Volume 15, Number 8—August 2009
Research
Bordetella pertussis Strains with Increased Toxin Production Associated with Pertussis Resurgence
Cite This Article
Citation for Media
Abstract
Before childhood vaccination was introduced in the 1940s, pertussis was a major cause of infant death worldwide. Widespread vaccination of children succeeded in reducing illness and death. In the 1990s, a resurgence of pertussis was observed in a number of countries with highly vaccinated populations, and pertussis has become the most prevalent vaccine-preventable disease in industrialized countries. We present evidence that in the Netherlands the dramatic increase in pertussis is temporally associated with the emergence of Bordetella pertussis strains carrying a novel allele for the pertussis toxin promoter, which confers increased pertussis toxin (Ptx) production. Epidemiologic data suggest that these strains are more virulent in humans. We discuss changes in the ecology of B. pertussis that may have driven this adaptation. Our results underline the importance of Ptx in transmission, suggest that vaccination may select for increased virulence, and indicate ways to control pertussis more effectively.
Bordetella pertussis causes whooping cough or pertussis, a respiratory disease that is most severe in infants. Before childhood vaccination was introduced in the 1950s, pertussis was a major cause of infant deaths worldwide. Widespread vaccination of children reduced the incidence of illness and deaths caused by pertussis (1). However, globally pertussis remains 1 of the top 10 causes of death in children (2). Further, in the 1990s a resurgence of pertussis was observed in several countries with highly vaccinated populations (3,4), and pertussis has become the most prevalent vaccine-preventable disease in industrialized countries. In the Netherlands, the estimated incidence of infection was 6.6% per year for the 3–79-year age group from 1995 through 1996 (5). Similar percentages have been found in the United States (6). One of the hallmarks of the pertussis resurgence is a shift in disease prevalence toward older persons who have waning vaccine-induced immunity (7).
The reemergence of pertussis has been attributed to various factors, including increased awareness, improved diagnostics, decreased vaccination coverage, suboptimal vaccines, waning vaccine-induced immunity, and pathogen adaptation. The relative contribution of these factors may differ between countries and is the subject of ongoing debate. Pathogen adaptation is supported by several observations. We and others have shown that antigenic divergence has occurred between vaccine strains and clinical isolates with respect to surface proteins, which confer protective immunity: pertussis toxin (Ptx), pertactin (Prn), and fimbriae (8,9). Strain variation was shown to affect vaccine efficacy in a mouse model (10–13). Because adaptation may involve the structure of virulence factors (by antigenic variation) and their regulation, we extended our studies on the evolution of B. pertussis by investigating polymorphism in the promoter of Ptx (ptxP), a major virulence factor and component of all pertussis vaccines (1). We provide evidence that expansion of strains with increased Ptx production has contributed to the resurgence of pertussis in the Netherlands.
Pertussis Notifications
Pertussis became a notifiable disease in the Netherlands in 1976. Notifications are submitted online by local health authorities. Other notifiable diseases are also monitored through this system, which falls under the responsibility of the Dutch National Institute of Health and Environment (3).
Bacterial Strains
B. pertussis strains examined were obtained from 1935 through 2004. A total of 1,566 isolates, 879 from the Netherlands and 687 from other countries, were analyzed for polymorphism in ptxP (Technical Appendix). Eight strains isolated from patients in the Netherlands from 1999 through 2001 were selected to study Ptx and Prn production: B1834 (ptxP1), B1868 (ptxP1), B1878 (ptxP1), B1920 (ptxP1), B1836 (ptxP3), B1865 (ptxP3), B1917 (ptxP3), and B2030 (ptxP3) (Table 1).
Sequencing
The primers 5′-AATCGTCCTGCTCAACCGCC-3′ and 5′-GGTATACGGTGGCGGGAGGA-3′ were used for amplification and sequencing of ptxP and correspond, respectively, to bases 60–79 and 633–614 of the ptx sequence with GenBank accession no. M14378. The ptx gene cluster from the strains B1834 (ptxP1), B1920 (ptxP1), B1917 (ptxP3), and B1831 (ptxP3) was sequenced completely. The sequences of the ptx gene clusters from strains B1834, B1920, B1917, and B1831 can be found under the following accession numbers, respectively: FN252334, FN252335, FN252336, and FN252333. The ptxP1-ptxP11 sequences have been assigned accession nos. FN252323, FN252322, FN252324, FN252325, FN252326, FN252327, FN252328, FN252329, FN252330, FN252331, and FN252332.
