Volume 17, Number 4—April 2011
Dispatch
Rapid Genotyping of Swine Influenza Viruses
Cite This Article
Citation for Media
Abstract
The emergence of pandemic (H1N1) 2009 virus highlighted the need for enhanced surveillance of swine influenza viruses. We used real-time reverse–transcription PCR–based genotyping and found that this rapid and simple genotyping method may identify reassortants derived from viruses of Eurasian avian-like, triple reassortant-like, and pandemic (H1N1) 2009 virus lineages.
Co-infection of influenza A viruses enables viral gene reassortments, thereby generating progeny viruses with novel genotypes. Such reassortants may pose a serious public health threat, as exemplified by the emergence of pandemic influenza (H1N1) in 2009 (1). Transmission of pandemic (H1N1) 2009 virus from humans to pigs has been reported (2–5). We recently identified a reassortment between pandemic (H1N1) 2009 virus and swine influenza viruses in pigs (6). These results emphasize the potential role of pigs as a mixing vessel for influenza viruses and the need for screening tests that can identify major reassortment events in pigs.
We previously developed 8 monoplex SYBR green–based quantitative reverse transcription–PCRs to detect all 8 gene segments derived from the pandemic (H1N1) 2009 virus or virus segments that are closely related to this lineage (i.e., neuraminidase [NA] and matrix protein from the Eurasian avian-like swine linage and polymerase basic protein [PB] 2, PB1, polymerase acidic protein [PA], hemagglutinin [HA], nucleocapsid protein [NP], and nonstructural protein [NS]) from triple reassortant swine linage (5). Using these PCRs, we identified swine viruses of atypical genotypes. However, with the exception of the HA-specific assay, the melting-curve signals of pandemic (H1N1) 2009 virus may be indistinguishable from the positive signals generated from its sister clade as indicated above. To differentiate between these closely related groups of viruses, we further optimized these assays by adding sequence-specific hydrolysis probes in the SYBR green assays.
For this study, all SYBR green assays were modified from the previously described assays (5), with the exception of the reverse primers for the newly designed PB1 and NS segments (Table 1). The subtype H1N1 swine influenza viruses isolated in Hong Kong during the past few years were mainly derived from the Eurasian avian-like swine lineage (6,7). To generate more precise genotyping data for our ongoing surveillance, the NA segment–specific assay was specifically designed to react with the pandemic (H1N1) 2009 virus and a portion of Eurasian avian-like swine viruses that are circulating in southeastern China (Technical Appendix Figure 1). To avoid overlapping the emission spectrum of SYBR green, we labeled all pandemic (H1N1) 2009 virus–specific hydrolysis probes (Integrated DNA Technologies, Inc., Coralville, IA, USA) with cyanine 5 (Cy5) and Black Hole Quencher-2 dyes at their 5′ and 3′ ends, respectively (Table 1). To enable use of short oligonucleotide sequences without compromising the annealing temperature of these probes, we modified the probes with locked nucleic acids (7). RNA extraction and complimentary DNA synthesis were identical to the protocols described (5,8). One microliter of 10-fold diluted complimentary DNA sample was amplified in a 20-µL reaction containing 10 µL of Fast SYBR Green Master Mix (Applied Biosystems, Foster City, CA, USA) and the corresponding primer probe set (0.5 µmol/L each). All reactions were optimized and performed simultaneously in a 7500 Sequence Detection System (Applied Biosystems) with the following conditions: 20 s at 95°C, followed by 30 cycles of 95°C for 3 s and 62°C for 30 s. SYBR green and Cy5 signals from the same reaction were captured simultaneously at the end of each amplification cycle. The expected PCR results of virus segment derived from different swine viral lineages are shown in Table 2.
