Volume 11, Number 8—August 2005
Dispatch
Bartonella quintana in Domestic Cat
Cite This Article
Citation for Media
Abstract
We recovered Bartonella quintana DNA from dental pulp of a domestic cat. This study, the first to detect B. quintana in a nonhuman mammal, changes our understanding of the epidemiology of this infection and proposes that cats may be an emerging source of human infection.
The α-proteobacterium Bartonella quintana is a fastidious, gram-negative organism; humans are the only known reservoir, and the human body louse, Pediculus humanus corporis, is the only known vector (1). Body lice infestation is linked to poor hygiene in homeless persons and persons engaged in war, as has been reported in several circumstances since trench fever was first described during World War I. B. quintana causes trench fever, chronic bacteremia, and endocarditis in homeless and alcoholic patients (2) and bacillary angiomatosis in both HIV-infected and immunocompetent patients (3). Rare cases of chronic lymph node infection caused by B. quintana were also reported (4,5). These patients were initially diagnosed with cat-scratch disease; they lived in conditions with high hygienic standards and had no evidence of infestation by body lice; they did have close contacts with cats and flea-infested kittens, however. Similarly, the source of B. quintana remains unknown in a few patients with B. quintana bacillary angiomatosis and endocarditis. Another investigation found a 4.5% prevalence of B. quintana in cat fleas collected in France (6). What is missing from these puzzling cases of B. quintana infection, however, is documentation of B. quintana in a cat. In this study, by using dental pulp of domestic cats to detect Bartonella spp. by polymerase chain reaction (PCR) that targets fragments of the pap31 gene, the 16S–23S internal transcribed spacer (ITS) (6,7), and 2 other genomic regions (8), we identified B. quintana in a cat.
Nine domestic cats collected in Marseille were euthanized for medical indications unrelated to infectious diseases. We collected 32 cuspid teeth from these cats (Table 1), although only 1 tooth from each cat was tested for Bartonella DNA. Dental pulp was extracted by using an original protocol involving external decontamination by 70% ethanol and setting the entire decontaminated tooth in sterile resin (Resin Polyester Sody 33, ESCIL, Chassieu, France). After polymerization at room temperature, the apex was removed from the tooth by using a sterilized disk, and the opened canal system was inserted upside down into a sterile Eppendorf tube and centrifuged at 8,000 rpm for 10 min to recover the dental pulp. Total DNA was then extracted according to standard phenol-chloroform protocol. A negative control (sterile water) was processed in parallel exactly as described above.
PCR amplifications were performed in a 25-μL reaction mixture containing 5 pmol of each primer (Eurogentec, Seraing, Belgium), 200 μmol/L each dNTP (Invitrogen, Cergy-Pontoise, France) in 10 mmol/L Tris-HCl (pH 8.3), 50 mmol/L KCl, 1.5 mmol/L MgCl2, 0.2 μg bovine serum albumin (Roche, Mannheim, Germany), 1 U Taq DNA polymerase (EuroblueTaq, Eurobio, Les Ulis, France), and 2 μL DNA. Primers PAPn1/PAPn2 targeting pap31 were previously described (7). Primers URBarto1/URBarto2 amplified a 639-bp/722-bp ITS fragment of B. henselae and B. quintana, respectively. This fragment has 67.7% sequence similarity between B. henselae and B. quintana (6). We also amplified 2 intergenic fragments, no. 336 (597 bp) and no. 894 (383 bp), which are specific for B. quintana and have been incorporated into multispacer typing of B. quintana (8). PCR included an initial 3-min step of denaturation at 94°C followed by 41 cycles of 30 s denaturation at 94°C, 30 s primer annealing at 58°C for pap31 primers (50°C for ITS primers), and 90 s elongation at 72°C. Amplification was completed by holding the reaction mixture at 72°C for 7 min. PCR products separated by 1.5% agarose gel electrophoresis were visualized by ethidium bromide staining, purified by using MultiScreen-PCR Filter Plate (Millipore, Saint-Quentin en Yvelines, France), and sequenced in both directions by using the d-Rhodamine Terminator Cycle Sequencing Ready Reaction kit (PerkinElmer, Coignières, France). Sequencing products were resolved in an Applied Biosystem automatic sequencer model 3100 (PerkinElmer).
