Volume 16, Number 12—December 2010
Dispatch
Wild Chimpanzees Infected with 5 Plasmodium Species
Cite This Article
Citation for Media
Abstract
Data are missing on the diversity of Plasmodium spp. infecting apes that live in their natural habitat, with limited possibility of human-mosquito-ape exchange. We surveyed Plasmodium spp. diversity in wild chimpanzees living in an undisturbed tropical rainforest habitat and found 5 species: P. malariae, P. vivax, P. ovale, P. reichenowi, and P. gaboni.
Despite ongoing and, in some regions, escalating morbidity and mortality rates associated with malaria-causing parasites, the evolutionary epidemiology of Plasmodium spp. is not well characterized. Classical studies of the blood pathogens of primates have found protozoa resembling human malaria parasites in chimpanzees and gorillas (1); however, these studies were limited to microscopy, negating conclusions regarding evolutionary relationships between human and ape parasites. Recent studies that used molecular approaches showed that captive and wild chimpanzees (Pan troglodytes) and lowland gorillas (Gorilla gorilla), as well as captive bonobos (Pan paniscus), harbor parasites broadly related to P. falciparum (2–5); wild and captive gorillas and captive bonobos and chimpanzees are sometimes infected with P. falciparum itself (4–6). Further, captive chimpanzees and bonobos have been shown to have malaria parasites related to human P. ovale and P. malariae (6–8); P. vivax has been identified in various monkeys and 1 semiwild chimpanzee (5,9). Recently, P. knowlesi, a simian malaria species, became the fifth human-infecting species (10), highlighting the possibility of transmission of new Plasmodium spp. from wild primates to humans.
To investigate the prevalence of different Plasmodium spp. in wild great apes living in their natural habitat (tropical rainforests), we analyzed tissue samples from 16 wild West African chimpanzees that died primarily of anthrax or respiratory disease in Taï National Park, Côte d’Ivoire. A generic real-time PCR that detects all known Plasmodium spp. was used to test all samples for the parasite. Sequence analysis of the CytB gene and small subunit rRNA genes was conducted for real-time PCR–positive samples to determine the strain present; 1,140 bp of the CytB gene and 765 bp of the 18S gene of the Plasmodium genome were amplified by classic PCR. Resulting products were sequenced either directly or after cloning for rRNA gene and when initial sequence information showed the possible presence of 2 different species (Table).
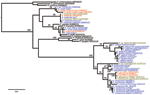
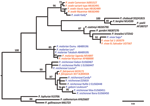
Phylogenetic analyses of sequences obtained confirmed the presence of 5 species: P. reichenowi and P. gaboni, which had been found previously (2,3); but also P. vivax, P. ovale, and P. malariae–like strains (Figure 1, Figure 2). The most prevalent species was P. reichenowi (6/16), which had representatives in subclusters P. gaboni and P. reichenowi. The other species were rare, seen only 1 (P. ovale and P. vivax) or 2 (P. malariae) times. Two chimpanzees showed co-infections with multiple Plasmodium spp. (Figure 1, Figure 2), 1 infected with P. reichenowi and a P. malariae–like strain and the other with P. reichenowi and P. gaboni.
Is the observed high prevalence of Plasmodium spp. typical for wild chimpanzees or related to reduced immune function associated with the severe infection that was the primary cause of death in each case? To investigate this question, we tested DNA extracted from fecal samples of apparently healthy chimpanzees collected over the past 8 years (n = 30) (11) of the same study population by using the generic real-time PCR followed by amplification of the CytB gene. Of these samples, 21 (70%) were positive for Plasmodium spp. by real-time PCR. Because of low copy numbers in feces, phylogentic analyses were limited to 2 samples in which P. reichenowi of the P. gaboni subcluster was confirmed.
To determine if the observed high prevalence of plasmodia was a site- or chimpanzee subspecies–specific phenomenon, we tested 30 randomly selected fecal samples of individually known apparently healthy wild Eastern chimpanzees from the Budongo Forest in Uganda. Overall prevalence of Plasmodium spp. was lower than in West African chimpanzees but still relatively high (40%); P. reichenowi and P. gaboni were identified in 3 samples.
