Volume 16, Number 3—March 2010
Dispatch
Banna Virus, China, 1987–2007
Cite This Article
Citation for Media
Abstract
Banna viruses (BAVs) have been isolated from pigs, cattle, ticks, mosquitoes, and human encephalitis patients. We isolated and analyzed 20 BAVs newly isolated in China; this finding extends the distribution of BAVs from tropical zone to north temperate climates and demonstrate regional variations in BAV phylogeny and mosquito species possibly involved in BAV transmission.
Banna virus (BAV), the prototype species of genus Seadornavirus within the family Reoviridae, has a genome composed of 12 segments of double-stranded RNA (1). BAV was initially isolated from persons with encephalitis and fever in Xishuangbanna, Yunnan Province, People’s Republic of China, in 1987 (2). Since then, BAV isolates have been obtained from pigs, cattle, and ticks in China (3,4) and from mosquitoes in Indonesia, China, and Vietnam. (5–7). BAV is a BioSafety Level 3 arboviral agent that is pathogenic to humans and may well be an emerging pathogen or undiagnosed cause of human viral encephalitis in some areas (1). Our objective was to describe new BAV isolates from China and to define the geographic distribution and the phylogenetic relationships of these isolates with reference to the previously described isolates.
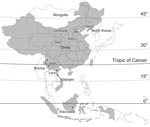
In this study, 20 new BAV isolates were obtained from mosquitoes collected from July through September during 2006 to 2007 at sites in Gansu Province (latitude 32°–35°N, 104°–107°E), Liaoning Province (39°–41°N, 123°–125°E), Shanxi Province (37°–38°N, 111°–113°E), and Inner Mongolia Province (41°–43°N, 121°–123°E) (Table, Figure 1). Mosquito samples were collected by using 12 V, 200 mA mosquito-trapping lamps (Wuhan Lucky Star Environmental Protection Tech Co., Ltd., Hubei, China) and by collecting mosquitoes from 8:00
Viruses were isolated and BAV isolates were identified using described procedures (8). Trizol reagent category no. 10296-028 (Invitrogen, Carlsbad, CA, USA) was used to extract total RNA. cDNA was prepared by using Ready-to-Go You-Prime First-Strand Beads Kit (Amersham Pharmacia Biotech, Piscatawy, NJ, USA) according to the manufacturer’s protocol. An 850-bp gene fragment from the 12th segment, which codes for the double-stranded RNA binding protein, was amplified from the cDNA of the BAV isolates by using previously published primers (9). PCR products were recovered by using purification kits (QIAGEN, Valencia, CA, USA), and then were inserted into pGEM-T easy vector (Promega, Madison, WI, USA). The insert sequence was determined by using M13 universal primers and an ABI Prism 3730 sequence analyzer (ABI, Shirley, NY, USA).
The genomic sequences of the 12th segment for the 20 new BAV strains were determined (GenBank accession nos. GQ331954–GQ331973). Phylogenetic trees were constructed from the amplified region of the 12th segment sequence by using the molecular evolutionary genetics analysis (MEGA) version 4 software (www.megasoftware.net) from aligned nucleotide sequences. We used neighbor-joining algorithms with 1,000 replicates for bootstrap support of tree groupings.
In this study, 38 BAV strains isolated during 1987–2007 were analyzed, which included 30 strains isolated in China (including 20 new BAV isolates first reported in this study and 10 previously described isolates from China (8,10–12), 3 strains from Indonesia, and 5 strains from Vietnam) (Table). Initial BAVs were isolated from Indonesia and Yunnan Province of China, which belong to tropical and subtropical zones (2,5).The new BAV isolates in our study were observed in Gansu, Shanxi, Liaoning, and Inner Mongolia provinces of China (northern China), which belong to the northern temperate zone. These strains represent a geographic distribution ranging from near the equator to latitude 45°N, extending from the tropical zone to the northern temperate zone (Figure 1). These data show that the distribution of BAVs is not limited to Southeast Asia but that it extends into northeast Asia as well.
Before our study, BAV had been isolated from 7 mosquito species in 2 genera (Culex tritaeniorhynchus, Cx. pipiens pallens, Cx. annulus, Cx. pseudovishnui, Cx. modestus, Anopheles sinensis, and Aedes vagus). To this list we now add 3 species in the genus Aedes (Ae. albopictus, Ae. vexans, and Ae. dorsalis) (Table), which are widely distributed in China and elsewhere.
Phylogenetic analysis based on the complete coding sequence (624 nt) of the 12th segment of the BAV genome indicated that the BAV isolates evaluated in this study could be divided into 2 phylogenetically different groups (Figure 2). Isolates from China and Vietnam are included in group A, and the strains from Indonesia are in group B. Group A could be further divided into 2 subgroups, A1 and A2. Subgroup A1 includes 4 independent clades that group according to the location of collection and represent viruses from northern China (Gansu, Shanxi, and Liaoning Provinces) as well as the Vietnam isolates. Subgroup A2 includes isolates mainly from southern China (Yunnan Province) and Vietnam, which is contiguous with Yunnan Province of China, as well as 2 isolates from northern China (BJ95-75/Beijing, and NM0706/Inner Mongolia) (Figure 1).
Our results demonstrate that BAV strains are distributed from the tropics of Southeast Asia to the northern temperate regions of China. These observations suggest that the distribution of BAV is wider than previously recognized and may be increasing. Consistent with previous observations (9), we report that BAV isolates from China cluster in group A and separate into subgroups mainly according to the geographic origin of the isolate; subgroup A1 is found in the north and subgroup A2 in the south. However, 2 isolates from northern China grouped in subgroup A2 (south), and 3 isolates from Vietnam grouped in subgroup A1 (north).
