Volume 16, Number 9—September 2010
Research
Worldwide Diversity of Klebsiella pneumoniae That Produce β-Lactamase blaKPC-2 Gene1
Abstract
Klebsiella pneumoniae isolates that produce carbapenemases (KPCs) are rapidly disseminating worldwide. To determine their genetic background, we investigated 16 blaKPC-2-harboring K. pneumoniae isolates from 5 countries. The isolates were multidrug resistant, possessed the blaKPC-2 gene, and differed by additional β-lactamase content. They harbored a naturally chromosome-encoded bla gene (blaSHV-1 [12.5%], blaSHV-11 [68.7%], or blaOKP-A/B [18.8%]) and several acquired and plasmid-encoded genes (blaTEM-1 [81.3%], blaCTX-M-2 [31.3%], blaCTX-M-12 [12.5%], blaCTX-M-15 [18.7%], and blaOXA-9 [37.5%]). The blaKPC-2 gene was always associated with 1 of the Tn4401 isoforms (a, b, or c). Tn4401 was inserted on different-sized plasmids that belonged to different incompatibility groups. Several blaKPC-containing K. pneumoniae clones were found: 9 different pulsotypes with 1 major (sequence type 258) and 7 minor distinct allelic profiles. Different clones harboring different plasmids but having identical genetic structure, Tn4401, could be at the origin of the worldwide spread of this emerging resistance gene.
Resistance of Klebsiella pneumoniae to carbapenems is mainly associated with acquired carbapenem-hydrolyzing β-lactamases (1). These β-lactamases can be metallo β-lactamases (IMP, VIM), expanded-spectrum oxacillinases (OXA-48), or Ambler class A enzymes (NMCA, IMI, SME, GES, and KPC) (1–4). The most common class A carbapenemases in K. pneumoniae are the K. pneumoniae carbapenemases (KPCs) (4). KPCs in carbapenem-resistant K. pneumoniae strains were first reported in 2001 in North Carolina (5), and until 2005, the geographic distribution of these enzymes in Enterobacteriaceae, including K. pneumoniae, was limited to the eastern part of the United States (5,6). Now, KPC-producing K. pneumoniae isolates are frequently identified among nosocomial pathogens (7). Recently, dissemination of a single major clone of KPC-producing K. pneumoniae (sequence type [ST] 258) in the eastern United States has been suggested (8). KPCs have been observed more rarely among other gram-negative rods such as Pseudomonas spp (9,10).
Outside the United States, KPC-producing K. pneumoniae are also being reported more often. The first case of KPC-producing K. pneumoniae infection was reported in 2005 in France and had a US origin (11). The first outbreak of KPC-producing K. pneumoniae outside the United States was in Israel (12). In South America, dissemination of KPC-producing K. pneumoniae was initially reported in 2006 in Colombia (13) and then in Brazil and Argentina (14,15). KPC enzymes have also been identified in P. aeruginosa isolates from Colombia (16). In the People’s Republic of China, KPC enzymes in several enterobacterial species are being increasingly reported (17). Finally, in Europe a few cases of KPC-producing K. pneumoniae infection have been described, but in Greece, outbreaks have occurred (18). In Europe, different variants of KPCs (KPC-2 and KPC-3) have been described; some patients carrying KPC-positive isolates had been transferred from the United States, Israel, or Greece (19–21).
Reports of this β-lactamase being found in novel locations are increasing worldwide, probably signaling active spread. The genetic element carrying the blaKPC-2 gene, Tn4401, was recently elucidated (22). Three isoforms of this Tn3-like transposon (a, b, and c) are known. Several other genetic environments of blaKPC gene have been described; other insertion sequences have been found upstream of the blaKPC gene (23,24). Nevertheless, the downstream sequences of the blaKPC gene matched perfectly with Tn4401, which suggests that these insertion sequences have been inserted into Tn4401.
Insertion sequences may play major roles in the evolution of Tn4401, but little information is available about the bacterial strains and the plasmids that may explain this rapid spread. Our goal, therefore, was to characterize the genetic background of several blaKPC-2- harboring K. pneumoniae isolates from various geographic origins.
