Volume 16, Number 2—February 2010
Letter
Origin of Highly Pathogenic Porcine Reproductive and Respiratory Syndrome Virus, China
Cite This Article
Citation for Media
To the Editor: A highly pathogenic porcine reproductive and respiratory syndrome virus (HP-PRRSV), which affected >2 million pigs, emerged in early 2006 in the People’s Republic of China. The disease was characterized by high fever (41°C), high illness rates (50%–100%), and high death rates (20%–100%) for pigs of all ages (1). A number of HP-PRRSVs have been isolated from 2006 through 2009 from infected pigs in different provinces of China and confirmed to be the causative agent of the new outbreaks (1,2). These HP-PRRSVs have a deletion of 30 amino acids in the nonstructural 2 (NSP-2). However, the evolutionary origin and path of the HP-PRRSV remain unknown.
We analyzed the full-length sequences of 67 PRRSVs: 35 HP-PRRSVs (HuN4 and LNSY-08-1 isolated in our laboratory and 33 viruses isolated in other laboratories), 28 classic PRRSVs (18 viruses isolated from China and 10 viruses representing other Asian countries and North America), and 4 commercially available attenuated live PRRSV vaccine viruses. Except for the 2 viruses we isolated (HuN4 and LNSY-08-1), the full-length sequences of the other 65 viruses were obtained from GenBank. Nucleotide and deduced amino acid sequences of these PRRSVs were aligned and compared by using previous methods (3,4).
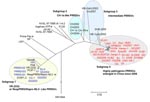
Whole genome–based phylogenetic analysis showed that these 67 PRRSVs could be divided into 4 subgroups (Appendix Figure). Ten classic PRRSVs from China, together with the North American prototype virus VR-2332 and the vaccine virus RespPRRS/Repro modified live vaccine, were classified into subgroup 1. The first Chinese isolate, CH-1a, and its 3 derivatives (CH2002, CH2003, and CH2004) were classified into subgroup 2. All 35 HP-PRRSVs were classified into subgroup 4, and they shared high homology (>99%) in their genomic sequences. The other 4 Chinese PRRSVs, including HB-1(sh)/2002, HB-2(sh)/2002, Em2007, and SHB, belonged to subgroup 3, an intermediate subgroup between subgroups 2 and 4. Phylogenetically, HP-PRRSVs had a close relationship with subgroups 2 and 3.
Four conserved deletions were shown among all HP-PRRSVs, including an adenosine deletion at position 122 in the 5′-untranslated region, a guanosine deletion at position 15,278 in the 3′-untranslated region, and 2 discontinuous deletions in the NSP-2, including a single amino acid deletion at position 482 (L482) and a second deletion of 29 amino acids between positions 533 and 561 (S533–A561). The presence of these 4 deletions among subgroup 4 viruses is a unique phenomenon, which may be used as a distinctive molecular marker for HP-PRRSVs.
The occurrence of these 4 deletions might be explained as a stepwise accumulation from subgroup 2 to subgroup 4. None of the 4 deletions were found in subgroup 2. Among viruses in subgroup 3, one, 2, or 3 of the 4 deletions occurred. For example, a single deletion was present at 122 nt in Em2007, double deletions at 122 nt and 15,278 nt in HB-1(sh)/2002 and SHB, and triple deletions at 122 nt, 15,278 nt, and 482 aa in GD3-2005 (this sequence was not submitted to GenBank until now). In 2008, Ma et al. compared GD3-2005 with several PRRSVs and reported the homology within them, pointing out that the 2 deletions in NSP-2 were identical to the HP-PRRSV (5). After careful analysis, we found the GD3-2005 more interesting than what was reported in Ma et al.; it belongs to an intermediate group, and shares the characters of gradual evolution. Eventually, all 4 deletions occurred in subgroup 4. This obvious pattern suggests that these 4 conserved deletions might have evolved step by step.
