Volume 13, Number 7—July 2007
Dispatch
Virus Detection and Monitoring of Viral Load in Crimean-Congo Hemorrhagic Fever Virus Patients
Figure 2
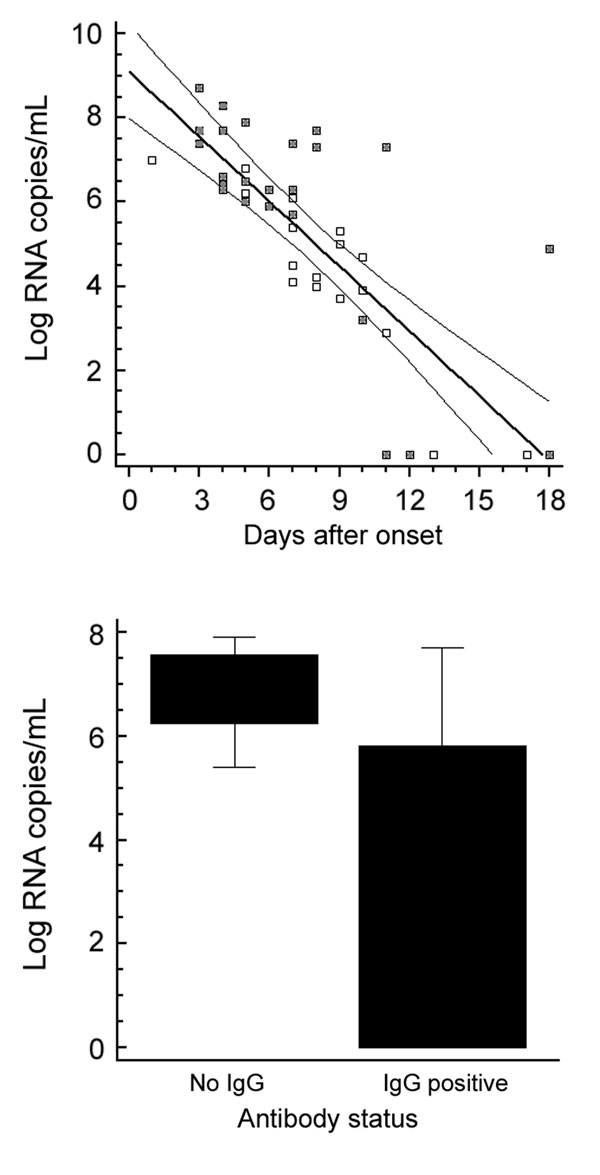
Figure 2. Clinical evaluation. Upper panel: plasma viral load over time in 44 samples from 17 patients. Samples from patients who died are marked with a filled square. Lower panel: plasma viral load in relation to antibody status in 16 samples with and 27 samples without detectable immunoglobulin G (IgG) antibodies. Only IgG status was taken as reference because only 2 patients had IgM without IgG. The difference of means between the 2 groups was highly significant (t test, p = 0.00005). Boxes indicate 25th through 75th percentiles; error bars indicate 5th and 95th percentiles. A) Oligonucleotides. The Figure 1 shows oligonucleotide binding sites from representative strains taken from the full CCHFV alignment used for assay design. The full alignment can be accessed at www.bni-hamburg.de. All sequences show the plus (coding) sense cDNA (i.e., the reverse complement of virus RNA). The forward primer RWCF and probes SE01 and SE03 are located on the plus strand. The reverse primer RWCR and probe SE0A are located on the minus strand. B) Analytical evaluation. Left and middle panel: amplification of a representative set of virus RNA (strain identity in legend) with probe SE01 only (left panel) and with the full set of 3 probes (middle). Note that all strains were detected with probe SE01 alone, but some strains showed weak signal. The additional probes increased the signal for such weak strains without lowering the signal for others. The 2 completely flat lines on both panels are negative controls. Left panel: probit analysis performed with the final assay protocol. The x-axis shows the calculated input concentration of synthetic virus RNA of strain BT-958 (EF123122), the y-axis shows the observed hit rates in 5 replicate reactions per concentration and the calculated detection probabilities according to the dose-response model. Upper and lower curves represent 95% confidence limits of the model. C) Clinical evaluation. Left panel: plasma viral load over time in 44 samples from 17 patients. Samples from patients who died are marked with a filled square. Right panel: plasma viral load in relation to antibody status in 16 samples with and 27 samples without detectable IgG antibodies. Only IgG status was taken as reference because only 2 patients had IgM without IgG. The difference of means between the 2 groups was highly significant (t test, p = 0.00005). Boxes indicate 25th through 75th percentiles; error bars indicate 5th and 95th percentiles. CCHFV, Crimean-Congo hemorrhagic fever virus.