Pertussis Toxin and Pertactin Production
B. pertussis strains were grown on Bordet-Gengou agar plates supplemented with 15% sheep blood and incubated for 3 days at 35°C. Cells were harvested and suspended in 2 mL Verwey medium (14) per plate. Cells from 1 mL were collected by centrifugation and resuspended in Verwey medium to a concentration of 5 × 106 bacteria/mL. Subsequently, 100 µL of this suspension (5 × 105 CFU) was plated on Bordet-Gengou agar plates. After an incubation of 48 to 60 hours at 35°C, cells were harvested in 2.5 mL Verwey medium. The cell suspension was heat-inactivated for 30 min at 56°C and stored at 4°C. An ELISA was used to quantify Ptx and Prn. For Ptx, Maxisorp 96-well plates (Nunc International, Rochester, NY, USA) were coated with 100 µL of 0.04 mg/mL fetuin (Sigma-Aldrich, St. Louis, MO, USA) in 0.04 M carbonate buffer, pH 9.6, overnight at 4°C. For Prn, polystyrene 96-well plates (Immunolon II; Dynatech, Chantilly, VA, USA) were coated with 100 µL of a 2,000-fold dilution of polyclonal rabbit anti-Prn immunoglobulin (Ig)G (15) in 0.04 M carbonate buffer, pH 9.6, overnight at 20°C. Plates were blocked by incubation with 130 µL 1% bovine serum albumin (Sigma-Aldrich) in phosphate-buffered saline (PBS) for 1 hour at 37°C, after which plates were washed twice with PBS supplemented with 0.05% Tween. A 3-fold serial dilution of the heat-inactivated cell suspensions was made in 100 µL PBS supplemented with 0.1% Tween (PBST); 1 µg/mL of Prn and Ptx were used as reference. The suspensions were incubated for 1 hour at 37°C followed by 2 washings. The Prn monoclonal antibody (MAb) (PeM85) that was used binds to the linear epitope GGFGPGGFGP present in the repeat region 1 of all known Prn variants, except Prn13 (15). The Ptx MAb (3F10) binds to a conformational epitope in the PtxA subunit (16). All strains selected for the ELISA experiments produced Prn2 and PtxA1 (Table 1). The MAbs were diluted in PBST, added to the wells, and incubated for 1 hour at 37°C, followed by 2 washings. To detect bound MAbs, plates were incubated with horseradish peroxidase–conjugated polyclonal rabbit anti-mouse IgG (DakoCytmation, Glostrup, Denmark), diluted in PBST, for 1 hour at 37°C, and followed by 2 washings. The optical density at 450 nm was measured with a plate reader (PowerWave HT 340; Biotek, Winooski, VT, USA) and the amount of produced Ptx and Prn were calculated using the KC4 program (Biotek).The ratio of Ptx and Prn production by ptxP1 and ptxP3 strains was calculated as follows: Ptx (or Prn) production ptxP3 strains divided by Ptx (or Prn) production ptxP1 strains.
Statistical Analyses
The significance of the increases in illness and death were calculated with the Fisher exact test. Ptx and Prn production was analyzed on the basis of the following considerations: 1) that there are random variations among experiments that influence Ptx and Prn production; 2) that there is a correlation between Ptx and Prn production; and 3) that the distribution Ptx and Prn measurements were skewed. To take into account these considerations regarding sources of random variation, a random intercept model was used and a logarithmic transformation was used before further analysis. Logarithmically transformed Ptx and Prn values were first analyzed with a random intercept model by using SAS PROC MIXED (SAS, Cary, NC, USA) and by using experiment as a random effect. We first tested whether there were differences between ptxP1 and ptxP3 strains in the production of Ptx and Prn by analyzing the logarithm of Ptx production and Prn production, respectively, as a dependent variable, and by using experiment as random effect and incubation time (in classes) and type (ptxP1 or ptxP3) as fixed effects. To determine whether the ratio of production in ptxP3 versus ptxP1 strains differ significantly for Prn and Ptx, we further fitted a multivariate model with both factors (Ptx and Prn) as dependent variables, again using experiment as random effect, and allowing all variance parameters to be factor (Ptx or Prn) specific. In this model the interaction between type (ptxP1 or ptxP3) with factor (Ptx or Prn) then gives the required P value.
