Volume 17, Number 11—November 2011
CME ACTIVITY - Research
Close Similarity between Sequences of Hepatitis E Virus Recovered from Humans and Swine, France, 2008−2009
Figure 3
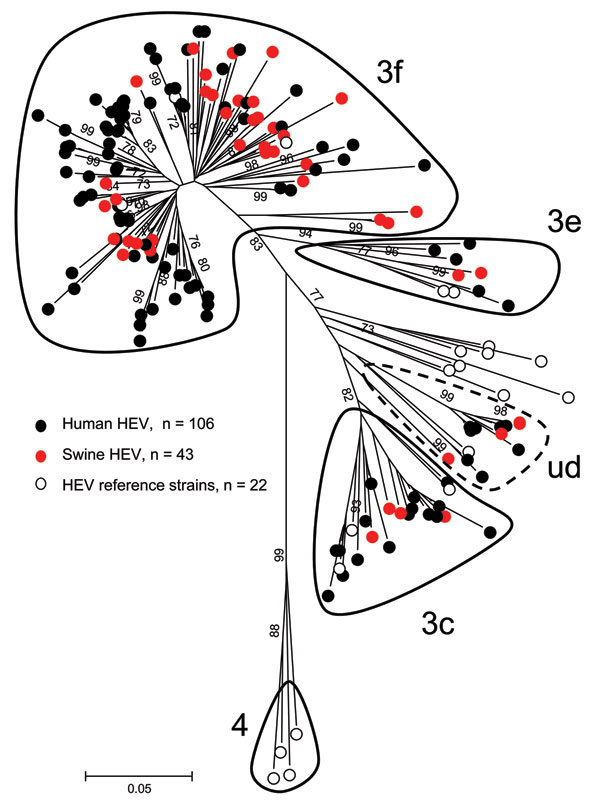
Figure 3. Phylogenetic tree of hepatitis E virus (HEV) detected in human and swine constructed by the neighbor-joining method with a bootstrap of 1,000 replicates based on the ClustalW alignment (MEGA4, www.megasoftware.net) of 204- to 306-nt sequences within open reading frame 2. The 106 HEV sequences recovered from patients from France are displayed as black dots (GenBank accession nos. JF730329−JF730434), the 43 HEV sequences recovered from swine from France are displayed as red dots (accession nos. JF718787−JF718829), and the 22 reference strains from GenBank are displayed as white dots (GenBank accession nos. in Table A1). Genotype 4, subtypes 3f, 3c, and 3e, as defined by Lu et al. (8), are encircled by a solid black line; undefined subtype is encircled by a dashed black line. Bootstrap values >70% are indicated on respective branches. Scale bar represents nucleotide substitutions per site.
References
- Panda SK, Thakral D, Rehman S. Hepatitis E virus. Rev Med Virol. 2007;17:151–80. DOIPubMedGoogle Scholar
- Purcell RH, Emerson SU. Hepatitis E: an emerging awareness of an old disease. J Hepatol. 2008;48:494–503. DOIPubMedGoogle Scholar
- Dalton HR, Bendall R, Ijaz S, Banks M. Hepatitis E: an emerging infection in developed countries. Lancet Infect Dis. 2008;8:698–709. DOIPubMedGoogle Scholar
- Emerson SU, Anderson D, Arankalle A, Meng XJ, Purdy M, Schlauder GG, Virus taxonomy VIIIth report of the ICTV. Hepevirus; 2004; 851–3.