The dissociation kinetics of PCR amplicons were studied by a melting curve analysis at the end of the PCR (60°C–95°C; temperature increment 0.1°C/s). We also tested various probe and SYBR green concentrations under different PCR conditions. The condition described above gave the most robust and consistent DNA amplification (data not shown). We tested 31 human pandemic (H1N1) 2009 and 63 human seasonal influenza viruses (33 subtype H1N1, 30 subtype H3N2) as controls. As expected, all human pandemic influenza viruses were double positive (i.e., positive with SYBR green and Cy5) and all seasonal influenza samples were double negative in all 8 assays.
To evaluate the sensitivity of the assays, we tested serial diluted plasmid DNA of the corresponding segments of influenza A/California/4/2009 virus as a standard. The fluorescent signals generated from the SYBR green reporter dye in all assays were highly similar to those previously reported (5), and the modified assays had a linear dynamic detection range from 102 to 108 copies/reaction (Technical AppendixFigure 2). As expected, the threshold cycle values deduced from the Cy5 reporter signal were generally higher than those from the SYBR green reporter (Technical AppendixFigure 2) (9). This finding can be partly explained by the nature of these 2 kinds of real-time PCR chemistries: a single Cy5 fluorophore of the hydrolysis probe was released from quenching for each amplicon synthesized while multiple SYBR green dyes bound to a single amplicon (10). After 35 PCR amplification cycles, the linear dynamic detection range of Cy5 signals generated from these reactions was 102 to 108 copies/reaction (data not shown). However, to avoid nonspecific SYBR green signals, we purposely limited the number of amplification cycles to 30.
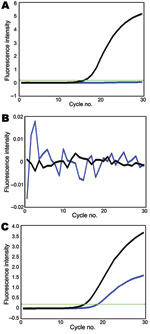
Using these assays, we tested 41 swine virus isolates collected during January 2009–January 2010. In all 8 reactions, 10 pandemic (H1N1) 2009 virus samples transmitted from humans to pigs (6) were double positive (online Technical AppendixFigure 1, pink). In these assays, gene segments of another 31 swine isolates were either SYBR green positive/Cy5 negative (Technical AppendixFigure 1, yellow) or double negative (Technical Appendix Figure 1, green), indicating that these virus segments were derived from the sister clade of pandemic (H1N1) 2009 virus or other swine lineages (except NA), respectively. For example, the reassortant of pandemic (H1N1) 2009 virus (A/swine/Hong Kong/201/2010 [H1N1]) was double positive for NA, double negative for HA and matrix protein, and SYBR green positive/Cy5 negative for PB2, PB1, PA, NP, and NS (Figure; other data not shown). All genotyping results of the studied viruses were consistent with results of previous phylogenetic analyses (5,6), indicating that our modified probes and SYBR green assays can provide more accurate genotyping results. With these genotyping data, viruses with atypical positive signal patterns might suggest a novel viral reassortment event and can be highlighted for investigation with sequencing-based methods.
To demonstrate the potential use of these assays in studying swine viruses circulating in other geographic locations, we tested 7 recent swine isolates (1 pandemic influenza subtype H1N1, 4 subtype H1N2, and 2 subtype H3N2) collected in the United States. Genotyping results agreed 100% with data deduced from sequence analyses (online Technical Appendix Table 1). We also analyzed all 436 contemporary (2008–2010) US swine virus segments available from the National Center for Biotechnology Information influenza virus sequence database. On the basis of the in silico analysis of sequences targeted by our primers and probes, 95% of the sequences (n = 413) are predicted to yield the expected results (online Technical AppendixTable 2).
The emergence of pandemic (H1N1) 2009 has highlighted the need for global systematic influenza surveillance in swine. Our results demonstrated that the addition of locked nucleic acid hydrolysis probes specific for pandemic (H1N1) 2009 virus into previously established SYBR green assays can help differentiate segments of pandemic (H1N1) 2009, Eurasian avian-like, and triple reassortant virus lineages. These assays might provide a rapid and simple genotyping method for identifying viruses that need to be fully genetically sequenced and characterized. They may also help provide better understanding of the viral reassortment events and viral dynamics in pigs. Although at present, genes derived from human seasonal viruses cannot be characterized with our modified assays, the performance of our assays warrants similar investigations for genotyping human influenza viruses.
