Volume 19, Number 8—August 2013
Dispatch
Detection of Novel Rotavirus Strain by Vaccine Postlicensure Surveillance
Cite This Article
Citation for Media
Abstract
Surveillance for rotavirus-associated diarrhea after implementation of rotavirus vaccination can assess vaccine effectiveness and identify disease-associated genotypes. During active vaccine postlicensure surveillance in the United States, we found a novel rotavirus genotype, G14P[24], in a stool sample from a child who had diarrhea. Unusual rotavirus strains may become more prevalent after vaccine implementation.
Active vaccine postlicensure surveillance for rotavirus-associated diarrhea is informative for determination of vaccine effectiveness and for characterization of disease-associated rotavirus genotypes (1–5). Most rotaviruses circulating in the United States belong to a limited number of strains, routinely characterized by serologic or genetic identification of the outer capsid protein antigens viral protein (VP) 7, which defines G types, and VP4, which defines P types (6,7). Of circulating strains in the United States, 85% contain a G or P antigen that is included in both US-licensed rotavirus vaccines (2,8). However, >70 G and P antigen combinations have been reported, and uncommon strains may suddenly appear in a new geographic area (1,2,5,9). Ongoing active surveillance is conducted through the Centers for Disease Control and Prevention’s New Vaccine Surveillance Network, a prospective, population-based surveillance program for acute gastroenteritis among children <5 years old, the details of which have been published (3–5). This surveillance has detected the emergence of G12P[8] and G9P[8] rotavirus genotypes, as well as 3 reported instances of US children infected with G8P[4] rotavirus (3–5,10). During the 2009 winter season (December 2008–June 2009) in Rochester, New York, 54 (30%) of 183 enrolled children with acute gastroenteritis had rotavirus infection. Fifty (94%) of 51 rotavirus strains were typical US strains, with G or P antigens contained in the licensed rotavirus vaccines; 3 were G8P[4] (10). One strain, however, appeared to be an unusual reassortant not previously reported in human infection. We describe this novel rotavirus genotype, G14P[24], found along with enteric adenovirus in a stool sample from a child with diarrhea.
A 36-month-old girl was brought to the emergency department of the Golisano Children’s Hospital at University of Rochester Medical Center with a 4-day history of emesis (2 times/day) and low-grade fever (37.7°C). During the previous 2 days, she also had experienced diarrhea (8 loose stools/day) and lethargy. Physical examination was only remarkable for mild dehydration; there were no other abnormalities. She was previously healthy, born after a full-term gestation, and breast-fed for the first 7–12 months of life. The child had not been vaccinated against rotavirus. She lived in the Rochester metropolitan area and had no unusual dietary or travel exposures. She had contact with pet dogs and cats at home and at a childcare setting. Approximately 1 month before her illness, she had visited a petting zoo at which farm animals but no nonhuman primates were present; her mother could not remember whether horses or cows were present but recalled the child petting sheep.
After oral rehydration, the child’s activity increased, and she was discharged to home. She continued to experience intermittent emesis and diarrhea for 1 month, although she maintained her weight. No other family members (1 sibling, 2 parents) became ill.
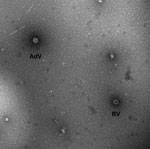
The child was enrolled, with parental informed consent, into the New Vaccine Surveillance Network. A stool sample taken during the hospital visit was positive for rotavirus antigen by enzyme immunoassay (Premier Rotaclone; Meridian Bioscience, Inc., Cincinnati, OH, USA). The specimen was analyzed at the Centers for Disease Control and Prevention by transmission electron microscopy, reverse transcription PCR genotyping, and nucleotide sequencing, as described (11). Electron microscopy showed 2 types of virions, 1 characteristic of rotavirus and 1 of enteric adenovirus (Figure 1).
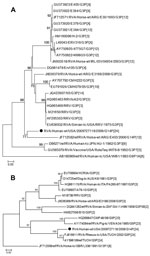
Analyses of VP7 and VP4 sequences using RotaC 2.0 (12) identified the rotavirus strain as genotype G14P[24] (6,7) (Figure 2). Phylogenetic analyses indicated monophyly of the VP7 gene with an equine rotavirus strain from Argentina and clustering of the VP4 gene with the simian rotavirus strain TUCH (Figure 2). The novel strain was designated as RVA/Human-wt/USA/2009727118/2009/G14P[24], in accordance with guidelines from the Rotavirus Classification Working Group (6,7). Full genome sequencing is in progress; the preliminary 11-gene genotype is G14-P[24]-I9-R2-C3-M3-A9-N3-T3-E3-H6, which indicates that this novel strain may be a reassortant containing genes from equine, simian, human, and bovine rotaviruses. In particular, the VP7 gene seems to be most related to equine G14 strains; the VP4 gene, to simian P[24]; the VP1 gene, to bovine R2; and the nonstructural protein 3 gene, to human T3 strains.
Human rotavirus infection is commonly associated with ≈6 of the >70 known human G and P antigen combinations reported among >160 known rotavirus strains (1,6). The G14P[24] strain we found had not been reported in human infection, but interspecies transmission of both reassorted and nonreassorted animal viruses has been described (9). The emergence of unusual reassortant animal strains raises questions about the effectiveness of current rotavirus vaccines, which might share neither G nor P types with such viruses. However, immunity to rotavirus is believed to be polygenic and probably involves antigens in addition to G and P antigens (14).
