Volume 15, Number 9—September 2009
Research
Predicting Phenotype and Emerging Strains among Chlamydia trachomatis Infections
Figure 4
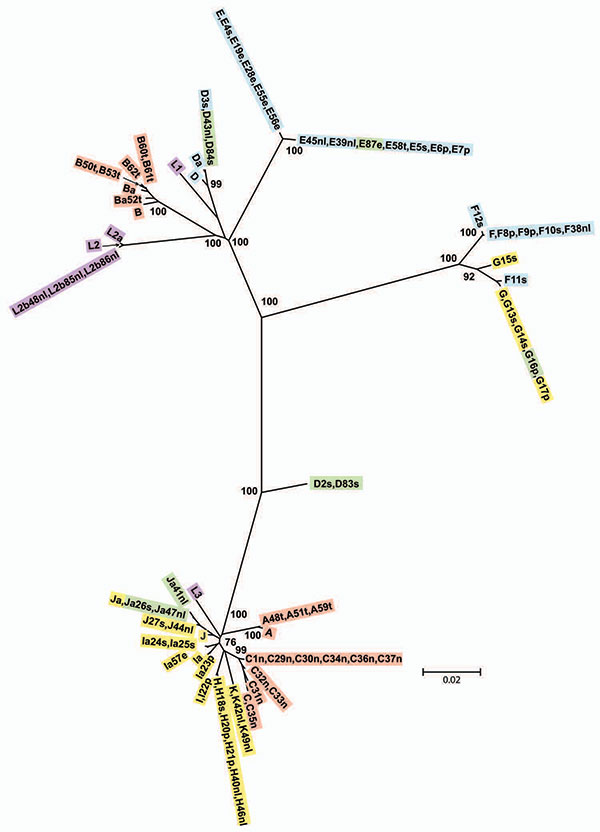
Figure 4. Minimum evolution tree for ompA. The tree was constructed using the matrix of pairwise differences between the 87 sequences by using the maximum composite likelihood method for estimating genetic distances. Numbers are bootstrap values (1,000 replicates) >70%. Lavender, invasive lymphogranuloma venereum (LGV); gold, noninvasive, nonprevalent sexually transmitted infection (STI) strains; red, trachoma strains; blue, noninvasive, highly prevalent STI strains; green, putative recombinant stains. Scale bar indicates number of substitutions per site.
Page created: December 07, 2010
Page updated: December 07, 2010
Page reviewed: December 07, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.