Bacterial Strains
K. pneumoniae isolates used in this study and their origin are listed in Table 1 (11,13,16,21,25). Electrocompetent Escherichia coli DH10B (Invitrogen, Eragny, France) was used as a recipient in electroporation experiments. E. coli J53AzR, which is resistant to sodium azide, was used for conjugation experiments. E. coli 50192 was used as a reference strain for plasmid extraction (22).
Antibiograms and MIC Determinations
Antibiograms were created by using the disk-diffusion method on Mueller-Hinton agar (Bio-Rad Laboratories, Marnes-La-Coquette, France), and susceptibility break points were determined as previously described and interpreted as recommended by the Clinical and Laboratory Standards Institute (22,26). All plates were incubated at 37°C for 18 h. MICs of β-lactams were determined by using the Etest technique (bioMérieux, Marcy l’Etoile, France).
Electroporation and Plasmid Extraction
Direct transfer of resistance into azide-resistant E. coli J53 was attempted as reported (22). Plasmids were introduced by electroporation into E. coli DH10B (22) by using a Gene Pulser II (Bio-Rad Laboratories).
Plasmid DNA was extracted by using a QIAGEN Plasmid Maxi Kit (QIAGEN, Courtaboeuf, France) and analyzed by agarose gel electrophoresis (Invitrogen, Paris, France). Natural plasmids were extracted by using the Kieser extraction method (27) and subsequently analyzed by electrophoresis on a 0.7% agarose gel.
Hybridization
DNA–DNA hybridization was performed as described by Sambrook et al. (28) with Southern transfer of an agarose gel containing Kieser method–extracted total DNA. The probe consisted of a 796-bp PCR-generated fragment from recombinant plasmid pRYC-1 (22) and was internal to the blaKPC-2 gene. Labeling of the probe and detection of signal were conducted by using an ECL nonradioactive labeling and detection kit according to the manufacturer's instructions (Amersham Biosciences, Orsay, France).
PCR Amplification and Sequencing
The blaCTX-M-, blaSHV-, blaTEM-, and blaOXA-1/9-like genes were searched for and characterized as described (21). PCR experiments were performed on an ABI 2700 thermocycler (Applied Biosystems, Les Ulis, France) by using laboratory-designed primers (Table 2). PCR products were then analyzed on agarose gel and sequenced.
Both strands of the PCR products were sequenced by using laboratory-designed primers with an automated sequencer (ABI PRISM 3100; Applied Biosystems). The nucleotide and the deduced protein sequences were analyzed by using software from the National Center for Biotechnology Information (www.ncbi.nlm.nih.gov).
Isoelectrofocusing
Crude β-lactamase extracts, obtained as described (21) from 10-mL cultures of clinical isolates and their E. coli transconjugants or electroporants were subjected to analytical isoelectrofocusing on an ampholine-containing polyacrylamide gel, pH 3.5–9.5 (Ampholine PAG plate; GE Healthcare, Orsay, France) for 90 min at 1,500 volts, 50 milliamps, and 30 watts. The focused β-lactamases were detected by overlaying the gel with 1 mmol nitrocefin (Oxoid, Dardilly, France). Isoelectric points were determined and compared with those of known β-lactamases (22).
Pulsed-field Gel Electrophoresis
Pulsed-field gel electrophoresis (PFGE) was performed by using XbaI (GE Healthcare) as described (29). XbaI-macrorestriction patterns were interpreted according to the recommendations of Tenover et al. (30).
Multilocus Sequence Typing
Multilocus sequence typing (MLST) with 7 housekeeping genes (rpoB, gapA, mdh, pgi, phoE, infB, and tonB) was performed according to Diancourt et al. (31). Allele sequences and STs were verified at http://pubmlst.org/kpneumoniae. A different allele number was given to each distinct sequence within a locus, and a distinct ST number was attributed to each distinct combination of alleles.
