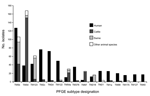
Distribution of PFGE subtypes differed by species and year (Figures 2 and 4). CGB subtypes occurred predominantly in cattle and accounted for 67% of cattle isolates. As with AKSSuT isolates, CGB subtype isolates were numerous in cattle from 1997 to 1998, but the number dropped markedly in 2002 and 2003 (Figure 2). CGA subtype isolates increased in swine from 2000 to 2003 and substantially outnumbered CGA cattle isolates during those years. CGA isolates in cattle were most common from 1997 to 1998 and then declined to a relatively stable, low level (Figure 2).
Of 9 MDR avian isolates, 5 were in CGA and 1 was in CGB. Of 19 MDR equine isolates, 4 were in CGA and 5 were in CGB. Of 15 MDR feline isolates, 8 were in CGA and 6 were in CGB.
Animal-Human Isolate Comparison
Combining the 1,028 human and 716 animal S. Typhimurium study isolates, 395 PFGE subtypes were identified. Sixty-six subtypes occurred both in animals and humans. These 66 subtypes represented 673 (65%) of human and 537 (75%) of animal isolates. Eighteen (27%) of shared subtypes were in CGA, and 12 (18%) were in CGB.
Combining the 296 MDR human isolates and the 580 MDR animal isolates, 183 PFGE subtypes were identified. Of these subtypes, 31 occurred both among human and animal MDR isolates. These 31 subtypes represented 237 (80%) human MDR isolates and 442 (76%) animal MDR isolates. Eighteen of the 31 shared MDR subtypes were in CGA, and 7 were in CGB. Of the 296 MDR human isolates, 177 (60%) had a CGA subtype that also occurred among MDR animal isolates, and 51 (17%) had a CGB subtype that also occurred among MDR animal isolates. Of the 296 MDR human isolates, 243 (82%) belonged to CGA (64%) or CGB (19%). Of the 580 MDR animal isolates, 506 (87%) belonged to CGA (44%) or CGB (43%).
The 6 most common individual subtypes in animals, all of which were in CGA or CGB (Figure 3), were represented among human isolates (Figure 4). TM5b, the second most common animal subtype, was the most common human subtype. TM54, the most common animal subtype, was sixth in humans. TM123 was the third most common animal subtype and fifth in humans (Figure 4).
This study provides a comprehensive comparison of clinical human and animal S. Typhimurium isolates from the same area. Overall, 29% of human S. Typhimurium isolates in Minnesota were MDR. Isolates with at least R-types ACSSuT or AKSSuT made up almost all (95%) of MDR S. Typhimurium in humans. Resistance phenotypes that were at least ACSSuT predominated. The level of multidrug resistance in human isolates decreased from 1997 to 2003, corresponding to a decrease in R-type AKSSuT isolates. Resistance to at least ACSSuT was stable over time. The level of multidrug resistance observed in human isolates in Minnesota was slightly lower than that observed through the National Antimicrobial Resistance Monitoring System (NARMS) through 2002; however, multidrug resistance trends for S. Typhimurium generally paralleled NARMS findings (4,23).
Increasing resistance to ceftriaxone documented in human isolates in Minnesota indicated that ceftriaxone resistance continues to emerge in S. Typhimurium in the United States (13,24). The 1.8% resistance to nalidixic acid observed in human isolates from 2000 to 2003 was not substantially higher than the 1% resistance among NARMS isolates from 2000 to 2002 (23) but was significantly higher than that seen in our isolates from 1997 to 1999. Most of the isolates that were resistant to both ceftriaxone and nalidixic acid were from 2000 or later. Resistance to these antimicrobial agents, as well as gentamicin and trimethoprim-sulfamethoxazole, frequently occurred in isolates that were also resistant to >5 other antimicrobial drugs; this finding was true for all isolates that were resistant to ceftriaxone or nalidixic acid. Resistance to these clinically important antimicrobial drugs was associated most frequently with ACSSuT resistance rather than AKSSuT resistance.