Ms Mak is a postgraduate student in the Department of Microbiology, The University of Hong Kong. Her research focuses on molecular diagnosis of influenza virus.
Acknowledgment
This study was supported by the Area of Excellence Scheme of the University Grants Committee Hong Kong (grant AoE/M-12/06), the Research Grant Council of Hong Kong (HKU 773408M to L.L.M.P.), the Seed Funding for Basic Research (Hong Kong University), Research Fund for the Control of Infectious Disease Commissioned Project Food and Health Bureau (Hong Kong), and the National Institutes of Health (National Institute of Allergy and Infectious Diseases contract HHSN266200700005C).
References
- Novel Swine-Origin Influenza A (H1N1) Virus Investigation Team, Dawood FS, Jain S, Finelli L, Shaw MW, Lindstrom S, . Emergence of a novel swine-origin influenza A (H1N1) virus in humans. N Engl J Med. 2009;360:2605–15. DOIPubMedGoogle Scholar
- Pasma T, Joseph T. Pandemic (H1N1) 2009 infection in swine herds, Manitoba, Canada. Emerg Infect Dis. 2010;16:706–8.PubMedGoogle Scholar
- Pereda A, Cappuccio J, Quiroga MA, Baumeister E, Insarralde L, Ibar M, Pandemic (H1N1) 2009 outbreak on pig farm, Argentina. Emerg Infect Dis. 2010;16:304–7.PubMedGoogle Scholar
- Howden KJ, Brockhoff EJ, Caya FD, McLeod LJ, Lavoie M, Ing JD, An investigation into human pandemic influenza virus (H1N1) 2009 on an Alberta swine farm. Can Vet J. 2009;50:1153–61.PubMedGoogle Scholar
- Poon LL, Mak PW, Li OT, Chan KH, Cheung CL, Ma ES, Rapid detection of reassortment of pandemic (H1N1) 2009 influenza virus. Clin Chem. 2010;56:1340–4. DOIPubMedGoogle Scholar
- Vijaykrishna D, Poon LL, Zhu HC, Ma SK, Li OT, Cheung CL, Reassortment of pandemic (H1N1) 2009 influenza A virus in swine. Science. 2010;328:1529. DOIPubMedGoogle Scholar
- Johnson MP, Haupt LM, Griffiths LR. Locked nucleic acid (LNA) single nucleotide polymorphism (SNP) genotype analysis and validation using real-time PCR. Nucleic Acids Res. 2004;32:e55. DOIPubMedGoogle Scholar
- Poon LL, Chan KH, Smith GJ, Leung CS, Guan Y, Yuen KY, Molecular detection of a novel human influenza (H1N1) of pandemic potential by conventional and real-time quantitative RT-PCR assays. Clin Chem. 2009;55:1555–8. DOIPubMedGoogle Scholar
- Papin JF, Vahrson W, Dittmer DP. SYBR green–based real-time quantitative PCR assay for detection of West Nile virus circumvents false-negative results due to strain variability. J Clin Microbiol. 2004;42:1511–8. DOIPubMedGoogle Scholar
- Buh Gašparič M, Tengs T, La Paz JL, Holst-Jensen A, Pla M, Esteve T, Comparison of nine different real-time PCR chemistries for qualitative and quantitative applications in GMO detection. Anal Bioanal Chem. 2010;396:2023–9. DOIPubMedGoogle Scholar
Figure
Tables
Cite This ArticleTable of Contents – Volume 17, Number 4—April 2011
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Leo L. M. Poon, Department of Microbiology, University Pathology Building, Queen Mary Hospital, Pokfulam, Hong Kong Special Administrative Region, People’s Republic of China
Top