Polymorphism of the Pertussis Toxin Promoter
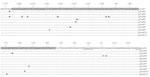
The synthesis and export of Ptx requires 14 genes, which are co-transcribed from ptxP (17). ptxP comprises a region of ≈170 bases upstream of the Ptx subunit gene ptxA and contains the RNA polymerase binding site and 6 binding sites for the BvgA dimer (18). BvgA is a global regulator of B. pertussis virulence genes, and cooperative binding of BvgA to ptxP is required for efficient transcription of ptx (18). We investigated polymorphism in ptxP by sequencing a DNA region of ≈380 bases upstream of ptxA by using a collection of 1,566 B. pertussis strains from 12 countries isolated during 1935–2004. Polymorphism was found to be restricted to the DNA region implicated in binding of RNA polymerase and BvgA. Eleven ptxP alleles were identified (Figure 1).
Geographic and Temporal Differences in ptxP Frequencies
Next we investigated geographic and temporal differences in ptxP frequencies. The following geographic regions were distinguished: the Netherlands, the continents of Africa, Asia, Europe (excluding the Netherlands), North America, and South America. Two periods, chosen on the basis of the appearance of ptxP3 strains in the Netherlands, were compared: 1935 through 1990 and 1991 through 2004 (Table 2). Only strains from the later period were available from South America. Two ptxP alleles were found to predominate worldwide, ptxP1 and ptxP3, and the remaining 9 alleles were pooled. Strains with ptxP1 predominated in the first period and were found in lower frequencies in the second period (global frequencies were 88% and 47%, respectively). A reverse trend was observed for the ptxP3 strains (global frequencies, 3% and 52%, respectively). In the first period, ptxP3 strains were only detected in the Netherlands and the United States (frequencies were 3% and 13%, respectively). The only region in which ptxP3 strains were not detected was Africa, where only ptxP1 strains were found. The minor ptxP alleles were observed in higher frequencies during 1935–1990 compared with 1991–2004 (global frequencies were 9% and 1%, respectively). The differences in ptxP allele frequencies may be due to sampling bias, geographic factors, or differences in vaccines, vaccination history, and vaccination coverage. Nevertheless, these data provide strong evidence that, in most parts of the world, ptxP3 strains emerged recently and replaced the resident ptxP1 strains. The ptxP3 allele was first detected in a strain isolated in the United States in 1984.
To investigate if ptxP1 and ptxP3 alleles were linked to other polymorphisms in ptx genes, the gene clusters from 2 ptxP1 and 2 ptxP3 strains were sequenced. The ptx sequences were identical, except for a single point mutation in ptxC. The single nucleotide polymorphism (SNP) in ptxC has been described previously, does not result in a change in amino acid sequence, and is therefore most likely selectively neutral (19). To study the linkage, ptxC was sequenced in 249 ptxP1 and 142 ptxP3 strains. Linkage between ptxP1-ptxC1 and ptxP3-ptxC2 was 100% and 98%, respectively. Only 3 strains harbored the combination ptxP3-ptxC1.
Association of the ptxP3 Allele with the Resurgence of Pertussis in the Netherlands
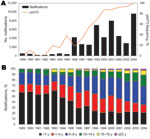
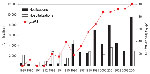
The availability of a large strain collection allowed us to analyze temporal trends in the Netherlands in more detail. From 1989 through 2004, a total of 99% of the strains harbored ptxP1 or ptxP3. In this period, ptxP1 was gradually replaced by ptxP3, which increased in frequency from 0% in 1989 to 100% in 2004. A close temporal relationship was shown between the increase in ptxP3 frequency and mandatory pertussis notifications (Figure 2, panel A). Increased notifications were found in all age groups, however, the largest increase was among persons >5 years of age (Figure 2, panel B). The shift toward older age categories coincided with emergence of ptxP3 strains. There was no change in age distribution from 1989 through 1992, which preceded the emergence of the ptxP3 allele.