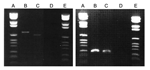
No amplification was observed for the negative controls in any PCR experiment. We obtained pap31 amplicons with DNA extracted from the teeth of cat 1 and cat 2. A 222-bp sequence derived from the tooth of cat 2 shared complete identity with that of B. quintana pap31 (GenBank accession no. AF308171), and a 237-bp sequence derived from the tooth of cat 1 shared 99% similarity with that of B. henselae ZF-1 (Houston genotype) pap31 (GenBank accession no. AF321116). One mutation, resulting in a glycine → aspartic acid shift at codon 137, differentiated query and reference sequences (GenBank accession no. AY839861) (Figure 1). ITS amplicons obtained from the same teeth (Figure 2) shared a complete sequence identity with B. henselae ITS (GenBank accession no. AF312496) in cat 1 and B. quintana ITS (GenBank accession no. AF368395) in cat 2 (Table 1). Sequences of fragment 336 in cat 2 shared 100% similarity with B. quintana reference sequence (GenBank accession no. AY660705) with 3 best BLAST scores ≥1,142 (E-value 0) and 82% similarity with B. henselae strain Houston-1 with BLAST score of 92 (E-value 2e–15). Sequence of fragment 894 from the same tooth shared 100% similarity with B. quintana reference sequence (GenBank accession no. AY660713) with 5 best BLAST scores ≥385 (E-value ≤e–104).
We found B. quintana and B. henselae DNA in the dental pulp of 2 domestic cats in France. To prevent contamination, we recovered pulp after the entire tooth was set in sterile resin. No amplification was obtained from controls, and no positive control was used. Amplicons were consistently obtained during separate PCR experiments targeting 4 different regions of the Bartonella genome. A unique mutation in the pap31 sequence derived from a specimen definitely ruled out contamination by modern laboratory Bartonella DNA. We previously detected B. henselae DNA in dental pulp from 13th- to 16th-century domestic cats (9) and from cats buried for 1 year (10). This study is, however, the first detection of B. henselae ZF-1, Houston genotype outside of cat-scratch disease lymph nodes (7).
B. quintana identity was confirmed by amplification of 2 genomic fragments not subject to genomic transfer and by high BLAST scores with 4 different molecular targets. Until now, B. quintana has been detected only in humans (2,3,5) and human body lice (1). We unexpectedly recovered B. quintana DNA from a cat's dental pulp, which gives a prevalence of 2.5% among 39 cats tested in 3 studies, including this one (9,10). B. henselae was found in 23% of cats, and B. clarridgeiae was the most prevalent species in cat fleas. These observations agree with a 4.5% prevalence of B. quintana recently observed in cat fleas in France (6) (Table 2), whereas it was not detected in biting flies from California (11). We suspected as early as 1994 that cats may play a role in B. quintana infection (4). We described 2 patients with either B. quintana chronic peripheral (4) or mediastinal adenomegaly (5) who lived in good hygienic conditions and had no evidence of body lice infestation but did have close contact with cats. Ongoing PCR and sequence-based survey of lymph nodes in patients suspected of cat-scratch disease in Marseille found 11.2% B. henselae and 1 additional case of B. quintana (Table 2). A few additional patients have been reported (12). Likewise, 1 of 14 patients with B. quintana bacillary angiomatosis did not have risk factors, including low income, homelessness, and exposure to lice, but did have contact with cats (3,13). The same observation holds true for 3 of 38 patients with B. quintana endocarditis who did not have risk factors, including homelessness, alcoholism, and exposure to body lice, but did have contact with cats or cat fleas. These data led us to hypothesize that a B. quintana bacteremic domestic cat could be a rare source for B. quintana human infection. If confirmed, these data may lead to a recommendation that immunocompromised patients and patients at risk for endocarditis avoid contact with cats.
Present data reinforce the idea that dental pulp is a suitable specimen on which to base PCR detection of bloodborne bacteria. In addition to our work on feline bartonellosis, we detected B. quintana in the dental pulp of a homeless patient with previous bacteremia (14) and in a 4,000-year-old cadaver (15). One may speculate on a common ancestor of B. henselae and B. quintana in cats, with B. quintana evolution toward a more specific niche. Further use of cat dental pulp to detect and genotype B. quintana may confirm these data and refine cat-based epidemiology and diagnosis of poorly understood clinical forms of B. quintana human infection.
