Volume 18, Number 5—May 2012
Letter
Clonal Spread of Geomyces destructans among Bats, Midwestern and Southern United States
To the Editor: Bat geomycosis (white nose syndrome) is caused by the psychrophilic fungus Geomyces destructans, which has rapidly spread in the United States and Canada since it was first reported from Albany, New York (1,2). In 2011, a single genotype of G. destructans was found in bats with geomycosis in different parts of New York (3). The findings raised the possibility of clonal spread of a new pathogen with serious implications for the survival of the affected bat populations (4). To provide information for devising conservation measures, we explored whether this emerging infectious disease is caused by a novel pathogen (5). To do so, we genotyped G. destructans isolates from the midwestern and southern United States.
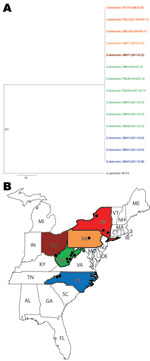
During 2010 and 2011, a total of 11 cultures of G. destructans were isolated and identified: 1 each from Pennsylvania and Ohio, 3 from North Carolina, and 6 from West Virginia (Figure). The cultures came from 8 little brown bats (Myotis lucifugus) and 3 tri-colored bats (Perimyotis subflavus). Two recent G. destructans isolates from New York and 1 G. pannorum isolate were included as controls. Genomic DNA was prepared from fungal growth by the conventional glass bead treatment, phenol–chloroform extraction, and ethanol precipitation. PCR amplifications of 8 G. destructans gene fragments (ALR, Bpntase, DHC1, GPHN, PCS, POB3, SRP72, and VPS13) were performed as described (3). The amplicons were sequenced and nucleotides were aligned by Sequencher 4.8 (www.genecodes.com); phylogenetic analyses were done using PAUP*4.0 software (www.sinauer.com).
A total of 4,722 nt sequences were obtained from 8 gene fragments of 13 G. destructans isolates (GenBank accession nos. JQ029780–JQ029883) and 1 G. pannorum isolate (GenBank accession nos. HQ834330, HQ834347, HQ834364, HQ834381, HQ834398, HQ834415, HQ834432, and HQ834449). Multiple alignments of these sequences showed 100% identity, and the aligned nucleotides matched perfectly with those of earlier G. destructans sequences for the same gene fragments analyzed from New York isolates (3). The nucleotide alignments of 8 sequences showed differences from those obtained from the closely related fungus, G. pannorum. Maximum-parsimony trees were generated by using sequences from each gene fragment. These trees showed a single clade of G. destructans strains distinct from G. pannorum; similar topologies were obtained when different phylogenetics methods were used for analysis (details not shown). A consensus maximum-parsimony tree derived from the 8 concatenated gene fragments also showed a single clade of G. destructans isolates from New York and the midwestern and southern United States (Figure).
The data obtained in this study strongly indicate further clonal spread of G. destructans from its origin near Albany, New York. The locations in which G. destructans was detected in the current study were spread across 5 states, which were >800 miles from Albany. The test isolates were compared with a New York isolate from 2008, which provided a 4-year temporal variation in our sampling. Bats of 2 species were positive for G. desctructans in the current samples, and they yielded the same G. destructans genotype. Thus, there is evidence for host-independent spread of a single clone of G. destructans.
These data would support the novel-pathogen hypothesis for the origin of bat geomycosis (5). However, these conclusions are based on limited sampling because isolations of G. destructans from affected bats are uncommon. The demonstration of pure fungal culture in the affected animals is still not the standard for geomycosis diagnostics, and most geomycosis is confirmed by bat morphologic appearance or histopathologic examination. Additionally, our phylogenetics analyses were limited to ≈5 kbp of fungal genomes, which could lead to sampling bias (3). Ideally, a large number of G. destructans isolates, including isolates from Europe, and additional polymorphic markers would be needed to determine the novel or local origin of this pathogen (6,7).
The environmental factors that led to introduction or reemergence of G. destructans in mines and caves remain unknown, and their contribution in the spread of the fungus through air, water, and soil is yet to be determined (8). Although no direct evidence has emerged, a role for anthropomorphic activities (occupational or recreational) in this spread is a distinct possibility (9). We provide genetic evidence for further spread of a single genotype of G. destructans from Albany, New York, to locations in the midwestern and southern United States. Experimental transmission of geomycosis from infected bats to healthy bats by direct contact has recently been confirmed (10). Therefore, G. destructans might be rapidly spreading along summer and winter migration routes of bats, which present ample opportunities for mixing of healthy and diseased animals.
Acknowledgments
We thank Xiaojiang Li and Sudha Chaturvedi for fungal characterization and comments on the manuscript.
This study was supported in part with funds from the US Fish and Wildlife Service.
References
- Blehert DS, Hicks AC, Behr M, Meteyer CU, Berlowski-Zier BM, Buckles EL, Bat white-nose syndrome: an emerging fungal pathogen? Science. 2009;323:227. DOIPubMedGoogle Scholar
- Chaturvedi V, Springer DJ, Behr MJ, Ramani R, Li X, Peck MK, Morphological and molecular characterizations of psychrophilic fungus Geomyces destructans from New York bats with white nose syndrome (WNS). PLoS ONE. 2010;5:e10783. DOIPubMedGoogle Scholar
- Rajkumar SS, Li X, Rudd RJ, Okoniewski JC, Xu J, Chaturvedi S, Clonal genotype of Geomyces destructans among bats with white nose syndrome, New York, USA. Emerg Infect Dis. 2011;17:1273–6. DOIPubMedGoogle Scholar
- Frick WF, Pollock JF, Hicks AC, Langwig KE, Reynolds DS, Turner GG, An emerging disease causes regional population collapse of a common North American bat species. Science. 2010;329:679–82. DOIPubMedGoogle Scholar
- Rachowicz LJ, Hero J-M, Alford RA, Taylor JW, Morgan JAT, Vredenburg VT, The novel and endemic pathogen hypotheses: competing explanations for the origin of emerging infectious diseases of wildlife. Conserv Biol. 2005;19:1441–8. DOIGoogle Scholar
- Puechmaille SJ, Wibbelt G, Korn V, Fuller H, Forget F, Muhldorfer K, Pan-European distribution of white-nose syndrome fungus (Geomyces destructans) not associated with mass mortality. PLoS ONE. 2011;6:e19167. DOIPubMedGoogle Scholar
- Archie EA, Luikart G, Ezenwa VO. Infecting epidemiology with genetics: a new frontier in disease ecology. Trends Ecol Evol. 2009;24:21–30. DOIPubMedGoogle Scholar
- Harvell CD, Mitchell CE, Ward JR, Altizer S, Dobson AP, Ostfeld RS, Climate warming and disease risks for terrestrial and marine biota. Science. 2002;296:2158–62. DOIPubMedGoogle Scholar
- Lebarbenchon C, Brown SP, Poulin R, Gauthier-Clerc M, Thomas F. Evolution of pathogens in a man-made world. Mol Ecol. 2008;17:475–84. DOIPubMedGoogle Scholar
- Lorch JM, Meteyer CU, Behr MJ, Boyles JG, Cryan PM, Hicks AC, Experimental infection of bats with Geomyces destructans causes white-nose syndrome. Nature. 2011;480:376–8. DOIPubMedGoogle Scholar
Figure
Cite This ArticleRelated Links
Table of Contents – Volume 18, Number 5—May 2012
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Vishnu Chaturvedi, Mycology Laboratory, Wadsworth Center, New York State Department of Health, 120 New Scotland Ave, Albany, NY 12208, USA
Top