The increasing resistance to ceftriaxone and nalidixic acid (an elementary quinolone) is of concern because extended-spectrum cephalosporins and fluoroquinolones are needed to treat serious Salmonella infections. Recent experiences in Denmark have shown treatment failures and excess deaths associated with quinolone-resistant S. Typhimurium (8,9). The addition of resistance to clinically useful antimicrobial drugs to already-pentaresistant R-types is added cause for concern because pentaresistant S. Typhimurium strains are more likely to cause infection (5) and adverse health outcomes (6,7) than drug-susceptible strains.
Despite the overall diversity observed among S. Typhimurium isolates by PFGE, human MDR isolates were highly clonal. Even when a relatively stringent definition of a clonal group (<3-band difference) was used, >80% of human MDR isolates composed 2 clonal groups. CGA isolates were characterized by ACSSuT resistance and represented most human MDR isolates. Of isolates from this study that were previously phage typed, those in CGA have all been in the DT104 complex (12,25,26). The clonal nature of ACSSuT/DT104 S. Typhimurium in the United States has been well documented (20,27).
CGB isolates were characterized by AKSSuT resistance. This group accounted for 19% of human MDR isolates overall but was more prevalent early in the study, after which a marked decline occurred. As with the ACSSuT/DT104 complex, AKSSuT isolates appear to be largely clonal in nature.
Most S. Typhimurium isolates from clinically ill animals in Minnesota were MDR, which emphasizes that MDR strains are prevalent animal pathogens (10). High resistance levels occurred in all species, throughout the state, and during the entire study period. As with humans, most MDR animal isolates were in either the CGA/ACSSuT (DT104) or CGB/AKSSuT clonal groups. PFGE subtypes found among human and animal MDR isolates were remarkably similar. This similarity is striking considering that Minnesota residents may be exposed to S. Typhimurium during travel or from food produced outside Minnesota.
Among animals, the CGB/AKSSuT clonal group was most common in cattle. The sharp decrease in CGB isolates in cattle was mirrored by a similar decrease in humans. The cause of this decrease in cattle is not known. The CGA/ACSSuT clonal group was distributed more evenly among all animal species but became more common in swine over time. The cause for the increase in swine CGA/ACSSuT isolates is not known.
MDR S. Typhimurium strains similar to those from our study have been recovered from food animals and retail meat products by other investigators, and multiple MDR S. Typhimurium outbreaks caused by foods of animal origin or animal contact have been documented (8,10,11,13–16,28,29). Our data provide additional evidence that food animals are the primary reservoir of MDR S. Typhimurium for humans; MDR S. Typhimurium that belong to CGA or CGB were documented in cattle or swine herds on hundreds of farms throughout Minnesota. Testing isolates with additional genetic subtyping methods and identifying resistance determinants would help further characterize the relationship between animal and human isolates (22,30). In addition, data on use of antimicrobial drugs in animal production (which are currently unavailable in the United States because requirements are lacking) would be helpful in assessing this issue.
Although the number of isolates was relatively small, the level of multidrug resistance was high in both cat and horse isolates. CGA/ACSSuT and CGB/AKSSuT isolates were observed in both species. The importance of these infections in companion animals has been demonstrated by recent MDR S. Typhimurium outbreaks in humans associated with small animal veterinary facilities, including a Minnesota outbreak of CGA/ACSSuT DT104 infections in persons who adopted infected kittens from a humane society (12).
The source of animal isolates for our study is a limitation in that Salmonella isolates from clinically ill animals overstate the level of antimicrobial resistance observed in isolates from healthy animals; therefore, strains from ill animals are not representative of strains carried by animals at slaughter (31,32). However, when we have evaluated S. Typhimurium isolates from other studies, the most prominent CGA and CGB subtypes from our study also have been found in healthy food animals or their environments. For example, TM5b and TM123 isolates were recovered from healthy, market-ready pigs at slaughter (J.B. Bender, unpub. data). Subtypes TM5b, TM123, and TM54 were represented among poultry isolates evaluated by Rajashekara et al. (28). In a study of Salmonella isolates on dairy farms in 4 states, including Minnesota, subtypes TM5b and TM54 were recovered from healthy dairy cows or environmental samples (33). Finally, MDR S. Typhimurium is present in the retail meat supply; in a recent study, almost all strains of S. Typhimurium recovered from ground meat (pork and chicken) were MDR phage types DT104 or DT208 (29).