Our results demonstrate that the prevalence of different Plasmodium spp. in wild chimpanzees is similar to that of untreated human populations in sub-Saharan Africa (www.who.int/malaria). Throughout sub-Saharan Africa, P. falciparum is more predominant in humans than are other Plasmodium spp. Considering the lack of clinical signs of malaria in chimpanzees from which fecal samples were collected and those that had died of respiratory disease or anthrax, Plasmodium spp. infections appear to be asymptomatic or at least nonlethal in wild chimpanzees. However, signs of illness are rarely observed in wild primates because infected animals often mask weakness to maintain social position and avoid attack by predators (12). Recently developed technologies for the noninvasive determination of temperature in wild chimpanzees may enable more effective examination of the relationship between the primary clinical feature of malaria (i.e., cyclical fevers) and Plasmodium spp. infection (13).
P. ovale was previously described from captive chimpanzees and P. malariae from captive chimpanzees and captive bonobos have been described (5–8). Our study results demonstrate that P. malariae and P. ovale occur in wild chimpanzees that inhabit pristine contiguous forest with extremely limited exposure to humans, suggesting the natural existence of these parasites in wild great apes.
Because of a Duffy-negative condition in 95%–99% of the human population in western and central continental Africa, transmission of P. vivax does not seem to occur. However, P. vivax infections are common in travelers returning from these areas (11). Even though we cannot totally exclude the possibility of introduction of P. vivax in the chimpanzee population through humans, our discovery of P. vivax in wild chimpanzees living exclusively within their natural habitat suggests that wild African apes may be a natural reservoir.
Our study shows the existence of P. reichenowi and related strains in wild chimpanzees as described for chimpanzees and gorilla by others (2–4,6). Infections with strains of the P. reichenowi group (sometimes referred to as the species P. gaboni, P. billbrayi, and P. billcollinsi) appear to occur widely in wild and captive great apes in Africa with some variation between chimpanzee subspecies from biogeographically distinct sites. The wild chimpanzees examined demonstrated no infections with classic human P. falciparum. This lack of infection is likely caused by low human presence in their habitat and, consequently, few or no infected vectors, low sample size, or a missing receptor in chimpanzees (14). More investigations are needed because recently P. falciparum infections have been described for 2 captive chimpanzees (6). The situation is clearer for captive and wild lowland gorillas (Gorilla gorilla) for which infections and receptors have recently been described (4). Infections have also been documented for captive bonobos (5).
Previous examination of the role of our closest phylogenetic relatives, the great apes, in the evolution and persistence of human plasmodia has been limited by a lack of data from wild ape populations where opportunities for human-mosquito-ape malaria exchange are minimal. Interpretation of patterns of malaria infection in captive ape populations, such as sanctuaries and zoos, must consider the ample opportunities for human-to-ape transmission of such parasites, negating the opportunity to investigate the evolutionary origins and public health–related risks of these parasites. Conversely, our examination of these parasites in wild chimpanzees with no contact to the periphery of the rainforest habitat (Technical Appendix Figure ) demonstrates that these apes are most likely naturally infected with P. ovale, P. vivax, and P. malariae, 3 types of plasmodia rarely observed in humans of the region. Whether wild great apes are the origin or reservoirs of these Plasmodium types requires further investigation. These results may have implications for global efforts to eradicate malaria in humans, including vaccine development based on animal variants of human parasites.
Mr Kaiser is a PhD candidate at the Robert Koch-Institute. He specializes in the development of PCR-based detection methods for various pathogens.
Acknowledgments
We thank the authorities of Côte d’Ivoire for long-term support, especially the Ministry of the Environment and Forests, the Ministry of Research, the directorship of the Taï National Park, and the Swiss Research Centre in Abidjan. In addition, we thank the Uganda Wildlife Authority and the Uganda National Council for Science and Technology for granting us permission to conduct this research, the assistants and students of the each of the chimpanzee projects for support during field observations, R. Wittig and J. Tesch for technical support, and S. Calvignac for phylogenetic analyses and helpful discussion.
