Volume 18, Number 10—October 2012
Dispatch
Influenza Virus Infection in Nonhuman Primates
Cite This Article
Citation for Media
Abstract
To determine whether nonhuman primates are infected with influenza viruses in nature, we conducted serologic and swab studies among macaques from several parts of the world. Our detection of influenza virus and antibodies to influenza virus raises questions about the role of nonhuman primates in the ecology of influenza.
Worldwide, infections with influenza A viruses are associated with substantial illness and death among mammals and birds. Public health and research have placed major focus on understanding the pathogenicity of different influenza virus strains and characterizing new influenza vaccines. Nonhuman primates (NHPs), including macaques, have become popular experimental models for studying the pathogenesis and immunology of seasonal and emerging influenza viruses. NHPs readily seroconvert after experimental inoculation with seasonal influenza virus and have been used to test candidate vaccines for strains of human and avian origin. Like humans, macaques infected with influenza virus exhibit fever, malaise, nasal discharge, and nonproductive cough; virus replication can be detected in the nasal passages and respiratory tract (1,2). However, whether NHPs are infected with influenza viruses in nature remains unknown.
Over the past decade, we have focused on the role of pet and performing monkeys in disease transmission throughout Asia. Commonly trapped in the wild, these monkeys might be sold at wet markets, the putative source of several zoonotic outbreaks (3), where they might be caged next to any number of animal species (Figure 1, panel A) (4). Pet and performing monkeys are likely conduits for cross-species transmission of respiratory pathogens like influenza viruses because of their close and long-term contact with their owners, audiences, domestic animals, wild animals, and birds (Figure 1, panel B) (5). However, the breadth and diversity of this interface presents a challenge for monitoring the emergence of infectious diseases. We have approached this challenge by conducting longitudinal studies at several sites and collecting biological samples and behavioral data representing various contexts of human–NHP contact (4–7). We used these historical and newly acquired samples, representing various countries and contexts of human–macaque contact, to determine whether NHPs are infected with influenza viruses in nature.
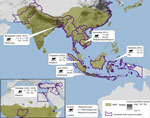
As part of our decade-long longitudinal studies, ≈200 serum samples were collected from macaques. These included pet macaques (Macaca nigra, M. nigrescens, M. hecki) from Sulawesi, Indonesia; performing macaques from Java, Indonesia (M. fascicularis) and from Bangladesh (M. mulatta); M. fascicularis macaques from the Bukit Timah and Central Catchment Nature Reserves in Singapore, where they freely interact with wild avian fauna and visitors (occasionally entering residential areas) (7); M. sylvanus macaques from the Upper Rock Nature Reserve in Gibraltar, where international tourists frequently use food to entice the macaques to climb about their heads and shoulders (6); and free-ranging macaques (M. fascicularis and M. nemestrina) at temple shrines or M. fascicularis macaques that range throughout a wildlife rescue center and nearby villages in Cambodia (Figure 2). Serum was collected and stored as described (8). All samples were stored on cold packs in the field and transferred to dry ice for shipment to the United States, where they were then stored at −80°C.
For initial screening for antibodies against influenza virus, serum samples were treated with receptor-destroying enzyme as described (9) and tested by using a multispecies Influenza A Virus NP Antibody Inhibition Test (Virusys Corporation, Taneytown, MD, USA) according to manufacturer’s instructions. ELISA results indicated nucleocapsid protein antibodies against influenza in samples from macaques from Cambodia (29.2%), Singapore (16.7%), Sulawesi (16.1%), Bangladesh (13.3%), and Java (6.0%) (Table 1). Antibodies were detected in animals 1–10 years of age at the time of sampling. No influenza virus–specific antibodies were detected from the 73 total samples from Gibraltar, perhaps because persons with influenza virus infection infrequently travel to the Upper Rock Reserve (healthy-visitor effect) (10) or perhaps because monkeys from Gibraltar are less susceptible to infection. Seroprevalence of antibodies against influenza A, by site and collection year, human and NHP population densities, and prevalence of avian influenza viruses are shown in Figure 2.