Another limitation of our study was the underrepresentation of poultry isolates. Minnesota is a leading poultry producer; however, most poultry diagnostics are conducted by the Minnesota Poultry Testing Laboratory. This laboratory has documented DT104 in Minnesota poultry (28). In our study, 3 of 4 nalidixic acid–resistant animal isolates were from turkeys, even though very few turkey isolates were tested. The role of poultry as a potential reservoir for MDR S. Typhimurium, including nalidixic acid–resistant strains, should be more thoroughly evaluated.
We agree with other investigators that the emergence of multidrug resistance in S. Typhimurium is associated with the widespread dissemination of clonal groups (27,34). The changing trends of MDR S. Typhimurium in cattle versus swine observed in our study and the presence of MDR strains in poultry indicate that more study of individual subtypes and resistance determinants (including specific mobile genetic elements) is required to understand the movement of these strains within and between animal species. Improved biosecurity practices to interrupt dissemination are undoubtedly the key in controlling these strains (27).
The potential role of the selection pressure of antimicrobial drugs used in animal agriculture in the dissemination of MDR S. Typhimurium clonal groups must be considered. The ability of MDR S. Typhimurium strains to accumulate additional resistances allows them to survive under a wide range of conditions when antimicrobial agents are used. Use of antimicrobial drugs to which MDR S. Typhimurium strains are already resistant may increase the number of animals infected with these strains and the number of animals that manifest clinical illness. This use is inherently likely to contribute to increased dissemination, both within and between farms. Thus, we encourage the judicious use of all antimicrobial drugs in animals as well as in humans. In particular, the recommendation (19) that nonessential uses of specific antimicrobial drugs in food animals should be eliminated (e.g., the use of tetracyclines and penicillins for growth promotion and feed efficiency) has merit. MDR S. Typhimurium strains are serious pathogens in food animals and humans. Restricting conditions that favor their dissemination should return the benefits of reduced incidence and severity of S. Typhimurium infections in both animals and humans.
Ms Wedel is an epidemiologist in the Minnesota Department of Health, Foodborne, Vectorborne, and Zoonotic Diseases Unit. Her professional interests include foodborne diseases, zoonotic diseases, molecular epidemiology, and antimicrobial resistance of foodborne bacterial pathogens.
Acknowledgments
We thank laboratory staff at the University of Minnesota Veterinary Diagnostic Laboratory, John Besser and staff at the Minnesota Department of Health Public Health Laboratory for their work with Salmonella isolates for this project, and staff from the Minnesota Department of Health Acute Disease Investigation and Control Section, who participated in data collection for this project or reviewed this manuscript.
This work was supported in part through cooperative agreements with the Centers for Disease Control and Prevention (CDC) Emerging Infections Program, Foodborne Diseases Active Surveillance Network (FoodNet) (U50/CCU511190-10) and the CDC Epidemiology and Laboratory Capacity for Infectious Diseases Program (U50/CCU519683-04-4).