This work was supported by the Robert Koch-Institute, the Max-Planck-Society, and Emory University. The Budongo Conservation Field Station receives core funding from the Royal Zoological Society of Scotland and the European Commission Ultra Sensitive Detection of Emerging Pathogens project (LSHB-CT-2006-037560).
References
- Coatney GR. The simian malarias: zoonoses, anthroponoses, or both? Am J Trop Med Hyg. 1971;20:795–803.
- Rich SM, Leendertz FH, Xu G, LeBreton M, Djoko CF, Aminake MN, The origin of malignant malaria. Proc Natl Acad Sci U S A. 2009;106:14902–7. DOIPubMedGoogle Scholar
- Ollomo B, Durand P, Prugnolle F, Douzery E, Arnathau C, Nkoghe D, A new malaria agent in African hominids. PLoS Pathog. 2009;5:e1000446. DOIPubMedGoogle Scholar
- Prugnolle F, Durand P, Neel C, Ollomo B, Ayala FJ, Arnathau C, African great apes are natural hosts of multiple related malaria species, including Plasmodium falciparum. Proc Natl Acad Sci U S A. 2010;107:1458–63. DOIPubMedGoogle Scholar
- Krief S, Escalante AA, Pacheco MA, Mugisha L, André C, Halwax M, On the diversity of malaria parasites in African apes and the origin of Plasmodium falciparum from bonobos. PLoS Pathog. 2010;6:e1000765. DOIPubMedGoogle Scholar
- Duval L, Fourment M, Nerrienet E, Rousset D, Sadeuh SA, Goodman SM, African apes as reservoirs of Plasmodium falciparum and the origin and diversification of the Laverania subgenus. Proc Natl Acad Sci U S A. 2010; [Epub ahead of print].
- Duval L, Nerrienet E, Rousset D, Sadeuh SA, Houze S, Fourment M, Chimpanzee malaria parasites related to Plasmodium ovale in Africa. PLoS ONE. 2009;4:e5520. DOIPubMedGoogle Scholar
- Hayakawa T, Arisue N, Udono T, Hirai H, Sattabongkot J, Toyama T, Identification of Plasmodium malariae, a human malaria parasite, in imported chimpanzees. PLoS ONE. 2009;4:e7412. DOIPubMedGoogle Scholar
- Escalante AA, Cornejo OE, Freeland DE, Poe AC, Durrego E, Collins WE, A monkey’s tale: the origin of Plasmodium vivax as a human malaria parasite. Proc Natl Acad Sci U S A. 2005;102:1980–5. DOIPubMedGoogle Scholar
- Galinski MR, Barnwell JW. Monkey malaria kills four humans. Trends Parasitol. 2009;25:200–4. DOIPubMedGoogle Scholar
- Jensen SA, Mundry R, Nunn CL, Boesch C, Leendertz FH. Non-invasive body temperature measurement of wild chimpanzees using fecal temperature decline. J Wildl Dis. 2009;45:542–6.PubMedGoogle Scholar
- Culleton RL, Mita T, Ndounga M, Unger H, Cravo PV, Paganotti GM, Failure to detect Plasmodium vivax in West and Central Africa by PCR species typing. Malar J. 2008;7:174. DOIPubMedGoogle Scholar
- Boesch C, Boesch-Achermann H. The chimpanzees of the Taï Forest: behavioural ecology and evolution. Oxford: Oxford University Press; 2000.
- Martin MJ, Rayner JC, Gagneux P, Barnwell JW, Varki A. Evolution of human–chimpanzee differences in malaria susceptibility: relationship to human genetic loss of N-glycolylneuraminic acid. Proc Natl Acad Sci U S A. 2005;102:12819–24. DOIPubMedGoogle Scholar
- Liu W, Li Y, Learn GH, Rudicell RS, Robertson JD, Keele BF, Origin of the human malaria parasite Plasmodium falciparum in gorillas. Nature. 2010;467:420–5. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleTable of Contents – Volume 16, Number 12—December 2010
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Fabian H. Leendertz, Robert Koch-Institute, Research Group Emerging Zoonoses, Nordufer 20, 13353, Berlin, Germany
Top