Pulsotypes
Molecular typing by PFGE identified 9 major pulsotypes among the isolates (Table 3). The first pulsotype (pulsotype A) corresponded to the strains from the United States and Greece. We found 4 different pulsotypes (B–E) among strains from Colombia, which suggested polyclonal diffusion inside this country. We also identified 2 different clones among strains from Brazil (pulsotypes F and G) and from Israel (pulsotypes H and I). These results indicate much heterogeneity among KPC-producing isolates from various geographic regions.
MLST of the 16 isolates resulted in 8 distinct allelic profiles: ST 258 (allelic profile 3–3-1–1-1–1-79) corresponding to isolates K. pneumoniae YC (United States), K. pneumoniae GR (Greece), and K. pneumoniae K271 (Greece); ST 14 (allelic profile 1–6-1–1-1–1-1) corresponding to isolates K. pneumoniae INC H1521–6 and K. pneumoniae INC H1516–6 (Colombia); ST 11 (allelic profile 3–3-1–1-1–1-4) corresponding to isolates from Brazil; ST 277 (allelic profile 3–1-1–1-1–1-43) corresponding to isolate K. pneumoniae 475 (Israel); novel ST 337 (allelic profile 2–1-11–1-1–1-13) corresponding to isolate K. pneumoniae KN2303 (Colombia); ST 338 (allelic profile 17–19–22–39–34–21–52) corresponding to isolates K. pneumoniae KN633 and K. pneumoniae HPTU 2020532 (Colombia); ST 339 (allelic profile 18–15–25–24–11–13–51) corresponding to isolate K. pneumoniae HPTU 27635 (Colombia); and ST 340 (allelic profile 3–3-1–1-1–1-18) corresponding to isolate K. pneumoniae 588 (Israel). The analysis of STs by eBURST (http://pubmlst.org) showed that ST 11 and ST 340 are single-locus variants of ST 258 and that ST 277 is a double-locus variant of ST 258. These results matched perfectly with PFGE results. One isolate from Brazil (K. pneumoniae A33504) showed a different pattern by PFGE but the same ST (ST 11) as other isolates from the same origin, which suggests a strong genetic relatedness.
Antimicrobial Drug Susceptibility
All isolates were resistant to penicillins and cephalosporins but showed varying levels of susceptibility to carbapenems (Table 4). Resistance to other drug classes varied among the isolates. For aminoglycosides, 2 clones (A and I) were susceptible to gentamicin only, 1 clone (H) was susceptible to amikacin only, and 3 clones (C, D, and G) were resistant to all tested aminoglycosides. Six clones (A, C, D, F, G, and I) showed resistance to fluoroquinolones. Percentages of nonsusceptible isolates to the non–β-lactam drugs were as follows: gentamicin, 75%; amikacin, 81.3%; ciprofloxacin, 81.5%; trimethoprim/sulfamethoxazole, 81.5%; and tetracycline, 87.5%. Two isolates were also resistant to colistin (K. pneumoniae GR and K. pneumoniae K271); each was from Greece, where this drug is often used (33).
β-Lactamase Genes
Positive results of CTX-M–, TEM-, SHV-, and OXA-specific PCRs are indicated in Table 1. All isolates possessed the blaKPC-2 gene and a naturally chromosome-encoded bla gene: blaSHV-1 (12.5%), blaSHV-11 (68.7%), or blaOKP-A/B (18.8%). The blaOKP genes are 1 of the 3 families of the chromosomal β-lactamase genes found in K. pneumoniae (34) with blaSHV and blaLEN and share 88% similarity with blaSHV-1. K. pneumoniae isolates also harbored several acquired and plasmid-encoded genes: blaTEM-1 (81.3%), blaCTX-M-2 (31.3%), blaCTX-M-12 (12.5%), blaCTX-M-15 (18.7%), and blaOXA-9 (37.5%).