References
- Mead PS, Slutsker L, Dietz V, McCaig LF, Bresee JS, Shapiro C, Food-related illness and death in the United States. Emerg Infect Dis. 1999;5:607–25. DOIPubMedGoogle Scholar
- Salmonellosis—technical information [monograph on the Internet]. 2003 Dec [cited 2005 Jan]. Available from http://www.cdc.gov/ncidod/dbmd/diseaseinfo/salmonellosis_t.htm
- Glynn MK, Bopp C, Dewitt W, Dabney P, Mokhtar M, Angulo FJ. Emergence of multidrug-resistant Salmonella enterica serotype Typhimurium DT104 infections in the United States. N Engl J Med. 1998;338:1333–8. DOIPubMedGoogle Scholar
- Rabatsky-Ehr T, Wichard J, Rossiter S, Holland B, Stamey K, Headrick ML, Multidrug-resistant strains of Salmonella enterica Typhimurium, United States, 1997–1998. Emerg Infect Dis. 2004;10:795–801.PubMedGoogle Scholar
- Glynn MK, Reddy V, Hutwagner L, Rabatsky-Ehr T, Shiferaw B, Vugia DJ, Prior antimicrobial use increases the risk of sporadic infections with multidrug-resistant Salmonella enterica serotype Typhimurium: a FoodNet case control study, 1996–1997. Clin Infect Dis. 2004;38(Suppl 3):S227–36. DOIPubMedGoogle Scholar
- Varma JK, Mølbak K, Barrett TJ, Beebe JL, Jones TF, Rabatsky-Ehr T, Antimicrobial-resistant nontyphoidal Salmonella is associated with excess bloodstream infections and hospitalizations. J Infect Dis. 2005;191:554–61. DOIPubMedGoogle Scholar
- Martin LJ, Fyfe M, Doré K, Buxton JA, Pollari F, Henry B, Increased burden of illness associated with antimicrobial-resistant Salmonella enterica serotype Typhimurium infections. J Infect Dis. 2004;189:377–84. DOIPubMedGoogle Scholar
- Mølbak K, Baggesen DL, Aarestrup FM, Ebbesen JM, Engberg J, Frydendahl K, An outbreak of multidrug-resistant, quinolone-resistant Salmonella enterica serotype Typhimurium DT104. N Engl J Med. 1999;341:1420–5. DOIPubMedGoogle Scholar
- Helms M, Vastrup P, Gerner-Smidt P, Mølbak K. Excess mortality associated with antimicrobial drug-resistant Salmonella Typhimurium. Emerg Infect Dis. 2002;8:490–5.PubMedGoogle Scholar
- Akkina JE, Hogue AT, Angulo FJ, Johnson R, Petersen KE, Saini PK, Epidemiologic aspects, control, and importance of multiple-drug resistant Salmonella Typhimurium DT104 in the United States. J Am Vet Med Assoc. 1999;214:790–8.PubMedGoogle Scholar
- Cody SH, Abbott SL, Marfin AA, Schulz B, Wagner P, Robbins K, Two outbreaks of multidrug-resistant Salmonella serotype Typhimurium DT104 infections linked to raw-milk cheese in northern California. JAMA. 1999;281:1805–10. DOIPubMedGoogle Scholar
- Wright JG, Tengelsen LA, Smith KE, Bender JB, Frank RK, Grendon JH, Multidrug-resistant Salmonella Typhimurium in four animal facilities. Emerg Infect Dis. 2005;11:1235–41.PubMedGoogle Scholar
- Fey PD, Safranek TJ, Rupp ME, Dunne EF, Ribot E, Iwen PC, Ceftriaxone-resistant salmonella infection acquired by a child from cattle. N Engl J Med. 2000;342:1242–9. DOIPubMedGoogle Scholar
- Wall PG, Morgan D, Lamden K, Griffin M, Threlfall EJ, Ward LR, Transmission of multi-resistant strains of Salmonella Typhimurium from cattle to man. Vet Rec. 1995;136:591–2. DOIPubMedGoogle Scholar
- Olsen SJ, Ying M, Davis MF, Deasy M, Holland B, Iampietro L, Multidrug-resistant Salmonella Typhimurium infection from milk contaminated after pasteurization. Emerg Infect Dis. 2004;10:932–5.PubMedGoogle Scholar
- Centers for Disease Control and Prevention. Multidrug-resistant Salmonella serotype Typhimurium—United States, 1996. MMWR Morb Mortal Wkly Rep. 1997;46:308–10.PubMedGoogle Scholar
- Davis MA, Hancock DD, Besser TE, Rice DH, Gay JM, Gay C, Changes in antimicrobial resistance among Salmonella enterica serovar Typhimurium isolates from humans and cattle in the Northwestern United States, 1982–1997. Emerg Infect Dis. 1999;5:802–6. DOIPubMedGoogle Scholar
- Swartz MN. Human diseases caused by foodborne pathogens of animal origin. Clin Infect Dis. 2002;34(Suppl 3):S111–22. DOIPubMedGoogle Scholar
- Facts about Antibiotics in Animals and the Impact on Resistance (FAAIR) Scientific Advisory Panel. Select findings and conclusions. Clin Infect Dis. 2004;34(Suppl 3):S73–7.