Ptx and Prn Production by ptxP1 and ptxP3 Strains
The effect of polymorphism in ptxP was assessed by determining the ratio of Ptx produced by ptxP1 and ptxP3 strains (Ptx produced by ptxP3 strain / Ptx produced by ptxP1 strain) after 48, 54, and 60 h growth on plates. In addition, we assessed the production of a second virulence factor, Prn, which is also regulated by bvg. No polymorphism was observed in the Prn promoter of the 8 strains analyzed. Data from 4 ptxP1 and 4 ptxP3 strains were pooled (Figure 3). The Prn ratios were slightly lower than 1, indicating that ptxP3 strains produce slightly less Prn than ptxP1 strains (average over all time points 0.94; p = 0.03). In contrast, the Ptx ratio was significantly larger than 1 (average over all time points 1.62; p<0.0001), indicating that ptxP3 strains produce more Ptx than ptxP1 strains under the growth conditions tested. The Ptx and Prn ratios were significantly different (p<0.0001).
Evidence for Increased Virulence of ptxP3 Strains in Humans
Ptx is assumed to contribute to the severity of infection. Therefore, we investigated whether ptxP3 strains were more virulent in humans than ptxP1 strains by comparing the incidence of hospitalizations, deaths, and lethality (ratio of deaths to hospitalizations) in the Netherlands during 2 periods, 1981 through 1992 and 1993 through 2004, with low (1.6%) and high (54.5%) ptxP3 frequencies, respectively (Table 3). All 3 parameters showed a statistically significant increase (1.41, 10.21, and 7.23 times respectively; p values <0.0001, 0.0058, and 0.03, respectively), suggesting that ptxP3 strains are more virulent in humans.
The persistence of pertussis in the face of intense vaccination is unexpected because B. pertussis is extremely homogeneous (19–21), implying a limited ability to adapt. However, the Ptx promoter showed a relatively high degree of polymorphism, suggesting that fine tuning of Ptx production has adaptive value. Globally, 11 ptxP alleles were found in 1,566 strains, 8 of which occurred in the B. pertussis population in the Netherlands. Polymorphism was restricted to a region required for transcription of ptx. Silent ptx genes are found in the closely related species B. parapertussis and B. bronchiseptica (17). The silencing of ptx genes indicates that production of Ptx involves benefits and costs. Thus, production of Ptx is beneficial for the pathogen by suppressing host defenses but also involves metabolic costs and increases the number of immunologic targets. Ptx is a major antigen of B. pertussis, and Ptx antibodies are used in diagnosing pertussis cases.
Globally, ptxP1 and ptxP3 were the most prevalent ptxP alleles. In the Netherlands, during 1989–2004, ptxP1 was completely replaced by ptxP3. The replacement of ptxP1 strains by ptxP3 strains in recent times is a global phenomenon because it has been observed in 11 countries representing 4 continents; Asia, Europe, and North and South America. Notably, ptxP3 strains were not observed in Africa. A broad current distribution of ptxP3 strains was also suggested by a recent study in which strains from 8 European countries were compared by pulsed-field gel electrophoresis (PFGE). One PFGE profile, BpSR11, predominated in 5 of the 8 European countries (22). We have found that in the Dutch population all BpSR11 strains carry the ptxP3 allele (N = 18).
In the Netherlands, emergence of ptxP3 strains was associated with increased notifications and a shift in disease prevalence toward older age categories. Changes in diagnostic procedures may have contributed to the latter 2 phenomena (3). However, hospitalizations, which are less sensitive to surveillance artifacts, also increased concurrently with the emergence of ptxP3 strains (Appendix Figure). Furthermore, an extensive analysis of surveillance data confirmed a true increase in the pertussis incidence after 1995 in the Netherlands (3). The expansion of ptxP3 strains was also associated with the resurgence of pertussis in Finland, where a large nationwide epidemic was observed in 2003 (23).
