Volume 16, Number 11—November 2010
Dispatch
Multidrug-Resistant Salmonella enterica Serovar Infantis, Israel
Figure 2
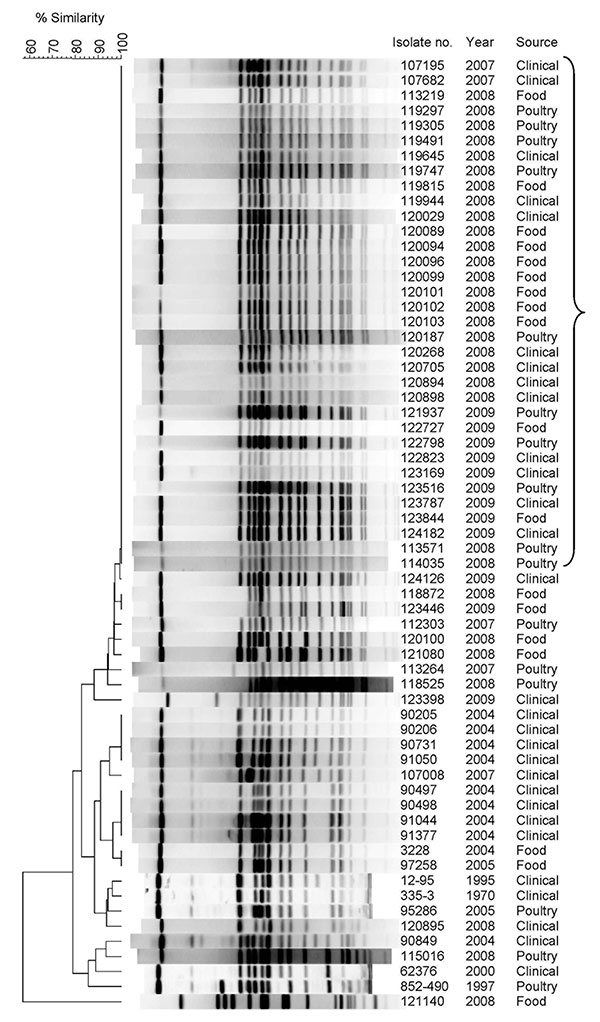
Figure 2. Pulsed-field gel electrophoresis (PFGE) patterns of Salmonella enterica serovar Infantis isolates from clinical, food, and poultry sources isolated in Israel, 1970–2009, showing a high degree of clonality. Isolate number, year of isolation, and source are indicated. Bracket indicates I1 pulsotype pattern. Macrodigestion performed using XbaI restriction enzyme and genetic similarity (in %) was based on dice coefficients. PFGE was conducted according to the standardized Salmonella protocol Centers for Disease Prevention and Control PulseNet as described (4) by using S. enterica ser. Braenderup H9812 strain as a molecular size standard. Because of space limitations, only 34/58 pulsotype I1 clones are shown. A complete list is provided in the Appendix Table.
References
- Tauxe RV. Salmonella Enteritidis and Salmonella Typhimurium: successful subtypes in the modern world. In: Scheld WM, Craig WA, Hughes JM, editors. Emerging infections 4. Washington: American Society for Microbiology; 1999. p. 37–52.
- Solnik-Isaac H, Weinberger M, Tabak M, Ben-David A, Shachar D, Yaron S. Quinolone resistance of Salmonella enterica serovar Virchow isolates from humans and poultry in Israel: evidence for clonal expansion. J Clin Microbiol. 2007;45:2575–9. DOIPubMedGoogle Scholar
- Weinberger M, Keller N. Recent trends in the epidemiology of non-typhoid Salmonella and antimicrobial resistance: the Israeli experience and worldwide review. Curr Opin Infect Dis. 2005;18:513–21. DOIPubMedGoogle Scholar
- Weinberger M, Solnik-Isaac H, Shachar D, Reisfeld A, Valinsky L, Andorn N, Salmonella enterica serotype Virchow: epidemiology, resistance patterns and molecular characterisation of an invasive Salmonella serotype in Israel. Clin Microbiol Infect. 2006;12:999–1005. DOIPubMedGoogle Scholar
- Fluit AC, Schmitz FJ. Class 1 integrons, gene cassettes, mobility, and epidemiology. Eur J Clin Microbiol Infect Dis. 1999;18:761–70. DOIPubMedGoogle Scholar
- Hopkins KL, Davies RH, Threlfall EJ. Mechanisms of quinolone resistance in Escherichia coli and Salmonella: recent developments. Int J Antimicrob Agents. 2005;25:358–73. DOIPubMedGoogle Scholar
- Gustafson RH. Antibiotics use in agriculture: an overview. In: Moats WA, editor. Agricultural uses of antibiotics. Washington: American Chemical Society; 1986. p. 1–6.
- Nogrady N, Kardos G, Bistyak A, Turcsanyi I, Meszaros J, Galantai Z, Prevalence and characterization of Salmonella Infantis isolates originating from different points of the broiler chicken–human food chain in Hungary. Int J Food Microbiol. 2008;127:162–7. DOIPubMedGoogle Scholar
- Nogrady N, Toth A, Kostyak A, Paszti J, Nagy B. Emergence of multidrug-resistant clones of Salmonella Infantis in broiler chickens and humans in Hungary. J Antimicrob Chemother. 2007;60:645–8. DOIPubMedGoogle Scholar
- Shahada F, Sugiyama H, Chuma T, Sueyoshi M, Okamoto K. Genetic analysis of multi-drug resistance and the clonal dissemination of beta-lactam resistance in Salmonella Infantis isolated from broilers. Vet Microbiol. 2010;140:136–41. Epub 2009 Jul 10. DOIPubMedGoogle Scholar