- Ribot EM, Wierzba RK, Angulo FJ, Barrett TJ. Salmonella enterica serotype Typhimurium DT104 isolated from humans, United States, 1985, 1990, and 1995. Emerg Infect Dis. 2002;8:387–91. DOIPubMedGoogle Scholar
- National Committee for Clinical Laboratory Standards. Zone diameter interpretive standards for Enterobacteriaceae. Wayne (PA): The Committee; 2003. p. 20–4.
- Tenover FC, Arbeit RD, Goering RV, Mickelsen PA, Murray BE, Persing DH, Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995;33:2233–9.PubMedGoogle Scholar
- Centers for Disease Control and Prevention. National Antimicrobial Resistance Monitoring System (NARMS): enteric bacteria. 2002 annual report. 2004 [cited 2005 Sep 26]. Available from http://www.cdc.gov/narms/annual/2002/2002ANNUALREPORTFINAL.pdf
- Dunne EF, Fey PD, Kludt P, Reporter R, Mostashari F, Shillam P, Emergence of domestically acquired ceftriaxone-resistant Salmonella infections associated with AmpC beta-lactamase. JAMA. 2000;284:3151–6. DOIPubMedGoogle Scholar
- Bender JB, Hedberg CW, Boxrud DJ, Besser JM, Wicklund JH, Smith KE, Use of molecular subtyping in surveillance for Salmonella enterica serotype Typhimurium. N Engl J Med. 2001;344:189–95. DOIPubMedGoogle Scholar
- Bender JB, Smith KE, Forfang J, Schaeffer L, Leano FT, Boxrud D, Outbreak of multidrug-resistant Salmonella Typhimurium DT104 in a daycare [abstract #2219]. In: Abstracts of the 39th International Conference on Antimicrobial Agents and Chemotherapy; San Francisco, California; 1999 Sep 26–29. Washington: American Society for Microbiology; 1999. p. 695.
- Davis MA, Hancock DD, Besser TE. Multiresistant clones of Salmonella enterica: the importance of dissemination. J Lab Clin Med. 2002;140:135–41. DOIPubMedGoogle Scholar
- Rajashekara G, Haverly E, Halvorson DA, Ferris KE, Lauer DC, Nagaraja KV. Multidrug-resistant Salmonella Typhimurium DT104 in poultry. J Food Prot. 2000;63:155–61.PubMedGoogle Scholar
- White DG, Zhao S, Sudler R, Ayers S, Friedman S, Chen S, The isolation of antibiotic-resistant salmonella from retail ground meats. N Engl J Med. 2001;345:1147–54. DOIPubMedGoogle Scholar
- Threlfall J, Hopkins KL, Ward LR. Diversification in Salmonella Typhimurium DT104 [letter]. Emerg Infect Dis. 2005;11:980–1.
- Dargatz DA, Fedorka-Cray PJ, Ladely SR, Ferris KE, Green AL, Headrick ML. Antimicrobial susceptibility patterns of Salmonella isolates from cattle in feedlots. J Am Vet Med Assoc. 2002;221:268–72. DOIPubMedGoogle Scholar
- Wells SJ, Fedorka-Cray PJ, Dargatz DA, Ferris K, Green A. Fecal shedding of Salmonella spp. by dairy cows on farm and at cull cow markets. J Food Prot. 2001;64:3–11.PubMedGoogle Scholar
- Fossler CP, Wells SJ, Kaneene JB, Ruegg PL, Warnick LD, Bender JB, Prevalence of Salmonella spp. on conventional and organic dairy farms. J Am Vet Med Assoc. 2004;225:567–73. DOIPubMedGoogle Scholar
- Liebana E, Garcia-Migura L, Clouting C, Clifton-Hadley FA, Lindsay E, Threlfall EJ, Multiple genetic typing of Salmonella enterica serotype Typhimurium isolates of different phage types (DT104, U302, DT204b, and DT49) from animals and humans in England, Wales, and Northern Ireland. J Clin Microbiol. 2002;40:4450–6. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 11, Number 12—December 2005
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Stephanie D. Wedel, Acute Disease Investigation and Control Section, Minnesota Department of Health, 625 Robert St N, PO Box 64975, St. Paul, MN 55164-0975, USA; fax: 651-201-5082
Top