Characterization Results for Tn4401
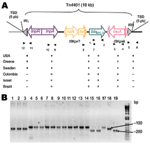
Primer couples specific for the different genes found on Tn4401 (Table 2; Figure 1, panel A) obtained similar-sized fragments for all strains, which suggests that the strains have a similar genetic organization. For only 1 primer pair, hybridizing in ISKpn7 and blaKPC gene (primers 7 and 8 in Figure 1, panel A), located upstream of the blaKPC gene, an ≈100-bp (K. pneumoniae YC, K. pneumoniae GR, and K. pneumoniae 271) or 200-bp (K. pneumoniae 475 and K. pneumoniae 588) shorter fragment was observed, compared with the Tn4401b structure, thus indicating that the 3 isoforms of Tn4401 were present in this collection of isolates (Figure 1, panel B).
To investigate the flanking sequences of Tn4401, we used PCR primers located in the Tn4401 structure and in the flanking sequences derived from K. pneumoniae YC (22). PCR products of expected size were obtained for K. pneumoniae GR and K. pneumoniae K271 isolates only. For all other strains, no PCR product could be obtained, suggesting that the Tn4401 insertion site might differ from that found in K. pneumoniae YC.
Genetic Support for blaKPC in the Isolates
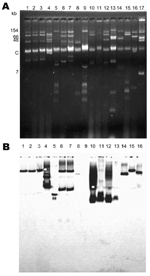
The carbapenem-resistant K. pneumoniae isolates contained several plasmids of different sizes, ranging from <5 kb to >170 kb (Figure 2, left panel). At least 1 plasmid hybridized with an internal probe for blaKPC-2 gene in each isolate, ranging from 13 kb to 80 kb (Figure 2, right panel; Table 3). We observed 2 hybridization signals (35 kb and 75 kb) for K. pneumoniae KN2303, as described (22). Plasmid location of the blaKPC genes was confirmed by electroporation of these plasmids into E. coli DH10B, but no transformant could be obtained for K. pneumoniae 2020532. The E. coli transformants had a β-lactam resistance pattern that corresponded to the expression of a blaKPC-like gene. Electroporation of 4 plasmids harboring the blaKPC-like gene into E. coli DH10B conferred resistance to at least an aminoglycoside molecule; pINC-H1521–6, pA33504, and p588 conferred resistance to all aminoglycosides except gentamicin, and electroporation of p475 into E. coli DH10B led to resistance to all aminoglycosides tested. No other antimicrobial drug resistance marker was cotransferred; the transformants remained susceptible to nalidixic acid, levofloxacin, ciprofloxacin, rifampin, tetracycline, trimethoprim/sulfamethoxazole, and colistin.
Mating-out assays showed that the ≈75–80-kb plasmids harboring blaKPC-2 from K. pneumoniae YC, GR, K271, and KN2303 were self-transferable to E. coli. The smaller plasmid from K. pneumoniae KN633 was not transferred to E. coli.
Origin of Replication
PCR-based replicon typing of the major plasmid incompatibility groups showed that the blaKPC-2-positive plasmids belonged to at least 3 incompatibility groups (IncFIIAS, IncN, and IncL/M) (Table 3). The plasmids of K. pneumoniae KN633, HPTU-2020532 from Colombia and K. pneumoniae A33504 from Brazil gave negative results with the Inc primers tested and could not be classified into a major plasmid incompatibility group.
Rapid spread of KPC-producing K. pneumoniae is a major clinical and public health concern. These broad-spectrum β-lactamases are increasing in new locations worldwide, indicating an ongoing process. Recently, a novel Tn3-based transposon, Tn4401, was identified in nonclonally related KPC-producing K. pneumoniae and P. aeruginosa isolates (22). This transposon is in most recently described isolates (20,35,36), although a recently characterized novel variant from China had another insertion sequence inserted upstream of blaKPC gene (24). Identification of Tn4401 inserted at different loci, on different plasmids, and flanked by different 5-bp target site duplications indicates a frequent and dynamic process of transposition. It has been suggested that this novel transposon is at the origin of blaKPC-like gene acquisition and dissemination (22). Sixteen K. pneumoniae isolates that express the blaKPC gene from 5 countries were characterized here.
