Volume 18, Number 5—May 2012
Letter
Clonal Spread of Geomyces destructans among Bats, Midwestern and Southern United States
Figure
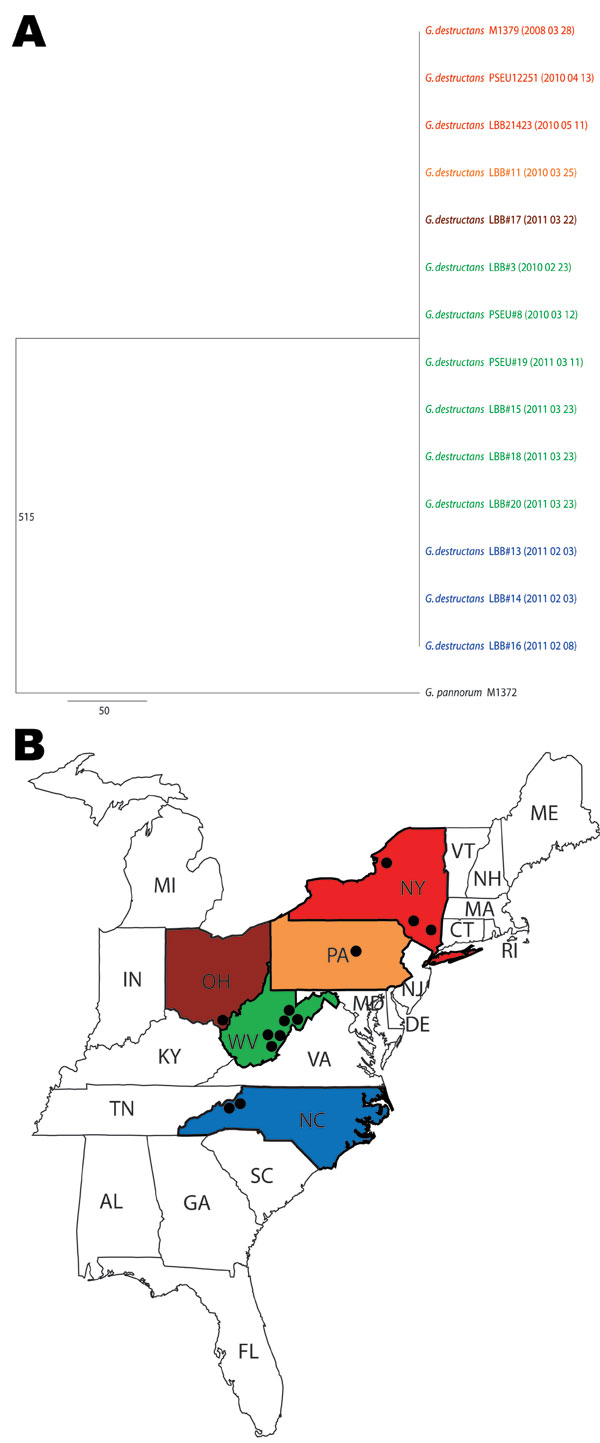
Figure. . . A) Consensus maximum-parsimony tree of 8 concatenated gene fragments of Geomyces destructans. Data were derived from 13 G. destructans test isolates. G. destructans M1379 and G. pannorum M1378 were used as controls in this study; they were described in an earlier report (3). The number 515 on the branch indicates the total number of variable nucleotide positions (of 4,722 nt) separating G. pannorum M1372 from the clonal genotype of G. destructans. Isolation dates are shown in parentheses (YYYY MMM DD). Scale bar indicates nucleotide substitutions per site. B) States color-matched to panel A to show where G. destructans isolates were found; dots indicate locations of positive test results.
References
- Blehert DS, Hicks AC, Behr M, Meteyer CU, Berlowski-Zier BM, Buckles EL, Bat white-nose syndrome: an emerging fungal pathogen? Science. 2009;323:227. DOIPubMedGoogle Scholar
- Chaturvedi V, Springer DJ, Behr MJ, Ramani R, Li X, Peck MK, Morphological and molecular characterizations of psychrophilic fungus Geomyces destructans from New York bats with white nose syndrome (WNS). PLoS ONE. 2010;5:e10783. DOIPubMedGoogle Scholar
- Rajkumar SS, Li X, Rudd RJ, Okoniewski JC, Xu J, Chaturvedi S, Clonal genotype of Geomyces destructans among bats with white nose syndrome, New York, USA. Emerg Infect Dis. 2011;17:1273–6. DOIPubMedGoogle Scholar
- Frick WF, Pollock JF, Hicks AC, Langwig KE, Reynolds DS, Turner GG, An emerging disease causes regional population collapse of a common North American bat species. Science. 2010;329:679–82. DOIPubMedGoogle Scholar
- Rachowicz LJ, Hero J-M, Alford RA, Taylor JW, Morgan JAT, Vredenburg VT, The novel and endemic pathogen hypotheses: competing explanations for the origin of emerging infectious diseases of wildlife. Conserv Biol. 2005;19:1441–8. DOIGoogle Scholar
- Puechmaille SJ, Wibbelt G, Korn V, Fuller H, Forget F, Muhldorfer K, Pan-European distribution of white-nose syndrome fungus (Geomyces destructans) not associated with mass mortality. PLoS ONE. 2011;6:e19167. DOIPubMedGoogle Scholar
- Archie EA, Luikart G, Ezenwa VO. Infecting epidemiology with genetics: a new frontier in disease ecology. Trends Ecol Evol. 2009;24:21–30. DOIPubMedGoogle Scholar
- Harvell CD, Mitchell CE, Ward JR, Altizer S, Dobson AP, Ostfeld RS, Climate warming and disease risks for terrestrial and marine biota. Science. 2002;296:2158–62. DOIPubMedGoogle Scholar
- Lebarbenchon C, Brown SP, Poulin R, Gauthier-Clerc M, Thomas F. Evolution of pathogens in a man-made world. Mol Ecol. 2008;17:475–84. DOIPubMedGoogle Scholar
- Lorch JM, Meteyer CU, Behr MJ, Boyles JG, Cryan PM, Hicks AC, Experimental infection of bats with Geomyces destructans causes white-nose syndrome. Nature. 2011;480:376–8. DOIPubMedGoogle Scholar
Page created: April 10, 2012
Page updated: April 10, 2012
Page reviewed: April 10, 2012
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.