The SNP distinguishing the ptxP1 and ptxP3 alleles is located in a region involved in binding of BvgA, the global regulator of virulence gene expression in B. pertussis. We hypothesize that the ptxP3 allele confers increased binding of BvgA compared to ptxP1, resulting in increased toxin production. When compared with ptxP1 strains, ptxP3 strains produced 1.62 times more Ptx. In contrast, the production of another bvg-regulated virulence factor, Prn, was slightly suppressed in ptxP3 strains compared with ptxP1 strains (factor 0.94), indicating that increased Ptx production cannot be explained by a global up-regulation of virulence genes.
The expansion of ptxP3 strains is remarkable and suggests that ptxP3 increases strain fitness or is linked to other genetic loci that do so. Although we cannot exclude that other loci are involved in the expansion of ptxP3 strains, several arguments underline the role of ptxP3. First, the high degree of polymorphism in the ptxP promoter indicates positive selection. Second, the increased Ptx production observed by ptxP3 strains provides a rationale for its emergence. It has been well established that Ptx plays a central role in immune suppression. Ptx enhances colonization of naive and immune mice by targeting macrophages and neutrophils (24,25). Ptx also suppresses antibody responses (26). The ptxP3 allele was found to be associated with 2 ptxC alleles, ptxC1 and ptxC2, which are distinguished by a silent SNP. This finding suggests that the ptxP3 allele is found in different genetic backgrounds, which may be explained by homoplasy or horizontal gene transfer. Both possibilities suggest that ptxP3 confers increased fitness. In most strains (98%), ptxP3 was linked to ptxC2. Furthermore, genomic profiling of Dutch B. pertussis strains indicates that ptxP3 strains are closely related, and are characterized by a chromosomal deletion (27). Thus, it is likely that, in the Netherlands, ptxP3 strains arose mainly by clonal expansion. We are analyzing a geographically more diverse strain collection to investigate this issue further.
Ptx has been suggested to increase severity of B. pertussis infections because the closely related B. parapertussis, which does not produce Ptx, generally causes less severe infections (28). Furthermore, Ptx causes leukocytosis in humans by inhibiting egression of leukocytes from the vasculature, and high levels of leukocytosis are associated with an increased mortality rate in infants due to pulmonary hypertension (29). Thus, the invasion of ptxP3 strains may result in increased illness and death. Consistent with this assumption, we found that the emergence of ptxP3 strains in the Netherlands was associated with increased incidence of hospitalizations and deaths and increased lethality. A recent Swedish study also suggested that B. pertussis strains differ in virulence. Infection with strains with PFGE profile BpSR11 was associated with a longer duration of hospital stay (30). As noted above, BpSR11 strains carry the ptxP3 allele. An association between Fim2 and increased disease severity was found in a study in the UK (31). In contrast, the Swedish study found no association between Fim type and virulence (30). Nevertheless, it is conceivable that other polymorphic loci in B. pertussis may also affect virulence.
An important issue is whether vaccination has selected for the ptxP3 strains. Several lines of evidence support this contention. First, ptxP3 strains were not found in the prevaccination era. Furthermore, although ptxP3 strains were found in high frequencies in vaccinated populations in the 1990s, they were not detected in Senegal, where vaccination was introduced in 1987 (32). Several studies have provided evidence that increased host immunity may select for higher virulence. Vaccination against 2 avian viruses, the Marek disease virus, and the infectious bursal disease virus, were associated with the emergence of more virulent strains (33). An important role of host immunity in selecting for virulence is also suggested by the co-evolution of the myxomatosis virus and rabbits (34). Furthermore, immune pressure was shown to select for more virulent Plasmodium chabaudi parasites in mice (35). Based on mathematical modeling, vaccines designed to reduce pathogen growth rate and/or toxicity may result in the evolution of pathogens with higher levels of virulence (36).