PFGE and MLST showed that several clones are currently spreading in different geographic locations. In Colombia, 3 pulsotypes could be identified. Overall, among the 16 isolates, 1 major ST (258) and its derivative ST 11 seemed to predominate. In a recent study that gathered isolates from 10 US states, ST 258 accounted for 70% of isolates, according to a database of KPC-producing K. pneumoniae PFGE results maintained by the Centers for Disease Control and Prevention (8). This ST has also been identified for KPC-producing K. pneumoniae in Sweden (in isolates imported from Greece and Israel) and more recently in Poland (36,37). These findings suggest possible international dissemination of KPC-producing ST 258. Apparently, the K. pneumoniae clone that contains the extended-spectrum β-lactamase (ESBL) determinant CTX-M-15 belongs to ST 11 (38).
KPC-producing K. pneumoniae contained diverse β-lactamases. All except 2 isolates harbored at least another β-lactamase; blaTEM-1 and a blaCTX-M-type ESBLs were expressed by >80% and 62.5% of isolates, respectively. KPC producers have already been associated with other β-lactamase genes, such as the widespread ESBL gene blaCTX-M-15 (17). SHV ESBLs have been found among isolates, as has been described for strains from the United States (39) and Norway (36). These additional β-lactamases are likely to complicate phenotype-based identification of KPC producers. Three isolates harbored the chromosome-encoded blaOKP-A/B genes and belonged to phylogenetic group KpII, which accounts for <10% of K. pneumoniae strains (34). Coexpression of OKP enzymes and ESBLs has rarely been reported.
Isolates also demonstrated diversity in their molecular features. In this study, the KPC-2 genes were encoded on a broad variety of plasmids, as shown by previous studies (22,35). These plasmids differed in size and incompatibility groups. Similar plasmids were observed among isolates with the same ST, whereas different plasmids were also associated with similar STs. Therefore, epidemiologic investigation of KPC producers should be performed at different molecular levels.
Tn4401 was present in all tested strains. The overall structure of Tn4401 seemed to be conserved, except for the 100-bp to 200-bp deletion. Of the 16 isolates, 11 encoded the full-length Tn4401b isoform, 3 encoded the Tn4401a isoform containing a 100-bp deletion (ST 258), and 2 encoded the Tn4401c isoform containing a 200-bp deletion upstream of the blaKPC gene. These types of transposons tend to evolve by capturing various insertion sequences, as illustrated for the vanA-containing Tn1546 transposon (40). For Tn4401, three descriptions have been published in which different insertion sequences were present upstream of blaKPC-2 (22–24). None of these atypical structures were found in our strains. Observation of Tn4401 on different plasmids further supports the hypothesis that this transposon contributes to the mobilization and dissemination of the blaKPC genes.
Our analysis of several K. pneumoniae isolates from 5 geographic origins indicates the spread of different clones that were harboring different plasmids but with an identical genetic structure, Tn4401, that sustained a blaKPC gene acquisition, which could likely be at the origin of the worldwide spread of this emerging resistance gene. Finally, taken together, our findings and those of recent studies report a major KPC-producing clone with ST 258, even if novel ST types could also be evidenced, especially from Colombia. Our data suggest that KPC genes benefit all molecular ingredients (transposon location, self-transferable plasmids, efficient STs) by facilitating their rapid spread to K. pneumoniae and other bacterial species.
Dr Cuzon is an assistant professor of microbiology at the University of Paris South Medical School. Her primary research interest is the genetic basis of antimicrobial drug resistance gene acquisition.
Acknowledgments
We thank Platform Genotyping of Pathogens and Public Health (Institut Pasteur) for coding MLST alleles and profiles.
This work was funded by the Institut National de la Santé et de la Recherche Médicale, France; by a grant from the Ministère de l’Education Nationale et de la Recherche (UPRES-EA3539), Université Paris XI, Paris; by the Assistance Publique-Hôpitaux de Paris, France; and by the European Community (7th PCRD, TROCAR contract HEALTH-F3-2008-223031).
