Volume 3, Number 2—June 1997
Synopsis
Hantaviruses: A Global Disease Problem
Figure
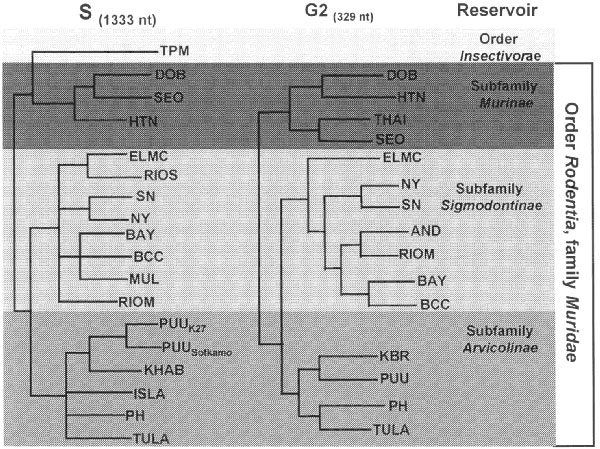
Figure. Phylogeny of hantaviruses and their relationships to natural reservoirs. The trees were constructed by comparing the complete coding regions of the S segments of hantaviruses or of 330 nucleotides corresponding to those of the M segment of Hantaan virus (strain 76118) from nucleotides 1987 to 2315. Abrreviations for viruses are as in Table 1. For each analysis, a single most parsimonious tree was derived by using PAUP 3.1.1 software. For the S segment tree, boostrap values resulting from 100 replications were all greater than 87% except for the branch leading to BCC (78%) and the branch leading to DOB (52%). The next most common placing of DOB was on a branch with HTN.
Page created: December 21, 2010
Page updated: December 21, 2010
Page reviewed: December 21, 2010
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.