- Lu L, Li C, Hagedorn CH. Phylogenetic analysis of global hepatitis E virus sequences: genetic diversity, subtypes and zoonosis. Rev Med Virol. 2006;16:5–36. DOIPubMedGoogle Scholar
- Pavio N, Meng X, Renou C. Zoonotic hepatitis E: animal reservoirs and emerging risks. Vet Res. 2010;41:46. DOIPubMedGoogle Scholar
- Li TC, Chijiwa K, Sera N, Ishibashi T, Etoh Y, Shinohara Y, Hepatitis E virus transmission from wild boar meat. Emerg Infect Dis. 2005;11:1958–60.PubMedGoogle Scholar
- Tei S, Kitajima N, Takahashi K, Mishiro S. Zoonotic transmission of hepatitis E virus from deer to human beings. Lancet. 2003;362:371–3. DOIPubMedGoogle Scholar
- Meng XJ, Purcell RH, Halbur PG, Lehman JR, Webb DM, Tsareva TS, A novel virus in swine is closely related to the human hepatitis E virus. Proc Natl Acad Sci U S A. 1997;94:9860–5. DOIPubMedGoogle Scholar
- Nakamura M, Takahashi K, Taira K, Taira M, Ohno A, Sakugawa H, Hepatitis E virus infection in wild mongooses of Okinawa, Japan: demonstration of anti-HEV antibodies and a full-genome nucleotide sequence. Hepatol Res. 2006;34:137–40. DOIPubMedGoogle Scholar
- Munné MS, Vladimirsky S, Otegui L, Castro R, Brajterman L, Soto S, Identification of the first strain of swine hepatitis E virus in South America and prevalence of anti-HEV antibodies in swine in Argentina. J Med Virol. 2006;78:1579–83. DOIPubMedGoogle Scholar
- dos Santos DRL, Vitral CL, de Paula VS, Marchevsky RS, Lopes JF, Gaspar AMC, Serological and molecular evidence of hepatitis E virus in swine in Brazil. Vet J. 2009;182:474–80. DOIPubMedGoogle Scholar
- Leblanc D, Ward P, Gagné M, Poitras E, Müller P, Trottier Y, Presence of hepatitis E virus in a naturally infected swine herd from nursery to slaughter. Int J Food Microbiol. 2007;117:160–6. DOIPubMedGoogle Scholar
- McCreary C, Martelli F, Grierson S, Ostanello F, Nevel A, Banks M. Excretion of hepatitis E virus by pigs of different ages and its presence in slurry stores in the United Kingdom. Vet Rec. 2008;163:261–5. DOIPubMedGoogle Scholar
- Banks M, Bendall R, Grierson S, Heath G, Mitchell J, Dalton H. Human and porcine hepatitis E virus strains, United Kingdom. Emerg Infect Dis. 2004;10:953–5.PubMedGoogle Scholar
- van der Poel WH, Verschoor F, van der Heide R, Herrera MI, Vivo A, Kooreman M, Hepatitis E virus sequences in swine related to sequences in humans, the Netherlands. Emerg Infect Dis. 2001;7:970–6. DOIPubMedGoogle Scholar
- Meng XJ, Halbur PG, Shapiro MS, Govindarajan S, Bruna JD, Mushahwar IK, Genetic and experimental evidence for cross-species infection by swine hepatitis E virus. J Virol. 1998;72:9714–21.PubMedGoogle Scholar
- Rutjes SA, Lodder WJ, Lodder-Verschoor F, van den Berg HHJL, Vennema H, Duizer E, Sources of hepatitis E virus genotype 3 in the Netherlands. Emerg Infect Dis. 2009;15:381–7. DOIPubMedGoogle Scholar
- Widén F, Sundqvist L, Matyi-Toth A, Metreveli G, Belák S, Hallgren G, Molecular epidemiology of hepatitis E virus in humans, pigs and wild boars in Sweden. Epidemiol Infect. 2011;139:361–71. DOIPubMedGoogle Scholar
- Reuter G, Fodor D, Forgách P, Kátai A, Szucs G. Characterization and zoonotic potential of endemic hepatitis E virus (HEV) strains in humans and animals in Hungary. J Clin Virol. 2009;44:277–81. DOIPubMedGoogle Scholar
- Zhang W, Yang S, Ren L, Shen Q, Cui L, Fan K, Hepatitis E virus infection in central China reveals no evidence of cross-species transmission between human and swine in this area. PLoS ONE. 2009;4:e8156. DOIPubMedGoogle Scholar