The primary neutralizing epitope (PNE), which is located on glycoprotein 5 and composed of the residues S37H(F/L)QLIYN with F/L39 as the binding site for the neutralizing antibody (6,7), also displayed similar changes at the 39 position among the 4 subgroups. The PNE residues in subgroups 1 (SHL39QLIYN) and 2 (SHF39QLIYN) were considerably conservative. Subgroup 3 contained either F39 or I39 (F39 in Em2007 and HB-2(sh)/2002, and I39 in both HB-1(sh)/2002 and SHB); subgroup 4 contained I39 only. The existence of either F39 or I39 in subgroup 3 PNE indicates its intermediate position between subgroups 2 and 4 in the evolution of HP-PRRSVs.
Pairwise comparison of subgroups 2, 3, and 4 did not find recombination or large fragment replacement, which suggests that all HP-PRRSVs originated from the same ancestor by gradual evolution. Notably, the recently isolated intermediate PRRSVs mentioned above (SHB, Em2007, and GD3-2005) were isolated in the region of South China where the outbreak of HP-PRRS initially occurred. Furthermore, the epidemiologic data show that the outbreak of HP-PRRSV emerged from 1 particular place and then spread widely. This evidence indicates that all HP-PRRSVs isolated in China likely originated from the same source.
In summary, our findings suggest that the newly emerged HP-PRRSVs originated from the Chinese CH-1a-like PRRSV. Further study is needed to determine what contributes to the increased pathogenicity of HP-PRRSV. Although the 4 deletions are conserved in all HP-PRRVs, the increased pathogenicity of HP-PRRSV may not merely be caused by the deletions; pathogenicity is affected by multigenetic factors.
Acknowledgment
This study was supported by grants from the National Basic Research Program of China (973 Program, no. 2005CB523200) and National Scientific Supporting Program of China (nos. 2006BAD06A04/03/018/01 and 2007BAD86B05/03).
References
- Tian K, Yu X, Zhao T, Feng Y, Cao Z, Wang C, Emergence of fatal PRRSV variants: unparalleled outbreaks of atypical PRRS in China and molecular dissection of the unique hallmark. PLoS One. 2007;2:e526. DOIPubMedGoogle Scholar
- Tong GZ, Zhou YJ, Hao XF, Tian ZJ, An TQ, Qiu HJ. Highly pathogenic porcine reproductive and respiratory syndrome, China. Emerg Infect Dis. 2007;13:1434–6.PubMedGoogle Scholar
- Cha SH, Choi EJ, Park JH, Yoon SR, Song JY, Kwon JH, Molecular characterization of recent Korean porcine reproductive and respiratory syndrome (PRRS) viruses and comparison to other Asian PRRS viruses. Vet Microbiol. 2006;117:248–57. DOIPubMedGoogle Scholar
- An TQ, Zhou YJ, Liu GQ, Tian ZJ, Li J, Qiu HJ, Genetic diversity and phylogenetic analysis of glycoprotein 5 of PRRSV isolates in mainland China from 1996 to 2006: coexistence of two NA-subgenotypes with great diversity. Vet Microbiol. 2007;123:43–52. DOIPubMedGoogle Scholar
- Ma P, Shi WD, Liu YG, Cai XH, Wang HF, Wang SJ, Isolation and characterization of Chinese PRRSV isolates and sequence analysis of structural genes. [in Chinese]. Progress in Veterinary Medicine. 2008;29:43–50.
- Ostrowski M, Galeota JA, Jar AM, Platt KB, Osorio FA, Lopez OJ. Identification of neutralizing and nonneutralizing epitopes in the porcine reproductive and respiratory syndrome virus GP5 ectodomain. J Virol. 2002;76:4241–50. DOIPubMedGoogle Scholar
- Plagemann PG. The primary GP5 neutralization epitope of North American isolates of porcine reproductive and respiratory syndrome virus. Vet Immunol Immunopathol. 2004;102:263–75. DOIPubMedGoogle Scholar
- Tamura K, Dudley J, Nei M, Kumar S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol. 2007;24:1596–9. DOIPubMedGoogle Scholar
Figure
Cite This ArticleRelated Links
Table of Contents – Volume 16, Number 2—February 2010
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Guang-Zhi Tong, Shanghai Veterinary Research Institute, CAAS, No. 518, Ziyue Rd, Minhang District, Shanghai 200241, People’s Republic of China
Top