References
- Nordmann P, Poirel L. Emerging carbapenemases in gram-negatives aerobes. Clin Microbiol Infect. 2002;8:321–31. DOIPubMedGoogle Scholar
- Queenan AM, Bush K. Carbapenemases: the versatile beta-lactamases. Clin Microbiol Rev. 2007;20:440–58. DOIPubMedGoogle Scholar
- Poirel L, Héritier C, Tolun V, Nordmann P. Emergence of oxacillinase-mediated resistance to imipenem in Klebsiella pneumoniae. Antimicrob Agents Chemother. 2004;48:15–22. DOIPubMedGoogle Scholar
- Nordmann P, Cuzon G, Naas T. The real threat of KPC carbapenemase–producing bacteria. Lancet Infect Dis. 2009;9:228–36. DOIPubMedGoogle Scholar
- Yigit H, Queenan AM, Anderson GJ, Domenech-Sanchez A, Biddle JW, Steward CD, Novel carbapenem-hydrolyzing β-lactamase KPC-1 from a carbapenem-resistant strain of Klebsiella pneumoniae. Antimicrob Agents Chemother. 2001;45:1151–61. DOIPubMedGoogle Scholar
- Bradford PA, Bratu S, Urban C, Visalli M, Mariano N, Landman D, Emergence of carbapenem-resistant Klebsiella species possessing the class A carbapenem-hydrolyzing KPC-2 and inhibitor-resistant TEM-30 β-lactamases in New York City. Clin Infect Dis. 2004;39:55–60. DOIPubMedGoogle Scholar
- Landman D, Bratu S, Kochar S, Panwar M, Trehan M, Doymaz M, Evolution of antimicrobial resistance among Pseudomonas aeruginosa, Acinetobacter baumannii and Klebsiella pneumoniae in Brooklyn, NY. J Antimicrob Chemother. 2007;60:78–82. DOIPubMedGoogle Scholar
- Kitchel B, Rasheed JK, Patel JB, Srinivasan A, Navon-Venezia S, Carmeli Y, Molecular epidemiology of KPC-producing Klebsiella pneumoniae in the United States: clonal expansion of MLST sequence type 258. Antimicrob Agents Chemother. 2009;53:3365–70. DOIPubMedGoogle Scholar
- Miriagou V, Tzouvelekis LS, Rossiter S, Tzelepi E, Angulo FJ, Whichard J. Imipenem resistance in a Salmonella clinical strain due to plasmid-mediated class A carbapenemase KPC-2. Antimicrob Agents Chemother. 2003;47:1297–300. DOIPubMedGoogle Scholar
- Wolter DJ, Khalaf N, Robledo IE, Vazquez GJ, Sante MI, Aquino EE, Surveillance of carbapenem-resistant Pseudomonas aeruginosa from Puerto Rico medical center hospitals: dissemination of KPC and IMP-18 beta-lactamases. Antimicrob Agents Chemother. 2009;53:1660–4. DOIPubMedGoogle Scholar
- Naas T, Nordmann P, Vedel G, Poyart C. Plasmid-mediated carbapenem-hydrolyzing β-lactamase KPC in a Klebsiella pneumoniae isolate from France. Antimicrob Agents Chemother. 2005;49:4423–4. DOIPubMedGoogle Scholar
- Leavitt A, Navon-Venezia S, Chmelnitsky I, Schwaber MJ, Carmeli Y. Emergence of KPC-2 and KPC-3 in carbapenem-resistant Klebsiella pneumoniae strains in an Israeli hospital. Antimicrob Agents Chemother. 2007;51:3026–9. DOIPubMedGoogle Scholar
- Villegas MV, Lolans K, Correa A, Suarez CJ, Lopez JA, Vallejo M; Colombian Nosocomial Resistance Study Group. First detection of the plasmid-mediated class A carbapenemase KPC-2 in clinical isolates of Klebsiella pneumoniae from South America. Antimicrob Agents Chemother. 2006;50:2880–2. DOIPubMedGoogle Scholar
- Monteiro J, Fernandes Santos A, Asensi MD, Peirano G, Gales AC. First report of KPC-2–producing Klebsiella pneumoniae strains in Brazil. Antimicrob Agents Chemother. 2009;53:333–4. DOIPubMedGoogle Scholar