We propose that the crucial event, which shifted the competitive balance between ptxP1 and ptxP3 strains, was the removal by vaccination of immunologically naive infants as the major source for transmission, selecting for strains, which are more efficiently transmitted by primed hosts. Recent studies and historical data indicate an important role of naïve infants in transmission in unvaccinated populations. In a previously unvaccinated population, infant vaccination resulted in a reduction in pertussis in the vaccinated and unvaccinated parts of the population (37). Furthermore, in unvaccinated populations, 60%–80% of the pertussis cases were found in children 0–5 years of age, most of whom were probably immunologically naive (32,38). In most countries infants receive their first vaccination at the age of 2 or 3 months, essentially eliminating transmission by immunologically naive hosts. In primed hosts, increased Ptx production may delay an effective immune response (24–26) enhancing transmission, and hence, pathogen fitness. Increased Ptx production may also be beneficial for the pathogen because the host requires higher levels of antibodies against Ptx for toxin neutralization. The antigenic divergence observed between vaccine strains and circulating strains (8,9) may act synergistically with the ptxP3 polymorphism by enhancing transmission by hosts primed by vaccination. Pertussis among recently vaccinated children is rare, indicating that pathogen adaptation does not play a role unless immunity has waned. Thus, we propose that waning immunity and pathogen adaptation have contributed to the resurgence of pertussis, although other factors such as increased awareness and improved diagnostics have also played a role.
The effect of pathogen adaptation on disease impact may depend on factors such as vaccine coverage and the quality of the vaccine used, which may differ between countries. A relatively weak vaccine used in the Netherlands may have exacerbated the effect of the emergence of ptxP3 strains on disease impact (3). Our results underline the important role of Ptx in the transmission of B. pertussis and suggest that an effective way to control pertussis is the improvement of current vaccines to induce Ptx-neutralizing antibodies which persist longer. An important question is whether other childhood vaccines also select for pathogens that are more efficiently transmitted by primed hosts, resulting in increased virulence.
Dr Mooi is a senior scientist at the National Institute of Public Health and the Environment,the Netherlands. His current interests include the molecular epidemiology and evolution of Bordetella spp., particularly in the context of vaccination.
Acknowledgments
We are grateful to Hendriek Boshuizen for the statistical analyses of Ptx and Prn production.
This study was supported by a grant from the Netherlands Ministry of Public Health, Welfare, and Sport.
References
- Edwards KM, Decker MD. Pertussis vaccine. In: Plotkin SA, Orenstein WA, editors. Vaccines. 4th ed. Philadelphia: Saunders; 2004. p. 471–528.
- Crowcroft NS, Stein C, Duclos P, Birmingham M. How best to estimate the global burden of pertussis? Lancet Infect Dis. 2003;3:413–8. DOIPubMedGoogle Scholar
- de Melker HE, Schellekens JF, Neppelenbroek SE, Mooi FR, Rumke HC, Conyn-van Spaendonck MA. Reemergence of pertussis in the highly vaccinated population of the Netherlands: observations on surveillance data. Emerg Infect Dis. 2000;6:348–57. DOIPubMedGoogle Scholar
- Edwards KM. Overview of pertussis: focus on epidemiology, sources of infection, and long term protection after infant vaccination. Pediatr Infect Dis J. 2005;24(Suppl):S104–8. DOIPubMedGoogle Scholar
- de Melker HE, Versteegh FG, Schellekens JF, Teunis PF, Kretzschmar M. The incidence of Bordetella pertussis infections estimated in the population from a combination of serological surveys. J Infect. 2006;53:106–13. DOIPubMedGoogle Scholar
- Cherry JD. The epidemiology of pertussis: a comparison of the epidemiology of the disease pertussis with the epidemiology of Bordetella pertussis infection. Pediatrics. 2005;115:1422–7. DOIPubMedGoogle Scholar
- Halperin SA. The control of pertussis—2007 and beyond. N Engl J Med. 2007;356:110–3. DOIPubMedGoogle Scholar
- Mooi FR, He Q, Guiso N. Phylogeny, evolution, and epidemiology of Bordetellae. In: Locht C, editor. Bordetella molecular microbiology, 1st ed. Norfolk (UK): Horizon Bioscience; 2007. p. 17–45.
