Volume 11, Number 7—July 2005
Research
Human Metapneumovirus Genetic Variability, South Africa
Figure
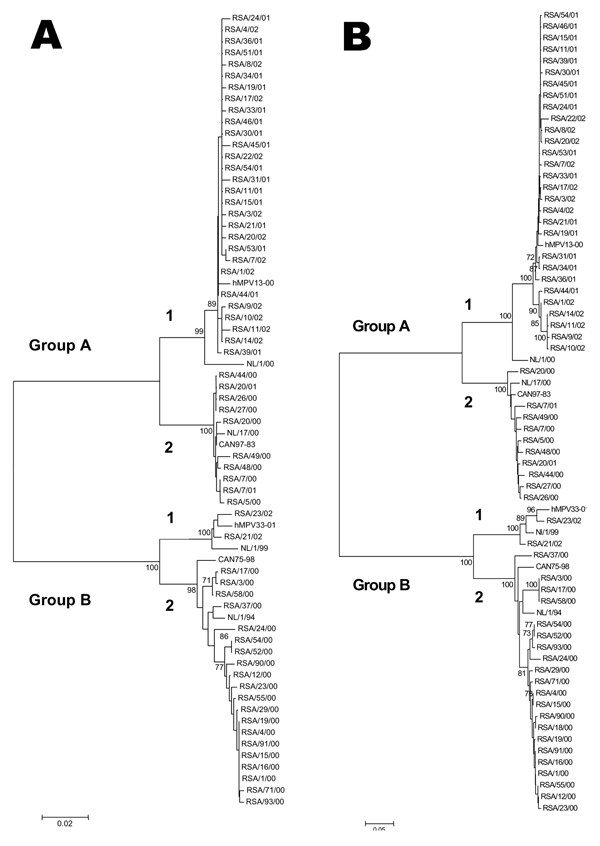
Figure. . . Neighbor-joining trees based on nucleotide sequences from A) the partial F gene and B) the G gene open reading frame from 61 South African human metapneumovirus (hMPV) isolates. The trees were computed with MEGA version 2.1 with the nucleotide Kimura 2-parameters. Bootstrap probabilities for 500 replicas are shown at the branch nodes. Only values of 70% to 100% are indicated. Isolates from South Africa are indicated by RSA, followed by the isolate number and year (e.g., RSA/18/02). The viruses from Canada (CAN97-83, hMPV13-00, CAN75-98, and hMPV33-01) and the Netherlands (NL/1/00, NL/17/00, NL/1/99, and NL/1/94) are prototypes from each subgroup.
Page created: April 24, 2012
Page updated: April 24, 2012
Page reviewed: April 24, 2012
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.