Considering that group A isolates are geographically located across the monsoon climate zone, where south-to-north winds are common during summer (13), BAV could be transferred in infected mosquitoes during this period by the prevailing winds that move from Southeast Asia to east Asia. In addition, bird migration, has been associated with the movement of other pathogens, and migration of infected birds through the east Asia–Australasia flyway (13), which traverses the region, may also account for this association. However, the transmission dynamics of BAVs are not well known. Further study is required to determine if winds and birds are involved in dispersal of the virus.
Our observations suggest that the public health impact of BAV may be underestimated. BAV appears to be actively circulating in areas where Japanese encephalitis virus (JEV) is endemic (14) and where C. tritaeniorhynchus, which is the main vector of JEV, is active. This mosquito also appears to be a common vector of BAV. The clinical symptoms of disease caused by the 2 viruses is similar, and BAV cases may be undetected during a JE outbreak. It has been reported that ≈14% of clinically diagnosed JE cases are BAV immunoglobulin (Ig) M positive (15), indicating that BAV epidemics may have occurred but have been clinically misdiagnosed as Japanese encephalitis. The apparent active transmission of BAV over a large geographic area, genetic variation between geographic regions, and the potential to cause severe disease underscore the need for additional surveillance, further characterization, and improved diagnostic systems worldwide.
Ms Liu is a PhD candidate at the State Key Laboratory for Infectious Disease Prevention and Control, Institute for Viral Disease Control and Prevention, Chinese Center for Disease Control and Prevention. She specializes in medical microbiology, and her current research interests include the detection and investigation of arboviruses and associated disease.
Acknowledgments
We thank Roger S. Nasci for consultation and assistance in the preparation and writing of this manuscript.
This work was supported by grants from the Ministry of Science and Technology, China (2003BA712A08-01; 2008ZX10004-008); Chinese CDC–US Centers for Disease Control Cooperative Agreement U19-GH000004; Development Grant of State Key Laboratory for Infectious Disease Prevention and Control (2008SKLID105); and Programme of Research Advance, China-France, B-06-04.
References
- Attoui H, Mohd Jaafar F, Micco P, Lamballerie X. Coltiviruses and seadornaviruses in North America, Europe, and Asia. Emerg Infect Dis. 2005;11:1673–9.PubMedGoogle Scholar
- Xu P, Wang Y, Zuo J, Lin J, Xu PM. New orbiviruses isolated from patients with unknown fever and encephalitis in Yunnan province [in Chinese]. Chin J Virol. 1990;6:27–33.
- Xu P, Wang Y, Zuo J, Lin J. Recovery of the same type of virus as human new orbivirus [in Chinese]. Chin J Virol. 1990;6:328–32.
- Li QP, Xe XC. First isolation of new orbivirus (Banna) from ticks and infected cattle sera in Xingjiang [in Chinese]. Endemic Dis Bull. 1992;7:64–9.
- Brown SE, Gorman M, Tesh B, Knudson L. Coltiviruses isolated from mosquitoes collected in Indonesia. Virology. 1993;196:363–7. DOIPubMedGoogle Scholar
- Chen B, Tao SJ. Arboviruses survey in China in recent ten years. Chin Med J (Engl). 1996;109:13–5.PubMedGoogle Scholar
- Nabeshima T, Thi Nga P, Guillermo P, Parquet MC, Yu F, Thanh TN, Isolation and molecular characterization of Banna virus from mosquitoes, Vietnam. Emerg Infect Dis. 2008;14:1276–9. DOIPubMedGoogle Scholar
- Zhai YG, Wang HQ, Xu HK, Meng WS, Chao YX, Fu SH, Investigation on arboviruses in Tianshui and Longnan regions of Gansu Province [in Chinese]. Chin J Zoonoses. 2008;24:59–63.
- Billoir F, Attoui H, Simon S, Gallian P, Micco P, Lamballerie X. Molecular diagnosis of group B coltiviruses infections. J Virol Methods. 1999;81:39–45. DOIPubMedGoogle Scholar
- Sun X, Fu S, Gong Z, Ge J, Meng W, Feng Y, Distribution of arboviruses and mosquitoes in northwestern Yunnan Province, China. Vector Borne Zoonotic Dis. 2009;PubMedGoogle Scholar
- Meng WS, Zhang JB, Sun XH, Liu QN, Chen Z, Zhai YG, Isolation and identification of arboviruses from mosquito pools in some regions of Liaoning Province, China [in Chinese]. Chin J Epidemiol. 2009;30:50–4.
- Xu LH, Tao SJ, Cao YX. Genotyping of the Chinese isolation of coltivirus [in Chinese]. Chin J Virol. 2003;17:346–9.
- Nga PT, del Carmen Parquet M, Cuong D, Ma P, Hasebe F, Inoue S, Shift in Japanese encephalitis virus (JEV) genotype circulating in northern Vietnam: implications for frequent introduction of JEV from Southeast Asia to East Asia. J Gen Virol. 2004;85:1625–31. DOIPubMedGoogle Scholar
- Erlanger TE, Weiss S, Keiser J, Utzinger J, Wiedenmayer K. Past, present, and future of Japanese encephalitis. Emerg Infect Dis. 2009;15:1–7. DOIPubMedGoogle Scholar
- Tao SJ, Chen BQ. Studies of coltivirus in China. Chin Med J (Engl). 2005;118:581–6.PubMedGoogle Scholar
Figures
Table
Cite This ArticleTable of Contents – Volume 16, Number 3—March 2010
| EID Search Options |
|---|
|
|
|
|
|
|