In summary, we identified infection with a novel G14P[24] rotavirus strain in a 36-month-old child with diarrhea. Whether this strain was responsible, entirely or in part, for the child’s symptoms is not certain, because enteric adenovirus was also identified. Co-infection with rotavirus and enteric adenovirus has been described, but it is unclear whether such co-infection is associated with more severe gastroenteritis (15). Nevertheless, the rotavirus strain we identified appears to be an unusual reassortant containing equine, human, simian, and bovine rotavirus genes. Further study of this and other unusual reassortant rotaviruses may lead to insight on rotavirus evolution. Continued surveillance is critical for assessing whether unusual genotypes of rotavirus become more prevalent after the implementation of rotavirus vaccination.
Dr Weinberg is a professor of pediatrics (infectious diseases) at the University of Rochester School of Medicine and Dentistry. His research interests include epidemiologic and laboratory aspects of potentially vaccine-preventable infections, especially those caused by rotavirus, parainfluenza virus, respiratory syncytial virus, and influenza virus.
Acknowledgment
We thank Charles Humphrey for performing electron microscopy; the physician, nursing, and laboratory technologist colleagues in the New Vaccine Surveillance Network; and the parents and children who enroll in New Vaccine Surveillance Network studies.
References
- Bányai K, Laszlo B, Duque J, Steele AD, Nelson EA, Gentsch JR, Systematic review of regional and temporal trends in global rotavirus strain diversity in the pre rotavirus vaccine era: insights for understanding the impact of rotavirus vaccination programs. Vaccine. 2012;30(Suppl 1):A122–30. DOIPubMedGoogle Scholar
- Gentsch JR, Hull JJ, Teel EN, Kerin TK, Freeman MM, Esona MD, G and P types of circulating rotavirus strains in the United States during 1996–2005: nine years of prevaccine data. J Infect Dis. 2009;200(Suppl 1):S99–105. DOIPubMedGoogle Scholar
- Payne DC, Staat MA, Edwards KM, Szilagyi PG, Gentsch JR, Stockman LJ, Active, population-based surveillance for severe rotavirus gastroenteritis in children in the United States. Pediatrics. 2008;122:1235–43. DOIPubMedGoogle Scholar
- Payne DC, Staat MA, Edwards KM, Szilagyi PG, Weinberg GA, Hall CB, Direct and indirect effects of rotavirus vaccination upon childhood hospitalizations in 3 US counties, 2006–2009. Clin Infect Dis. 2011;53:245–53. DOIPubMedGoogle Scholar
- Payne DC, Szilagyi PG, Staat MA, Edwards KM, Gentsch JR, Weinberg GA, Secular variation in United States rotavirus disease rates and serotypes: implications for assessing the rotavirus vaccination program. [Erratum in: Pediatr Infect Dis J. 2010;29:287–8]. Pediatr Infect Dis J. 2009;28:948–53. DOIPubMedGoogle Scholar
- Matthijnssens J, Ciarlet M, McDonald SM, Attoui H, Banyai K, Brister JR, Uniformity of rotavirus strain nomenclature proposed by the Rotavirus Classification Working Group (RCWG). Arch Virol. 2011;156:1397–413. DOIPubMedGoogle Scholar
- Matthijnssens J, Ciarlet M, Rahman M, Attoui H, Banyai K, Estes MK, Recommendations for the classification of group A rotaviruses using all 11 genomic RNA segments. Arch Virol. 2008;153:1621–9. DOIPubMedGoogle Scholar
- Cortese MM, Parashar UD. Prevention of rotavirus gastroenteritis among infants and children: recommendations of the Advisory Committee on Immunization Practices (ACIP). MMWR Recomm Rep. 2009;58(RR-2):1–25 .PubMedGoogle Scholar
- Martella V, Banyai K, Matthijnssens J, Buonavoglia C, Ciarlet M. Zoonotic aspects of rotaviruses. Vet Microbiol. 2010;140:246–55. DOIPubMedGoogle Scholar
- Weinberg GA, Payne DC, Teel EN, Mijatovic-Rustempasic S, Bowen MD, Wikswo M, First reports of human rotavirus G8P[4] gastroenteritis in the United States. J Clin Microbiol. 2012;50:1118–21. DOIPubMedGoogle Scholar
- Bányai K, Mijatovic-Rustempasic S, Hull JJ, Esona MD, Freeman MM, Frace AM, Sequencing and phylogenetic analysis of the coding region of six common rotavirus strains: evidence for intragenogroup reassortment among co-circulating G1P[8] and G2P[4] strains from the United States. J Med Virol. 2011;83:532–9. DOIPubMedGoogle Scholar
- Maes P, Matthijnssens J, Rahman M, Van Ranst M. RotaC: a web-based tool for the complete genome classification of group A rotaviruses. BMC Microbiol. 2009;9:238. DOIPubMedGoogle Scholar
- Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol. 2010;59:307–21. DOIPubMedGoogle Scholar
- Desselberger U, Huppertz HI. Immune responses to rotavirus infection and vaccination and associated correlates of protection. J Infect Dis. 2011;203:188–95. DOIPubMedGoogle Scholar
- Román E, Wilhelmi I, Colomina J, Villar J, Cilleruelo ML, Nebreda V, Acute viral gastroenteritis: proportion and clinical relevance of multiple infections in Spanish children. J Med Microbiol. 2003;52:435–40. DOIPubMedGoogle Scholar
Figures
Cite This ArticleTable of Contents – Volume 19, Number 8—August 2013
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Geoffrey A. Weinberg, Department of Pediatrics, University of Rochester School of Medicine and Dentistry, 601 Elmwood Ave, Box 690, Rochester, NY 14642, USA
Top