Volume 17, Number 2—February 2011
Dispatch
New Avian Influenza Virus (H5N1) in Wild Birds, Qinghai, China
Cite This Article
Citation for Media
Abstract
Highly pathogenic avian influenza virus (H5N1) (QH09) was isolated from dead wild birds (3 species) in Qinghai, China, during May–June 2009. Phylogenetic and antigenic analyses showed that QH09 was clearly distinguishable from classical clade 2.2 viruses and belonged to clade 2.3.2.
In May 2005, highly pathogenic avian influenza (HPAI) virus (H5N1) caused a disease outbreak in wild birds in the Qinghai Lake region of the People’s Republic of China (1). Subsequently, this virus (QH05, clade 2.2) disseminated from Asia to Europe and Africa, which has led to great concern and energetic debates about the role of migratory birds in influenza epidemics (1–5). In 2006, this virus was detected in migratory birds in Qinghai (6,7). In 2007, viruses similar to QH05 were isolated from surveyed anseriformes in Qinghai and showed only a short evolutionary distance from earlier viruses (8). Genetic diversity of avian influenza viruses (H5N1) was not detected in wild birds in Qinghai before 2008 (7,8). We report evidence that a second lineage of viruses, in addition to clade 2.2, has emerged in wild birds in Qinghai.
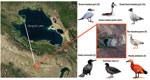
During May 8–June 15, 2009, a total of 273 wild birds died in the wetlands of Gengahai Lake, Qinghai Province, China (Figure 1) during an influenza outbreak. We obtained 224 great crested grebes (Podiceps cristatus), 16 great black-headed gulls (Larus ichthyaetus), 15 brown-headed gulls (Larus brunnicephalus), 9 bar-headed geese (Anser indicus), 5 ruddy shelducks (Tadorna ferruginea), 3 great cormorants (Phalacrocorax carbo), and 1 common coot (family Rallidae) (Figure 1).
To determine the pathogenesis of this outbreak, we obtained organs, including lung and brain, and cloacal swabs from 13 birds at different times. Tissue samples were inoculated into 10-day-old, embryonated, specific pathogen–free eggs for virus isolation. Hemagglutinin and neuraminidase subtypes were determined as described (9). Eleven avian influenza viruses (H5N1) were isolated from 3 species of wild birds: 4 from great crested grebes, 5 from great black-headed gulls, and 2 from brown-headed gulls. Results of virus isolation for samples from a bar-headed goose and a shelduck were negative. Samples from great cormorants and a common coot were not obtained and tested.
We sequenced genomes of the 11 viruses and found that the viruses (QH09) were closely related and showed 100% homology at the nucleotide level. Sequences of the entire genome of QH09 (representative strain A/great crested grebe/Qinghai/1/09) are available in GenBank (accession nos. CY063315–CY063322).
Hemagglutinin of QH09 virus had a series of basic amino acids (PQRERRRKR) at the cleavage site. Neuraminidase of this virus had a deletion of 20 aa at residues 49–68 in the stalk region. No amino acid substitutions conferred resistance to adamantane or neuraminidase inhibitors. Unlike many isolates related to Qinghai Lake strains of clade 2.2, QH09 virus does not have an E627K substitution in basic polymerase. Nonstructural protein 1 had a deletion of 5 aa at residues 80–84, which is commonly observed in HPAI viruses (H5N1) that are circulating in Southeast Asia.
Seven gene segments of QH09, except for the acidic polymerase (PA) gene, showed the greatest homology with 2 clade 2.3.2 viruses, A/little egret/Hongkong/8863/07(H5N1) and A/whooper swan/Hokkaido/1/08(H5N1); identities were 97.0%–99.5%. The strain that showed the greatest homology with QH09 PA was A/chicken/Yamaguchi/7/04 (H5N1), which was the precursor of the PA gene of the A/bar-headed goose/Qinghai/2/05 virus (1) (Table). Homology of the 8 gene segments of QH09 with QH05-like viruses was 92%–96.3%.