Serum samples that were positive by ELISA were also screened by hemagglutination-inhibition assay as described (9). Based on the year and location of NHP sample collection, the estimated ages of the NHPs at the time of sample collection, and the presence of avian H5 and H9 influenza viruses in many of these countries during the sampling period (11–13), a panel of human vaccine strains and avian influenza virus strains was used in the hemagglutination-inhibition assay. Although not all ELISA-positive serum samples could be subtyped, antibodies against seasonal subtype H1N1 and H3N2 influenza A strains were detected from macaques in Bangladesh, Singapore, Java, and Sulawesi (Table 2). Of the performing macaques in Bangladesh, 2 had antibodies against A/chicken/Bangladesh/5473/2010, a strain of G1 clade subtype H9N2 avian influenza virus. Subtype H9N2 influenza viruses are prevalent in poultry in Bangladesh (14) and have been detected in humans (12). We did not detect antibodies against highly pathogenic avian influenza subtype H5 viruses, which might not be surprising given our relatively small sample size (Table 2). Also given the small sample size, we were unable to perform microneutralization studies, which would be useful to perform with future samples.
In 2011, to determine whether any macaques were actively infected with influenza virus, we collected oral swabs from 48 monkeys in Cambodia to test for influenza virus by real-time reverse transcription PCR as described (8). In brief, the inside of the anesthetized and immobilized monkeys’ mouths (cheeks, tongue, and gums) were swabbed. Swabs were immediately placed into viral transport media, stored, and shipped as previously described. RNA was isolated by using an Ambion MagMAX-96 AI/ND Viral RNA Isolation Kit (Life Technologies Corporation, Grand Island, NY, USA) on a Kingfisher Flex system (Thermo Fisher Scientific, Waltham, MA, USA). Viral RNA was analyzed in a Bio-Rad CFX96 Real-Time PCR Detection System and a C1000 Thermocycler (Bio-Rad, Hercules, CA, USA) with TaqMan Fast Virus 1-Step Master Mix (Applied Biosystems, Foster City, CA, USA) and the InfA primer/probe sets as described (15). Of the 48 respiratory samples, 1 (2.1%) was positive for influenza virus; cycle threshold value was 38 (limit of detection is 40). Attempts to amplify longer PCR fragments of matrix, hemagglutinin, or neuraminidase genes or to isolate the virus by blind passage in embryonated chicken eggs or MDCK cells were unsuccessful.
Our results indicate that NHPs that have contact with humans can be naturally infected with seasonal endemic human influenza viruses and with emerging pandemic-risk avian influenza viruses. We found serologic evidence of infection in several countries, contexts, and macaque species. Preliminary real-time reverse transcription PCR results also pointed to the presence of virus in a buccal swab from an adult macaque from Cambodia, indicating active infection at the time of sampling. On the basis of results from this study, it seems that pet, performing, and free-ranging macaques are susceptible to influenza virus infection. Given the close relationship between humans and NHPs in areas of the world where avian and human influenza viruses cocirculate, further surveillance of these populations is warranted. The ability to detect and eventually isolate strains of influenza virus currently infecting NHPs and humans at the animal–human interface is paramount to understanding how NHP–human interactions can affect the genetics, transmission, and public health risk for infection with influenza A viruses.
Dr Karlsson is a postdoctoral research associate in the Department of Infectious Diseases at St. Jude Children’s Research Hospital, Memphis, Tennessee, USA. His research is focused on surveillance of respiratory and gastrointestinal viruses and on understanding how nutrition can affect virus–host interactions and emerging infectious diseases.
Acknowledgments
We thank R. Webby and R. Webster for critical discussions regarding the manuscript. We are particularly grateful to the following communities, temple committees, and government agencies in the areas where we have been sampling monkeys for years: Lembaga Ilmu Pengetahuan, Indonesia; S. Chan and the staff of the Central Nature Reserve, National Parks Board, Singapore; M. Pizarro, J. Cortes, and the staff of the Gibraltar Ornithological & Natural History Society and the Government of Gibraltar; S. Begum, K. Hasan, and the students and faculty of the Department of Zoology, Jahangirnagar University; and C. Kimleng, S. Bunnary, T. Sothearos, Sisiket, H. Davun, K. Pal, D. Cohn, A. Fuentes, J. Supriatna, R. Babo, Y. Paramastri, E. Iskandar, J. Froehlich, L. Engel, H. Engel, and L. Johnson for supporting and participating in this research. We also thank S. Krauss and M. Ducatez for graciously providing the panel of human vaccine strains and avian influenza virus strains that were used in the hemagglutination-inhibition assay.