- Boutrouille A, Bakkali-Kassimi L, Crucière C, Pavio N. Prevalence of anti-hepatitis E virus antibodies in French blood donors. J Clin Microbiol. 2007;45:2009–10. DOIPubMedGoogle Scholar
- Mansuy JM, Legrand-Abravanel F, Calot JP, Peron JM, Alric L, Agudo S, High prevalence of anti-hepatitis E virus antibodies in blood donors from South West France. J Med Virol. 2008;80:289–93. DOIPubMedGoogle Scholar
- Rose N, Lunazzi A, Dorenlor V, Merbah T, Eono F, Eloit M, High prevalence of hepatitis E virus in French domestic pigs. Comp Immunol Microbiol Infect Dis. 2001;34:419–27. DOIPubMedGoogle Scholar
- Colson P, Borentain P, Queyriaux B, Kaba M, Moal V, Gallian P, Pig liver sausage as a source of hepatitis E virus transmission to humans. J Infect Dis. 2010;202:825–34. DOIPubMedGoogle Scholar
- Ministére de l’Agriculture et de la Péche. Consommation de viande en France. AGRESTE, 2008. 2008/n°29 [cited 2008 Jun]. http://www.agreste.agriculture.gouv.fr/IMG/pdf/syntheseviande0806.pdf
- Cooper K, Huang FF, Batista L, Rayo CD, Bezanilla JC, Toth TE, Identification of genotype 3 hepatitis E virus (HEV) in serum and fecal samples from pigs in Thailand and Mexico, where genotype 1 and 2 HEV strains are prevalent in the respective human populations. J Clin Microbiol. 2005;43:1684–8. DOIPubMedGoogle Scholar
- Tamura K, Dudley J, Nei M, Kumar S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol. 2007;24:1596–9. DOIPubMedGoogle Scholar
- French Ministry of the Economy. Finance and Industry. Decree no. 2009-1707 of 30 December 2009. Paris: Official Journal of the French Republic, Direction de l’information légale et administrative; 2010.
- Legrand-Abravanel F, Mansuy J, Dubois M, Kamar N, Peron J, Rostaing L, Hepatitis E virus genotype 3 diversity, France. Emerg Infect Dis. 2009;15:110–4. DOIPubMedGoogle Scholar
- Renou C, Moreau X, Pariente A, Cadranel J, Maringe E, Morin T, A national survey of acute hepatitis E in France. Aliment Pharmacol Ther. 2008;27:1086–93. DOIPubMedGoogle Scholar
- Peralta B, Mateu E, Casas M, de Deus N, Martín M, Pina S. Genetic characterization of the complete coding regions of genotype 3 hepatitis E virus isolated from Spanish swine herds. Virus Res. 2009;139:111–6. DOIPubMedGoogle Scholar
- Adlhoch C, Wolf A, Meisel H, Kaiser M, Ellerbrok H, Pauli G. High HEV presence in four different wild boar populations in East and West Germany. Vet Microbiol. 2009;139:270–8. DOIPubMedGoogle Scholar
- Di Bartolo I, Ponterio E, Castellini L, Ostanello F, Ruggeri FM. Viral and antibody HEV prevalence in swine at slaughterhouse in Italy. Vet Microbiol. 2011;149:330–8. DOIPubMedGoogle Scholar
- Forgách P, Nowotny N, Erdélyi K, Boncz A, Zentai J, Szucs G, Detection of hepatitis E virus in samples of animal origin collected in Hungary. Vet Microbiol. 2010;143:106–16. DOIPubMedGoogle Scholar
- Grandadam M, Tebbal S, Caron M, Siriwardana M, Larouze B, Koeck JL, Evidence for hepatitis E virus quasispecies. J Gen Virol. 2004;85:3189–94. DOIPubMedGoogle Scholar
- de Deus N, Casas M, Peralta B, Nofrarías M, Pina S, Martín M, Hepatitis E virus infection dynamics and organic distribution in naturally infected pigs in a farrow-to-finish farm. Vet Microbiol. 2008;132:19–28. DOIPubMedGoogle Scholar
- Bouwknegt M, Rutjes SA, Reusken CB, Stockhofe-Zurwieden N, Frankena K, de Jong MCM, The course of hepatitis E virus infection in pigs after contact-infection and intravenous inoculation. BMC Vet Res. 2009;5:7. DOIPubMedGoogle Scholar