Phylogenetic analysis showed that 7 of 8 gene fragments of QH09, except for the PA gene, mapped with clade 2.3.2 viruses A/little egret/Hongkong/8863/07 and A/whooper swan/Hokkaido/1/08 and were in a different cluster than clade 2.2 viruses isolated in Qinghai during 2005–2007 (Figure 2, panel A). The PA gene of QH09 had the same lineage as A/chicken/Yamaguchi/7/04 (Figure 2, panel B).
Antigenic analyses by hemagglutination inhibition (HI) assay with chicken antiserum against a QH05 virus (A/bar-headed goose/Qinghai/3/05) (clade 2.2) and QH09 virus showed that the cross-reactive HI titer of QH09 virus (64) was 16-fold lower than that of homologous QH05 virus (1,024). The cross-reactive HI titer of QH05 virus (128) was also 16-fold lower than that of homologous QH09 virus (2,048).
Genetic and antigenic characterization of HPAI viruses (H5N1) from wild birds in Qinghai in 2009 suggests that these viruses are closely related to clade 2.3.2 and are clearly distinguishable from the classical QH05 clade 2.2. Previously reported QH05-like virus represented >4 genotypes, and no 2 QH05-like viruses were identical at the amino acid or nucleotide sequence levels (1,6). Although isolates of QH09 were obtained from 3 species at different times, 100% homology of the 11 isolates of QH09 was observed, which suggests that wild birds in Qinghai in 2009 were newly infected by 1 strain of virus.
Clade 2.3.2 viruses from wild nonpasserine bird species were reported in Hong Kong in 2007–2008 (10). The fact that a similar virus was isolated from a whooper swan (order Anseriformes) in Japan in 2008 showed that clade 2.3.2 was dispersed by migration of wild birds (11). Our results indicated that QH09 virus is a reassortant containing 7 gene segments of clade 2.3.2 viruses detected in wild birds and the PA gene of CK/Yamaguchi/7/04-like virus, which contributed the PA gene to 1 QH05 virus (1).
Similar genotypes of QH09-like clade 2.3.2 viruses were also detected in great-crested grebes and black-headed gulls in Russia in 2009 (12). Bar-headed geese, whooper swans, and other anseriforme birds in Mongolia were infected with QH09-like clade 2.3.2 viruses (13). Therefore, QH09-like clade 2.3.2 virus is likely adapted to wild birds and is similar to clade 2.2 viruses, and its presence in Qinghai suggests that wild birds have spread this virus to other regions.
The possibility that wild birds in Qinghai in 2009 were infected by domestic fowl that harbored clade 2.3.2 virus is low because of the location of Qinghai Province and the scarcity of poultry in this region. In addition, no influenza outbreak in poultry occurred in this region in 2009. Therefore, it is likely that wild birds spread the virus.
How wild birds transmit HPAI virus (H5N1) is not clear. The ecology, epidemiology, genetics, and evolution of this virus are not fully understood. QH05-like clade 2.2 viruses and clade 2.3.2 viruses have been detected in wild pikas (14). However, whether the 2 virus clades have propagated in other mammalian hosts is unknown.
Qinghai Lake is located near multiple avian flyways. Although there are no reports of detection of clade 2.3.2 virus in wild birds near Qinghai Lake, the finding of clade 2.3.2 virus in the Gengahai wetlands of Qinghai increases concerns about a potential pandemic and the likelihood that avian influenza virus (H5N1) will again spread and increase its genetic diversity. Therefore, determining movements of wild migratory waterfowl from Qinghai Lake and their virologic status is needed to assess potential avian vectors of HPAI virus (H5N1).
Dr Li is a veterinary microbiologist at the National Avian Influenza Reference Laboratory, Harbin, People’s Republic of China. Her research interests are the pathogenesis and molecular epidemiology of avian influenza viruses.
Acknowledgment
This study was supported by the 973 Program (2011CB505001, 2010CB534000); the Natural Science Foundation of China (30825032); the Animal Infectious Disease Control Program of the Ministry of Agriculture of China; the National Non-profit Special Fund for Research on Animal Diseases of Borders (200803026); grants from the Chinese National Science and Technology Plan (2006BAD06A05 and 2006BAD06A03); and the Disease Surveillance Program of the Ministry of Health of China (2009ZX10004-214).
