Volume 12, Number 7—July 2006
Dispatch
Replicon Typing of Plasmids Encoding Resistance to Newer β-Lactams
Abstract
Polymerase chain reaction–based replicon typing represents a novel method to describe the dissemination and follow the evolution of resistance plasmids. We used this approach to study 26 epidemiologically unrelated Enterobacteriaceae and demonstrate the dominance of incompatibility (Inc) A/C or Inc N-related plasmids carrying some emerging resistance determinants to extended-spectrum cephalosporins and carbapenems.
Understanding the molecular epidemiology of resistance plasmids has been a major issue since scientists became aware of plasmids' role in the spread of antimicrobial drug resistance. However, understanding this epidemiology has been complex because of the diversity and promiscuity of these elements. The plasmid replication system, which dictates the plasmid's behavior (host range, copy number) is the major plasmid landmark from a biologic standpoint; it is used for plasmid classification and identification (1). Plasmids were originally classified in incompatibility (Inc) groups (2). Inc is a manifestation of plasmid relatedness based on commonality of replication controls. The standard procedure for determining Inc groups requires laborious hands-on work, multiple conjugation, transformation assays, or hybridization experiments (1–3).
Our objective of understanding the relationship among resistance plasmids prompted us to develop a polymerase chain reaction (PCR)–based replicon typing method (4). Our study has 2 aims: 1) to investigate phylogenetic relatedness among plasmids carrying extended-spectrum cephalosporin (ESC) and carbapenem resistance determinants emerging in 3 different countries (Greece, Italy, and the United States) and 2) to ascertain the sensitivity of the method.
PCR-based replicon typing was applied to type the resistance plasmids carried by 26 Escherichia coli transconjugants or transformants obtained from epidemiologically unrelated clinical isolates of Enterobacteriaceae associated with community- or hospital-acquired infections in the United States or southern Europe (Italy and Greece). The resistance plasmids carried genes encoding β-lactamases of Ambler class A (SHV-12), B (VIM-1 or VIM-4), and C (CMY-2, CMY-4, or CMY-13) (Table), which represent key emerging resistance determinants to ESC and carbapenems.
Eighteen primer pairs were used to perform 5 multiplex and 3 simplex PCRs, which recognized FIA, FIB, FIC, HI1, HI2, I1-Iγ, L/M, N, P, W, T, A/C, K, B/O, X, Y, and FII replicons (4). All amplified replicons were sequenced by standard procedures and used as specific probes to confirm the replicon typing results by Southern blot hybridization on purified plasmid DNA (data not shown).
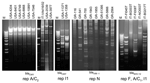
The plasmid donors from the United States consisted of 4 previously characterized ESC-resistant Salmonella isolates submitted to the National Antimicrobial Resistance Monitoring System (NARMS) from 1996 to 1998 (12) and 6 ESC-resistant Escherichia coli O157:H7 isolates collected by NARMS from 2000 to 2001 (5). During the study periods, participating state and local public health laboratories forwarded every tenth non-Typhi type Salmonella and every fifth E. coli O157 isolate they received to the Centers for Disease Control and Prevention for susceptibility testing. This collection includes representatives from sporadic and outbreak infections (5,12). The 6 Salmonella and 4 E. coli plasmid donors selected for this study were a small sample of epidemiologically unrelated isolates representative of those carrying a blaCMY-2 β-lactamase gene on plasmids classified as type A or B on the basis of the blaCMY-2 hybridization pattern (6,13). The PCR-based replicon typing method assigned the A/C and I1 replicons to type A and type B plasmids, respectively (Table), which was confirmed by DNA sequencing. The I1-type amplicon sequences were identical to the R64 IncI1 reference plasmid (no. AP005147), whereas the A/C-type amplicon sequences exhibited 26 nucleotide (nt) substitutions with respect to the RA1 IncA/C reference plasmid (no. X73674), which caused 3 amino acid variations. Therefore, the A/C-replicon from the US plasmids may represent a new replicon variant, which we designated repA/C2 (DNA sequence released under EMBL accession no. AM087198). The Figure shows conserved PstI restriction profiles obtained for the A/C2 plasmids that are different from those exhibited by the I1 plasmids.