- Pasteran FG, Otaegui L, Guerriero L, Radice G, Maggiora R, Rapoport M, Klebsiella pneumoniae carbapenemase-2, Buenos Aires, Argentina. Emerg Infect Dis. 2008;14:1178–80. DOIPubMedGoogle Scholar
- Villegas MV, Lolans K, Correa A, Kattan JN, Lopez JA, Quinn JP; Colombian Nosocomial Resistance Study Group. First identification of Pseudomonas aeruginosa isolates producing a KPC-type carbapenem-hydrolyzing β-lactamase. Antimicrob Agents Chemother. 2007;51:1553–5. DOIPubMedGoogle Scholar
- Cai JC, Zhou HW, Zhang R, Chen GX. Emergence of Serratia marscescens, Klebsiella pneumoniae, and Escherichia coli possessing the plasmid-mediated carbapenem-hydrolyzing β-lactamase KPC-2 in intensive care units from a Chinese hospital. Antimicrob Agents Chemother. 2008;52:2014–8. DOIPubMedGoogle Scholar
- Pournaras S, Protonotariou E, Voulgari E, Kristo I, Dimitroulia E, Vitti D, Clonal spread of KPC-2 carbapenemase-producing Klebsiella pneumoniae strains in Greece. J Antimicrob Chemother. 2009;64:348–52. DOIPubMedGoogle Scholar
- Navon-Venezia S, Leavitt A, Schwaber MJ, Rasheed JK, Srinivasan A, Patel JB, First report on hyper-epidemic clone of KPC-3 producing Klebsiella pneumoniae in Israel genetically related to a strain causing outbreaks in the United States. Antimicrob Agents Chemother. 2009;53:818–20. DOIPubMedGoogle Scholar
- Woodford N, Zhang J, Warner M, Kaufmann ME, Matos J, Macdonald A, Arrival of Klebsiella pneumoniae producing KPC carbapenemase in the United Kingdom. J Antimicrob Chemother. 2008;62:1261–4. DOIPubMedGoogle Scholar
- Cuzon G, Naas T, Demachy MC, Nordmann P. Plasmid-mediated carbapenem-hydrolyzing β-lactamase KPC in a Klebsiella pneumoniae isolate from Greece. Antimicrob Agents Chemother. 2008;52:796–7. DOIPubMedGoogle Scholar
- Naas T, Cuzon G, Villegas MV, Lartigue MF, Quinn JP, Nordmann P. Genetic structures at the origin of acquisition of the beta-lactamase blaKPC gene. Antimicrob Agents Chemother. 2008;52:1257–63. DOIPubMedGoogle Scholar
- Wolter DJ, Kurpiel PM, Woodford N, Palepou MF, Goering RV, Hanson ND. Phenotypic and enzymatic comparative analysis of the novel KPC variant KPC-5 and its evolutionary variants, KPC-2 and KPC-4. Antimicrob Agents Chemother. 2009;53:557–62. DOIPubMedGoogle Scholar
- Shen P, Wei Z, Jiang Y, Du X, Ji S, Yu Y, Novel genetic environment of the carbapenem-hydrolysing beta-lactamase KPC-2 among Enterobacteriaceae in China. Antimicrob Agents Chemother. 2009;53:4333–8. DOIPubMedGoogle Scholar
- Tegmark Wisell K, Haeggman S, Gazelius L, Thompson O, Gustafsson I, Ripa T, Identification of Klebsiella pneumoniae carbapenemase in Sweden. Euro Surveill. 2007;12:E071220.3.
- Clinical and Laboratory Standards Institute. Performance standards for antimicrobial susceptibility testing; fifteenth informational supplement. M100–S15. Wayne (PA): The Institute; 2005.
- Kieser T. Factors affecting the isolation of CCC DNA from Streptomyces lividans and Escherichia coli. Plasmid. 1984;12:19–36. DOIPubMedGoogle Scholar
- Sambrook J, Fritsch EF, Maniatis T. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor (NY): Cold Spring Harbor Laboratory Press; 1989.