References
- Chen H, Li Y, Li Z, Shi J, Shinya K, Deng G, Properties and dissemination of H5N1 viruses isolated during an influenza outbreak in migratory waterfowl in western China. J Virol. 2006;80:5976–83. DOIPubMedGoogle Scholar
- Liu J, Xiao H, Lei F, Zhu Q, Qin K, Zhang XW, Highly pathogenic H5N1 influenza virus infection in migratory birds. Science. 2005;309:1206. DOIPubMedGoogle Scholar
- Newman SH, Iverson SA, Takekawa JY, Gilbert M, Prosser DJ, Batbayar N, Migration of whooper swans and outbreaks of highly pathogenic avian influenza H5N1 virus in eastern Asia. PLoS ONE. 2009;4:e5729. DOIPubMedGoogle Scholar
- Olsen B, Munster VJ, Wallensten A, Waldenstrom J, Osterhaus AD, Fouchier RA. Global patterns of influenza a virus in wild birds. Science. 2006;312:384–8. DOIPubMedGoogle Scholar
- Salzberg SL, Kingsford C, Cattoli G, Spiro DJ, Janies DA, Aly MM, Genome analysis linking recent European and African influenza (H5N1) viruses. Emerg Infect Dis. 2007;13:713–8.PubMedGoogle Scholar
- Li Y, Shi J, Zhong G, Deng G, Tian G, Ge J, Continued evolution of H5N1 influenza viruses in wild birds, domestic poultry and humans in China from 2004 to 2009. J Virol. 2010;84:8389–97. DOIPubMedGoogle Scholar
- Wang G, Zhan D, Li L, Lei F, Liu B, Liu D, H5N1 avian influenza re-emergence of Lake Qinghai: phylogenetic and antigenic analyses of the newly isolated viruses and roles of migratory birds in virus circulation. J Gen Virol. 2008;89:697–702. DOIPubMedGoogle Scholar
- Kou Z, Li Y, Yin Z, Guo S, Wang M, Gao X, The survey of H5N1 flu virus in wild birds in 14 pProvinces of China from 2004 to 2007. PLoS ONE. 2009;4:e6926. DOIPubMedGoogle Scholar
- World Health Organization. WHO manual on animal influenza diagnosis and surveillance. 2002 [cited 2010 Sep 7]. http://www.who.int/csr/resources/publications/influenza/en/whocdscsrncs20025rev.pdf.
- Smith GJ, Vijaykrishna D, Ellis TM, Dyrting KC, Leung YH, Bahl J, Characterization of avian influenza viruses A (H5N1) from wild birds, Hong Kong, 2004–2008. Emerg Infect Dis. 2009;15:402–7. DOIPubMedGoogle Scholar
- Uchida Y, Mase M, Yoneda K, Kimura A, Obara T, Kumagai S, Highly pathogenic avian influenza virus (H5N1) isolated from whooper swans, Japan. Emerg Infect Dis. 2008;14:1427–9. DOIPubMedGoogle Scholar
- Sharshov K, Silko N, Sousloparov I, Zaykovskaya A, Shestopalov A, Drozdov I. Avian influenza (H5N1) outbreak among wild birds, Russia, 2009. Emerg Infect Dis. 2010;16:349–51.PubMedGoogle Scholar
- Kang H, Batchuluun D, Kim M, Choi J, Erdene-Ochir T, Paek M, Genetic analyses of H5N1 avian influenza virus in Mongolia, 2009 and its relationship with those of eastern Asia. Vet Microbiol. 2010 Jun 9; [Epub ahead of print].
- Zhou J, Sun W, Wang J, Guo J, Yin W, Wu N, Characterization of the H5N1 highly pathogenic avian influenza virus derived from wild pikas in China. J Virol. 2009;83:8957–64. DOIPubMedGoogle Scholar
Figures
Table
Cite This Article1These authors contributed equally to this article.
Table of Contents – Volume 17, Number 2—February 2011
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Hualan Chen, Harbin Veterinary Research Institute, Chinese Academy of Agricultural Sciences, 427 Maduan St, Harbin 150001, People’s Republic of China
Top