This work was supported by the National Institutes of Health National Institute of Allergy and Infectious Diseases, contract no. HHSN266200700005C, the American Lebanese Syrian Associated Charities, NIH National Center for Research Resources grant P51 RR000166RR 02S014, National Institutes of Health National Institute of Allergy and Infectious Diseases grants R01 AI078229 and R03 AI064865, Defense Advanced Research Projects Agency N66001-02-C-8072, the University of Toronto Connaught Fund, the Chicago Zoological Society, and the University of New Mexico Research Allocations Committee.
References
- Berendt RF. Simian model for the evaluation of immunity to influenza. Infect Immun. 1974;9:101–5.PubMedGoogle Scholar
- Bodewes R, Rimmelzwaan GF, Osterhaus AD. Animal models for the preclinical evaluation of candidate influenza vaccines. Expert Rev Vaccines. 2010;9:59–72. DOIPubMedGoogle Scholar
- Webster RG. Wet markets—a continuing source of severe acute respiratory syndrome and influenza? Lancet. 2004;363:234–6. DOIPubMedGoogle Scholar
- Schillaci MA, Jones-Engel L, Engel GA, Paramastri Y, Iskandar E, Wilson B, Prevalence of enzootic simian viruses among urban performance monkeys in Indonesia. Trop Med Int Health. 2005;10:1305–14. DOIPubMedGoogle Scholar
- Schillaci MA, Jones-Engel L, Engel GA, Kyes RC. Exposure to human respiratory viruses among urban performing monkeys in Indonesia. Am J Trop Med Hyg. 2006;74:716–9. PubMedGoogle Scholar
- Fuentes A, Shaw E, Cortes J. Qualitative assessment of macaque tourist sites in Padangtegal, Bali, Indonesia, and the Upper Rock Nature Reserve, Gibraltar. Int J Primatol. 2007;28:1143–58. DOIGoogle Scholar
- Sha JC, Gumert M, Lee B, Fuentes A, Chan S, Jones-Engel L. Status of the long-tailed macaque (Macaca fascicularis) in Singapore and implications for management. Biodivers Conserv. 2009;18:2909–26. DOIGoogle Scholar
- Jones-Engel L, Steinkraus KA, Murray SM, Engel GA, Grant R, Aggimarangsee N, Sensitive assays for simian foamy viruses reveal a high prevalence of infection in commensal, free-ranging Asian monkeys. J Virol. 2007;81:7330–7. DOIPubMedGoogle Scholar
- Palmer DF, Dowdle WR, Coleman MT, Schild GC. Advanced laboratory techniques for influenza diagnosis. Atlanta: Centers for Disease Control, US Department of Health, Education and Welfare.; 1975.
- Li CY, Sung FC. A review of the healthy worker effect in occupational epidemiology. Occup Med (Lond). 1999;49:225–9. DOIPubMedGoogle Scholar
- Perdue ML, Swayne DE. Public health risk from avian influenza viruses. Avian Dis. 2005;49:317–27. DOIPubMedGoogle Scholar
- Malik Peiris JS. Avian influenza viruses in humans. Rev Sci Tech. 2009;28:161–73.PubMedGoogle Scholar
- Kalthoff D, Globig A, Beer M. (Highly pathogenic) avian influenza as a zoonotic agent. Vet Microbiol. 2010;140:237–45. DOIPubMedGoogle Scholar
- Negovetich NJ, Feeroz MM, Jones-Engel L, Walker D, Alam SMR, Hasan K, Live bird markets of Bangladesh: H9N2 viruses and the near absence of highly pathogenic H5N1 influenza. PLoS ONE. 2011;6:e19311. DOIPubMedGoogle Scholar
- World Health Organization. CDC protocol of realtime RTPCR for swine influenza A(H1N1). 2009 Apr 30 [cited 2012 Apr 11]. http://www.who.int/csr/resources/publications/swineflu/CDCrealtimeRTPCRprotocol_20090428.pdf
Figures
Tables
Cite This ArticleTable of Contents – Volume 18, Number 10—October 2012
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Lisa Jones-Engel, University of Washington–NRPC, 1705 Pacific St NE, HSB I-039, Seattle, WA 98195, USA
Top