- Mooi FR, van Oirschot H, Heuvelman K, van der Heide HG, Gaastra W, Willems RJ. Polymorphism in the Bordetella pertussis virulence factors P.69/pertactin and pertussis toxin in the Netherlands: temporal trends and evidence for vaccine-driven evolution. Infect Immun. 1998;66:670–5.PubMedGoogle Scholar
- Gzyl A, Augustynowicz E, Gniadek G, Rabczenko D, Dulny G, Slusarczyk J. Sequence variation in pertussis S1 subunit toxin and pertussis genes in Bordetella pertussis strains used for the whole-cell pertussis vaccine produced in Poland since 1960: efficiency of the DTwP vaccine-induced immunity against currently circulating B. pertussis isolates. Vaccine. 2004;22:2122–8. DOIPubMedGoogle Scholar
- Fingermann M, Fernandez J, Sisti F, Rodriguez ME, Gatti B, Bottero D, Differences of circulating Bordetella pertussis population in Argentina from the strain used in vaccine production. Vaccine. 2006;24:3513–21. DOIPubMedGoogle Scholar
- Watanabe M, Nagai M. Effect of acellular pertussis vaccine against various strains of Bordetella pertussis in a murine model of respiratory infection. J Health Sci. 2002;48:560–4. DOIGoogle Scholar
- King AJ, Berbers G, van Oirschot HF, Hoogerhout P, Knipping K, Mooi FR. Role of the polymorphic region 1 of the Bordetella pertussis protein pertactin in immunity. Microbiology. 2001;147:2885–95.PubMedGoogle Scholar
- Verwey W, Thiele E, Sage D, Suchardt L. A simplified liquid culture medium for the growth of Haemophilus pertussis. J Bacteriol. 1949;58:127–34.
- Hijnen M, He Q, Schepp R, van Gageldonk P, Mertsola J, Mooi FR, Antibody responses to defined regions of the Bordetella pertussis virulence factor pertactin. Scand J Infect Dis. 2008;40:94–104. DOIPubMedGoogle Scholar
- Bartoloni A, Pizza M, Bigio M, Nucci D, Ashworth LA, Irons LI, Mapping of a protective epitope of pertussis toxin by in vitro refolding of recombinant fragments. Bio/Technology. 1988;6:709–12. DOIGoogle Scholar
- Hewlett EL, Donato GM. Bordetella toxins. In: Locht C, editor. Bordetella molecular microbiology. Norfolk (UK): Horizon Bioscience; 2007. p. 97–118.
- Stibitz S. The bvg regulon. In: Locht C, editor. Bordetella molecular microbiology. Norfolk (UK): Horizon Bioscience; 2007. p. 47–67.
- van Loo IH, Heuvelman KJ, King AJ, Mooi FR. Multilocus sequence typing of Bordetella pertussis based on surface protein genes. J Clin Microbiol. 2002;40:1994–2001. DOIPubMedGoogle Scholar
- Diavatopoulos DA, Cummings CA, Schouls LM, Brinig MM, Relman DA, Mooi FR. Bordetella pertussis, the causative agent of whooping cough, evolved from a distinct, human-associated lineage of B. bronchiseptica. PLoS Pathog. 2005;1:e45. DOIPubMedGoogle Scholar
- Parkhill J, Sebaihia M, Preston A, Murphy LD, Thomson N, Harris DE, Comparative analysis of the genome sequences of Bordetella pertussis, Bordetella parapertussis, and Bordetella bronchiseptica. Nat Genet. 2003;35:32–40. DOIPubMedGoogle Scholar
- Hallander H, Advani A, Riffelmann M, vonKonig CHW, Caro V, Guiso N, Bordetella pertussis strains circulating in Europe in 1999 to 2004 as determined by pulsed-field gel electrophoresis. J Clin Microbiol. 2007;45:3257–62. DOIPubMedGoogle Scholar
- Elomaa A, Advani A, Donnelly D, Antila M, Mertsola J, He Q, Population dynamics of Bordetella pertussis in Finland and Sweden, neighbouring countries with different vaccination histories. Vaccine. 2007;25:918–26. DOIPubMedGoogle Scholar