The plasmid donors from Italy consisted of 7 multidrug-resistant isolates of various species of Enterobacteriaceae carrying either blaSHV-12 or blaCMY-4 and blaVIM-4 plasmidborne β-lactamase genes (Table). These isolates had been collected from 2002 to 2003 at 4 different hospitals in northern or central Italy (7,8) and were epidemiologically unrelated, except for IT-VA416/02 and IT-VA417/02, which were from the same patient (7). PCR replicon typing of the 5 blaSHV-12–carrying plasmids detected 3 repFII (100% identical to the reference sequence no. M33752), 1 repI1 (100% identical to the R64 plasmid), and 1 repA/C1 (99% homologous to the RA1 plasmid) (Table), suggesting mobilization of this gene among different plasmid scaffolds. The blaSHV-12 plasmids showed different PstI restriction patterns, which confirmed their diversity (Figure). The 2 plasmids carrying blaVIM-4 and blaCMY-4 were assigned by PCR replicon typing to the A/C type. The sequence of these replicons showed the same 26 characteristic nucleotide substitutions of the A/C2-replicon identified in the US plasmids. These 2 A/C2-plasmids showed an apparently identical PstI restriction profile (data not shown), which was also very similar to that of some USA blaCMY-2 plasmids (see the 2039 and 3977 US plasmids and the Italian VA416/02 plasmid in the Figure). The 2 Italian A/C2 plasmids, in addition to blaCMY-4 (which is a blaCMY-2 variant different by only a single nucleotide substitution), also carried the blaVIM-4 carbapenemase gene, which has not been reported on blaCMY-2–carrying plasmids from the United States and may represent a novel acquisition. These findings indicate intercontinental spread of these plasmids and novel acquisition of resistance genes.
The plasmid donors from Greece consisted of a collection of 7 Klebsiella pneumoniae isolates carrying the blaVIM-1 gene (9) and 2 E. coli isolates carrying blaVIM-1 and blaCMY-13 genes (10). These isolates, randomly collected from 5 different hospitals in Athens and Piraeus from 2001 to 2003, are representative of the VIM-1–producing isolates circulating in Greece. No repetitive samples were taken from patients. All isolates exhibited decreased susceptibility to carbapenems. Restriction analysis of these plasmids classified them into 6 different groups on the basis of their restriction profiles (Figure). By replicon typing, all of these plasmids were assigned to the same repN-type replicon, which exhibited 2-nt point mutations (99% homology) in respect to the R46 IncN reference plasmid (no. NC_003292), an indication that they were phylogenetically related and probably evolved from a common ancestor. Although one might expect similar plasmid scaffolds to exist among isolates in Greece and Italy because of geographic proximity, this was not the case. This finding explains the great variability of resistance plasmids carrying different combinations of resistance genes.
Since the origin of replication is a constant and conserved part of a plasmid, replicon typing focused on this portion of the plasmid is a more sensitive and specific method for identifying phylogenetically related plasmids than restriction-based analysis of the entire plasmid. This fact is probably due to the presence of multiple mobile elements (IS elements, transposons, integrons) that can mediate rearrangements of the plasmid scaffolds, which leads to the formation of apparently divergent plasmids. In fact, this phenomenon was demonstrated for the GR-541 plasmid that contains multiple copies of insertion sequences and other mobile genetic elements within its scaffold (14).
A PCR-based replicon typing approach was successfully applied to relevant resistance plasmids. Coupled with sequencing, the approach allowed high-resolution typing of the plasmid replicons. Typing results provided original insights into the molecular epidemiology of resistance plasmids. For instance, the blaCMY-2–carrying plasmid circulating in the United States was also detected in Europe in the form of a derivative that also carries the VIM-4 carbapenemase determinant. This finding demonstrates that plasmids carrying resistance to clinically relevant antimicrobial agents can spread worldwide among bacteria responsible for both nosocomial and community-acquired infections. The heterogeneity among Italian plasmids encoding SHV-12 (the most prevalent SHV-type extended-spectrum β-lactamase in this country) (15) suggests a notable potential for this determinant to spread among different plasmid replicons. On the other hand, replicon typing indicated that the VIM-1–encoding plasmids from Greece were all related despite their different restriction profiles, which points out the common origin of these plasmids. The blaCMY-13 gene from Greece is located on the repN plasmid, whereas Italy and the United States share the A/C2 plasmid as a vehicle of the blaCMY gene, despite their geographic distance. Further research is necessary to determine the influences on plasmid trafficking as well as further similarities and differences. Replicon identification may provide useful clues to the evolution of these resistant plasmids. The ability to trace and screen plasmids by PCR may facilitate further understanding of the horizontal transfer of antimicrobial drug resistance.
Dr Carattoli is a researcher in the Department of Infectious, Parasitic, Immune-mediated Diseases at the Istituto Superiore di Sanità. Her areas of research interest include the genetic basis of multidrug resistance and plasmid characterization of gram-negative bacteria.
