Volume 32, Number 4—April 2026
Dispatch
Rapid Spread of Recombinant African Swine Fever Virus Genotypes I and II, Vietnam, 2023–2024
Cite This Article
Citation for Media
Abstract
Molecular analyses of African swine fever (ASF) outbreaks in northern and central Vietnam during 2023–2024 revealed a rapid expansion (14.1%–42.2%) of recombinant ASF virus genotypes I and II. Increased prevalence and resistance to commercial ASF vaccines underscore the urgent need for better ASF control and an updated vaccine in Vietnam.
African swine fever (ASF) is a highly contagious viral disease affecting domestic and wild pigs; mortality rates reach up to 100% (1). The causative agent, ASF virus (ASFV), is a large double-stranded DNA virus in the Asfarviridae family. ASFV strains have been classified into 24 genotypes, of which only genotypes I and II have spread beyond Africa (2).
In Asia, ASFV genotype II was first detected in China in 2018 (3) and in Vietnam in early 2019 (4). Since 2021, recombinant ASFV strains containing genetic elements from genotypes I and II (rASFV I/II) have emerged in China and subsequently in Vietnam in 2023, presenting new challenges for ASF control (5,6). Two live attenuated ASFV genotype II vaccines have been licensed in Vietnam; however, they provide protection only against homologous strains (7,8) and are ineffective against rASFV I/II (9). Studies conducted during 2019–2022 investigated the molecular epidemiology of ASFV in Vietnam (10,11), but little is known about circulating strains during 2023–2024. In our study, we aimed to characterize ASFV strains causing outbreaks in northern and central Vietnam during 2023–2024, focusing on the emergence and spread of rASFV I/II.
During January 2023–August 2024, we collected 182 clinical samples (spleen, lymph nodes, lung, and whole blood) from pigs suspected of having ASFV infection across 26 provinces in northern and central Vietnam (Appendix Figure 1). We extracted total nucleic acids and tested them by using real-time PCR, targeting the B646L (p72) gene (VDx ASFV qPCR version 2.1; Median Diagnostics, http://mediandiagnostics.com). All samples tested positive for ASFV DNA; cycle threshold values ranged from 14.96 to 32 (Appendix Table). We performed in-house genotyping assays (Phan G1 and G2 Realtime-PCR) to differentiate ASFV genotypes I and II, as previously described (9). Results showed that 133/182 samples (73.08%) contained genotype II p72 and 49/182 samples (26.92%) contained genotype I p72. To confirm those findings, we conducted conventional PCRs targeting partial B646L (p72), complete E183L (p54), complete EP402R (CD2v), intergenic region (IGR), and central variable region (CVR), as previously described (11). We performed Sanger sequencing on amplicons (1st BASE, https://base-asia.com) and analyzed them by using Geneious Prime 2024 (Geneious, https://www.geneious.com), GraphPad Prism 8.4.3 (GraphPad Software, https://www.graphpad.com), and MEGA 11 (MEGA Software, https://www.megasoftware.net). We visualized phylogenetic trees by using iTOL version 7.4 (Interactive Tree of Life, https://itol.embl.de). We submitted all sequences to GenBank (accession nos. PV975276–639 and PX205390–33).
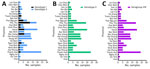
Sequence analysis confirmed that Phan G2–positive samples contained the p72 gene of ASFV genotype II, whereas Phan G1–positive samples carried the p72 gene of genotype I. Of note, all genotype I p72–positive samples also possessed the p54 gene of genotype II and the CD2v gene of serogroup VIII (Figure 1; Appendix Figure 2), consistent with previously reported rASFV I/II strains (5,6). We detected rASFV I/II in 17 of 26 provinces, whereas genotype II was present nationwide. We identified rASFV I/II in Bac Giang Province in May 2023, four months earlier than previously reported (5). A recent study demonstrated high levels of recombination between closely related ASFV genotypes in vitro and in vivo (12). In our study, the co-circulation of rASFV I/II and genotype II across multiple provinces suggests an increased risk for further recombination arising from co-infection, particularly in high-density production systems characterized by repeated exposure and prolonged environmental virus persistence. We further characterized ASFV strains on the basis of tandem repeat sequences in the IGR and amino acid variations in the CVR of the B602L gene. To date, 4 IGR variants (IGR I–IV) have been described, and in Vietnam, all 4 IGR types have been reported (10). In our study, among the 133 genotype II viruses analyzed, we identified 3 IGR variants (IGR I–III) that showed no intravariant nucleotide differences. IGR II remained dominant, being detected in all 26 provinces, whereas IGR III appeared in only 2 samples (from Bac Giang and Ha Nam Provinces). We found IGR I in 18 samples from 8 provinces (Appendix Figure 3). Those findings suggest a shift in IGR variant distribution among genotype II strains in Vietnam. In contrast, all rASFV I/II strains carried only the IGR II variant, consistent with previous reports (5).