Dr. La is a PhD student at Université de la Méditerranée—Unité des Rickettsies. He studies bacterial colonization and use of dental pulp to detect bloodborne pathogens.
Acknowledgment
The authors acknowledge Vijay A.K.B Gundi for expert review of the manuscript and Nash Laurent for assistance with animals.
References
- Maurin M, Raoult D. Bartonella (Rochalimaea) quintana infections. Clin Microbiol Rev. 1996;9:273–92.PubMedGoogle Scholar
- Brouqui P, Lascola B, Roux V, Raoult D. Chronic Bartonella quintana bacteremia in homeless patients. N Engl J Med. 1999;340:184–9. DOIPubMedGoogle Scholar
- Koehler JE, Sanchez MA, Garrido CS, Whitfeld MJ, Chen FM, Berger TG, Molecular epidemiology of Bartonella infections in patients with bacillary angiomatosis-peliosis. N Engl J Med. 1997;337:1876–83. DOIPubMedGoogle Scholar
- Raoult D, Drancourt M, Carta A, Gastaut JA. Bartonella (Rochalimaea) quintana isolation in patient with chronic adenopathy, lymphopenia, and a cat. Lancet. 1994;343:977. DOIPubMedGoogle Scholar
- Drancourt M, Moal V, Brunet P, Dussol B, Berland Y, Raoult D. Bartonella (Rochalimaea) quintana infection in a seronegative hemodialyzed patient. J Clin Microbiol. 1996;34:1158–60.PubMedGoogle Scholar
- Rolain JM, Franc M, Davoust B, Raoult D. Molecular detection of Bartonella quintana, B. koehlerae, B. henselae, B. clarridgeiae, Rickettsia felis, and Wolbachia pipientis in cat fleas, France. Emerg Infect Dis. 2003;9:338–42.PubMedGoogle Scholar
- Zeaiter Z, Fournier PE, Raoult D. Genomic variation of Bartonella henselae strains detected in lymph nodes of patients with cat scratch disease. J Clin Microbiol. 2002;40:1023–30. DOIPubMedGoogle Scholar
- Foucault C, La Scola B, Lindroos H, Andersson SG, Raoult D. Multi-spacer-typing (MST) for sequence-based typing of Bartonella quintana. J Clin Microbiol. 2005;43:41–8. DOIPubMedGoogle Scholar
- La VD, Clavel B, Lepetz S, Aboudharam G, Raoult D, Drancourt M. Molecular detection of Bartonella henselae DNA in the dental pulp of 800-year-old French cats. Clin Infect Dis. 2004;39:1391–4. DOIPubMedGoogle Scholar
- Aboudharam G, La VD, Davoust B, Drancourt M, Raoult D. Molecular detection of Bartonella spp. in the dental pulp of stray cats buried for a year. Microb Pathog. 2005;38:47–51. DOIPubMedGoogle Scholar
- Chung CY, Kasten RW, Paff SM, Van Horn BA, Vayssier-Taussat M, Boulouis HJ, Bartonella spp. DNA associated with biting flies from California. Emerg Infect Dis. 2004;10:1311–3.PubMedGoogle Scholar
- Azevedo ZM, Higa LY, Boechat PR, Boechat MB, Klaplauch F. Cat-scratch disease caused by Bartonella quintana in an infant: an unusual presentation. Rev Soc Bras Med Trop. 2000;33:313–7.PubMedGoogle Scholar
- Gasquet S, Maurin M, Brouqui P, Lepidi H, Raoult D. Bacillary angiomatosis in immunocompromised patients. AIDS. 1998;12:1793–803. DOIPubMedGoogle Scholar
- Aboudharam G, Fournier PE, Drancourt M, Raoult D, Foucault C, Brouqui P. Molecular detection of Bartonella quintana DNA in the dental pulp of a homeless patient. Eur J Clin Microbiol Infect Dis. 2004;23:920–2.PubMedGoogle Scholar
- Drancourt M, Tran-Hung L, Courtin J, De Lumley H, Raoult D. Bartonella quintana in a 4,000-year-old human tooth. J Infect Dis. 2005;191:607–11. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleTable of Contents – Volume 11, Number 8—August 2005
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Didier Raoult, Unité des Rickettsies, CNRS UMR 6020, IFR 48, Faculté de Médecine, Université de la Méditerranée, Marseille, France; fax: 33-491-38-77-72
Top