Acknowledgment
This work is supported by the MED-NET-VET (FOOD-CT-2004-506122, WP09) and DREPS2 (LSHM-CT-2005-018705) contracts with the European Commission and European Research Network within the Training and Mobility of Researchers program (HPRN-CT-2002-00264).
References
- Datta N, Hughes VM. Plasmids of the same Inc groups in Enterobacteria before and after the medical use of antibiotics. Nature. 1983;306:616–7. DOIPubMedGoogle Scholar
- Couturier M, Bex F, Bergquist PL, Maas WK. Identification and classification of bacterial plasmids. Microbiol Rev. 1988;52:375–95.PubMedGoogle Scholar
- Carattoli A, Bertini A, Villa L, Falbo V, Hopkins KL, Threlfall EJ. Identification of plasmids by PCR-based replicon typing. J Microbiol Methods. 2005;63:219–28. DOIPubMedGoogle Scholar
- Whichard JM, Joyce K, Fey P, Nelson JM, Angulo FJ, Barrett TJ. β-lactam resistance and Enterobacteriaceae, United States. Emerg Infect Dis. 2005;11:1464–6.PubMedGoogle Scholar
- Carattoli A, Tosini F, Giles WP, Rupp ME, Hinrichs SH, Angulo FJ, Characterization of plasmids carrying CMY-2 from expanded-spectrum cephalosporin-resistant Salmonella isolated in the United States between 1996 and 1998. Antimicrob Agents Chemother. 2002;46:1269–72. DOIPubMedGoogle Scholar
- Luzzaro F, Docquier JD, Colinon C, Endimiani A, Lombardi G, Amicosante G, Emergence in Klebsiella pneumoniae and Enterobacter cloacae clinical isolates of the VIM-4 metallo-β-lactamase encoded by a conjugative plasmid. Antimicrob Agents Chemother. 2004;48:648–50. DOIPubMedGoogle Scholar
- Luzzaro F, Mezzatesta M, Mugnaioli C, Perilli M, Stefani S, Amicosante G, Trends in the production of extended-spectrum β-lactamases among enterobacteria of medical interest. Report of the second Italian nationwide survey. J Clin Microbiol. 2006;44:1659–64. DOIPubMedGoogle Scholar
- Giakkoupi P, Xanthaki A, Kanelopoulou M, Vlahaki A, Miriagou V, Kontou S, VIM-1 metallo-β-lactamase-producing Klebsiella pneumoniae strains in Greek Hospitals. J Clin Microbiol. 2003;41:3893–6. DOIPubMedGoogle Scholar
- Miriagou V, Tzelepi E, Gianneli D, Tzouvelekis LS. Escherichia coli with a self-transferable, multi-resistant plasmid coding for the metallo-β-lactamase VIM-1. Antimicrob Agents Chemother. 2003;47:395–7. DOIPubMedGoogle Scholar
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial disk susceptibility tests; approved standard. 9th ed. Wayne (PA): The Institute; 2006.
- Dunne EF, Fey PD, Kludt P, Reporter R, Mostashari F, Shillam P, Emergence of domestically acquired ceftriaxone-resistant Salmonella infections associated with AmpC beta-lactamase. JAMA. 2000;284:3151–6. DOIPubMedGoogle Scholar
- Giles WP, Benson AK, Olson ME, Hutkins RW, Whichard JM, Winokur PL, DNA sequence analysis of regions surrounding blaCMY-2 from multiple Salmonella plasmid backbones. Antimicrob Agents Chemother. 2004;48:2845–52. DOIPubMedGoogle Scholar
- Miriagou V, Carattoli A, Tzelepi E, Villa L, Tzouvelekis LS. IS26-associated In4-type integrons forming multiresistance loci in enterobacterial plasmids. Antimicrob Agents Chemother. 2005;49:3541–3. DOIPubMedGoogle Scholar
- Perilli M, Dell'Amico E, Segatore B, De Massis MR, Bianchi C, Luzzaro F, Molecular characterization of extended-spectrum β-lactamases produced by nosocomial isolates of Enterobacteriaceae from an Italian nationwide survey. J Clin Microbiol. 2002;40:611–4. DOIPubMedGoogle Scholar
Figure
Table
Cite This ArticleTable of Contents – Volume 12, Number 7—July 2006
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Alessandra Carattoli, Department of Infectious, Parasitic, Immune-mediated Diseases, Istituto Superiore di Sanità, Rome, Viale Regina Elena 299, 00161 Rome, Italy
Top