- Carrër A, Lassel L, Fortineau N, Mansouri M, Anguel N, Richard C, Outbreak of CTX-M-15–producing Klebsiella pneumoniae in the intensive care unit of a French hospital. Microb Drug Resist. 2009;15:47–54. DOIPubMedGoogle Scholar
- Tenover FC, Arbeit R, Goering V, Mickelsen PA, Murray BE, Persing DH, Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: criteria for bacterial strain typing. J Clin Microbiol. 1995;33:2233–9.PubMedGoogle Scholar
- Diancourt L, Passet V, Verhoef J, Grimont PA, Brisse S. Multilocus sequence typing of Klebsiella pneumoniae nosocomial isolates. J Clin Microbiol. 2005;43:4178–82. DOIPubMedGoogle Scholar
- Carattoli A, Bertini A, Villa L, Falbo V, Hopkins KL, Threlfall EJ. Identification of plasmids by PCR-based replicon typing. J Microbiol Methods. 2005;63:219–28. DOIPubMedGoogle Scholar
- Falagas ME, Rafailidis PI, Ioannidou E, Alexiou VG, Matthaiou DK, Karageorgopoulos DE, Colistin therapy for microbiologically documented multidrug-resistant gram-negative bacterial infections: a retrospective cohort study of 258 patients. Int J Antimicrob Agents. 2010;35:194–9. Epub 2009 Dec 16. DOIPubMedGoogle Scholar
- Hæggman S, Löfdahl S, Paauw A, Verhoef J, Brisse S. Diversity and evolution of the class A chromosomal beta-lactamase gene in Klebsiella pneumoniae. Antimicrob Agents Chemother. 2004;48:2400–8. DOIPubMedGoogle Scholar
- Gootz TD, Lescoe MK, Dib-Hajj F, Dougherty BA, He W, Della-Latta P, Genetic organization of transposase regions surrounding blaKPC carbapenemase genes on plasmids from Klebsiella strains isolated in a New York City hospital. Antimicrob Agents Chemother. 2009;53:1998–2004. DOIPubMedGoogle Scholar
- Samuelsen O, Naseer U, Tofteland S, Skutlaberg DH, Onken A, Hjetland R, Emergence of clonally related Klebsiella pneumoniae isolates of sequence type 258 producing plasmid-mediated KPC carbapenemase in Norway and Sweden. J Antimicrob Chemother. 2009;63:654–8. DOIPubMedGoogle Scholar
- Baraniak A, Izdebski R, Herda M, Fiett J, Hryniewicz W, Gniadkowski M. The emergence of Klebsiella pneumoniae ST258 with KPC-2 in Poland. Antimicrob Agents Chemother. 2009;53:4565–7. DOIPubMedGoogle Scholar
- Damjanova I, Toth A, Paszti J, Hajbel-Vekony G, Jakab M, Berta J, Expansion and countrywide dissemination of ST11, ST15 and ST147 ciprofloxacin-resistant CTX-M-15–type beta-lactamase–producing Klebsiella pneumoniae epidemic clones in Hungary in 2005—the new “MRSAs”? J Antimicrob Chemother. 2008;62:978–85. DOIPubMedGoogle Scholar
- Endimiani A, Hujer AM, Perez F, Bethel CR, Hujer KM, Kroeger J, Characterization of blaKPC-containing Klebsiella pneumoniae isolates detected in different institutions in the eastern USA. J Antimicrob Chemother. 2009;63:427–37. DOIPubMedGoogle Scholar
- Willems RJ, Top J, Van den Braak N, Van Belkum A, Mevius DJ, Hendriks G, Molecular diversity and evolutionary relationships of Tn1546-like elements in enterococci from humans and animals. Antimicrob Agents Chemother. 1999;43:483–91.PubMedGoogle Scholar
Figures
Tables
Cite This Article1This study was presented in part at the Interscience Conference on Antimicrobial Agents and Chemotherapy; 2009 Sep 12–15; San Francisco, CA, USA.
Table of Contents – Volume 16, Number 9—September 2010
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Thierry Naas, Service de Bactériologie–Virologie, Hôpital de Bicêtre, 78 rue du Général Leclerc, 94275 Le Kremlin-Bicêtre CEDEX, France
Top