Analysis of the B602L CVR region revealed 2 variants (CVR14 and CVR15) among genotype II viruses and 13 variants among rASFV I/II strains (Appendix Figure 4). Genotype II viruses contained 7–10 tetrameric amino acid repeats, whereas rASFV I/II exhibited 42–70 repeats, reflecting greater genetic diversity. The CVR14 variant dominated genotype II strains (131/133), whereas CVR15 was rare. Among rASFV I/II, CVR4 and CVR11 were the most prevalent, being detected across several provinces. Provinces such as Tuyen Quang, Thai Nguyen, Phu Tho, Hanoi, Bac Giang, Nam Dinh, and Thanh Hoa harbored multiple CVR variants, indicating ongoing viral diversification. Some CVR variants showed homology with previously reported strains: CVR2 matched the rASFV strain pig/Henan/123014/2022 from China (6), CVR4 corresponded to the Vietnam strain VNUA/rASFV/HD1/23 (5), and CVR3 resembled genotype I ASFV OURT 88/3 (13). Detection of 13 CVR variants among rASFV I/II strains in 2024 indicates ongoing ASFV evolution and underscores the need to clarify the relationship between genetic diversity and virulence among ASFV strains currently circulating in Vietnam.
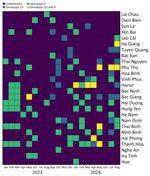
Spatial and temporal analyses showed a statistically significant increase (χ2 17.7; p<0.001) in rASFV I/II among ASFV-positive samples submitted for laboratory diagnosis and confirmation (Figure 2). We detected rASFV I/II in 17 of 26 provinces and observed the proportion increasing from 14.14% (14/99 samples) in 2023 to 42.17% (35/83 samples) in 2024 (Table). That trend suggests rapid geographic expansion and a potential replacement of ASFV genotype II by rASFV I/II. However, those results should be interpreted with caution because they are based on laboratory submissions rather than systematic field surveillance. The mechanisms underlying the shift are unclear but could be attributable to resistance to the vaccines, higher virulence, increased viral shedding, or greater transmission efficiency of the rASFV I/II strains.
Our study provides strong evidence that highly pathogenic rASFV I/II strains are rapidly evolving and spreading in northern and central Vietnam, showing indications of partial displacement of classical genotype II viruses. Detection of rASFV I/II as early as May 2023 suggests these recombinant strains circulated months before official reporting, highlighting the need for continuous genetic surveillance for early detection. The widespread co-circulation of genotype II and rASFV I/II across multiple provinces underscores the importance of genotyping-based real-time PCR in routine monitoring. Limited cross-protection of existing vaccines emphasizes the urgency of updated immunization strategies and novel vaccine platforms targeting rASFV I/II. Moreover, marked genetic heterogeneity, particularly in CVR regions, indicates ongoing viral evolution with potential effects on virulence and transmission. The co-circulation of multiple ASFV strains raises concern over the emergence of new recombinant variants, complicating disease control. Enhanced genomic surveillance, improved diagnostic tools, and systematic vaccine evaluation are therefore essential for effective ASF prevention and control in Vietnam and the region.
Mr. Nguyen is a veterinarian and a master’s student at Vietnam National University of Agriculture. His primary research interests include virology and animal pathology.