- Carbonetti NH, Artamonova GV, Van Rooijen N, Ayala VI. Pertussis toxin targets airway macrophages to promote Bordetella pertussis infection of the respiratory tract. Infect Immun. 2007;75:1713–20. DOIPubMedGoogle Scholar
- Kirimanjeswara GS, Agosto LM, Kennett MJ, Bjornstad ON, Harvill ET. Pertussis toxin inhibits neutrophil recruitment to delay antibody-mediated clearance of Bordetella pertussis. J Clin Invest. 2005;115:3594–601. DOIPubMedGoogle Scholar
- Mielcarek N, Riveau G, Remoue F, Antoine R, Capron A, Locht C. Homologous and heterologous protection after single intranasal administration of live attenuated recombinant Bordetella pertussis. Nat Biotechnol. 1998;16:454–7. DOIPubMedGoogle Scholar
- King AJ, van Gorkom T, Pennings JL, van der Heide HG, He Q, Diavatopoulos D, Comparative genomic profiling of Dutch clinical Bordetella pertussis isolates using DNA microarrays: identification of genes absent from epidemic strains. BMC Genomics. 2008;9:311. DOIPubMedGoogle Scholar
- Watanabe M, Nagai M. Whooping cough due to Bordetella parapertussis: an unresolved problem. Expert Rev Anti Infect Ther. 2004;2:447–54. DOIPubMedGoogle Scholar
- Pierce C, Klein N, Peters M. Is leukocytosis a predictor of mortality in severe pertussis infection? Intensive Care Med. 2000;26:1512–4. DOIPubMedGoogle Scholar
- Advani A, Gustafsson L, Carlsson RM, Donnelly D, Hallander HO. Clinical outcome of pertussis in Sweden: association with pulsed-field gel electrophoresis profiles and serotype. APMIS. 2007;115:736–42. DOIPubMedGoogle Scholar
- Van Buynder PG, Owen D, Vurdien JE, Andrews NJ, Matthews RC, Miller E. Bordetella pertussis surveillance in England and Wales: 1995–7. Epidemiol Infect. 1999;123:403–11. DOIPubMedGoogle Scholar
- Preziosi MP, Yam A, Wassilak SG, Chabirand L, Simaga A, Ndiaye M, Epidemiology of pertussis in a West African community before and after introduction of a widespread vaccination program. Am J Epidemiol. 2002;155:891–6. DOIPubMedGoogle Scholar
- Davison F, Nair V. Use of Marek’s disease vaccines: could they be driving the virus to increasing virulence? Expert Rev Vaccines. 2005;4:77–88. DOIPubMedGoogle Scholar
- Best SM, Kerr PJ. Coevolution of host and virus: the pathogenesis of virulent and attenuated strains of myxoma virus in resistant and susceptible European rabbits. Virology. 2000;267:36–48. DOIPubMedGoogle Scholar
- Mackinnon MJ, Read AF. Immunity promotes virulence evolution in a malaria model. PLoS Biol. 2004;2:E230. DOIPubMedGoogle Scholar
- Gandon S, Mackinnon MJ, Nee S, Read AF. Imperfect vaccines and the evolution of pathogen virulence. Nature. 2001;414:751–6. DOIPubMedGoogle Scholar
- Taranger J, Trollfors B, Bergfors E, Knutsson N, Sundh V, Lagergard T, Mass vaccination of children with pertussis toxoid – decreased incidence in both vaccinated and nonvaccinated persons. Clin Infect Dis. 2001;33:1004–10. DOIPubMedGoogle Scholar
- Luttinger P. The epidemiology of pertussi. Am J Dis Child. 1916;12:290–315.
Figures
Tables
Cite This ArticleTable of Contents – Volume 15, Number 8—August 2009
| EID Search Options |
|---|
|
|
|
|
|
|


Please use the form below to submit correspondence to the authors or contact them at the following address:
Frits R. Mooi, Laboratory for Infectious Diseases and Screening, Centre for Infectious Diseases Control, the Netherlands, National Institute for Public Health and the Environment, PO Box 1, 3720 BA Bilthoven, the Netherlands;
Top