Acknowledgment
This work was supported by the Vietnam National Project (project code DTDL.CN-53/19), South Korea’s Animal and Plant Quarantine Agency, Ministry of Agriculture, Food and Rural Affairs (grant no. I-1543085-2025-26-01), and the Canadian Food Inspection Agency RPS Project (grant no. N-000534).
References
- Penrith ML, Vosloo W. Review of African swine fever: transmission, spread and control. J S Afr Vet Assoc. 2009;80:58–62. DOIPubMedGoogle Scholar
- Bastos AD, Penrith ML, Crucière C, Edrich JL, Hutchings G, Roger F, et al. Genotyping field strains of African swine fever virus by partial p72 gene characterisation. Arch Virol. 2003;148:693–706. DOIPubMedGoogle Scholar
- Zhou X, Li N, Luo Y, Liu Y, Miao F, Chen T, et al. Emergence of African swine fever in China, 2018. Transbound Emerg Dis. 2018;65:1482–4. DOIPubMedGoogle Scholar
- Le VP, Jeong DG, Yoon SW, Kwon HM, Trinh TBN, Nguyen TL, et al. Outbreak of African swine fever, Vietnam, 2019. Emerg Infect Dis. 2019;25:1433–5. DOIPubMedGoogle Scholar
- Le VP, Nguyen VT, Le TB, Mai NTA, Nguyen VD, Than TT, et al. Detection of recombinant African swine fever virus strains of p72 genotypes I and II in domestic pigs, Vietnam, 2023. Emerg Infect Dis. 2024;30:991–4. DOIPubMedGoogle Scholar
- Zhao D, Sun E, Huang L, Ding L, Zhu Y, Zhang J, et al. Highly lethal genotype I and II recombinant African swine fever viruses detected in pigs. Nat Commun. 2023;14:3096. DOIPubMedGoogle Scholar
- Diep NV, Ngoc NT, Duc NV, Dang VX, Tiep TN, Quy CT, et al. Safety and efficacy profiles of the live attenuated vaccine AVAC ASF LIVE for preventing African swine fever in pigs. Transbound Emerg Dis. 2025;2025:
8623876 . DOIPubMedGoogle Scholar - Tran XH, Le TTP, Nguyen QH, Do TT, Nguyen VD, Gay CG, et al. African swine fever virus vaccine candidate ASFV-G-ΔI177L efficiently protects European and native pig breeds against circulating Vietnamese field strain. Transbound Emerg Dis. 2022;69:e497–504. DOIPubMedGoogle Scholar
- Diep NV, Duc NV, Ngoc NT, Dang VX, Tiep TN, Nguyen VD, et al. Genotype II live-attenuated ASFV vaccine strains unable to completely protect pigs against the emerging recombinant ASFV genotype I/II strain in Vietnam. Vaccines (Basel). 2024;12:1114. DOIPubMedGoogle Scholar
- Mai NTA, Dam VP, Cho KH, Nguyen VT, Van Tuyen N, Nguyen TL, et al. Emergence of a novel intergenic region (IGR) IV variant of african swine fever virus genotype II in domestic pigs in Vietnam. Vet Res Commun. 2023;47:1773–6. DOIPubMedGoogle Scholar
- Mai NTA, Vu XD, Nguyen TTH, Nguyen VT, Trinh TBN, Kim YJ, et al. Molecular profile of African swine fever virus (ASFV) circulating in Vietnam during 2019–2020 outbreaks. Arch Virol. 2021;166:885–90. DOIPubMedGoogle Scholar
- Kitamura T, Masujin K, Ikezawa M, Ambagala A, Kokuho T. Generation of Chimeric African swine fever viruses through in vitro and in vivo intergenotypic gene complementation. Vaccines (Basel). 2025;13:462. DOIPubMedGoogle Scholar
- Chapman DAG, Tcherepanov V, Upton C, Dixon LK. Comparison of the genome sequences of non-pathogenic and pathogenic African swine fever virus isolates. J Gen Virol. 2008;89:397–408. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleOriginal Publication Date: April 10, 2026
1These authors contributed equally to this article.
Table of Contents – Volume 32, Number 4—April 2026
| EID Search Options |
|---|
|
|
|
|
|
|