Volume 22, Number 3—March 2016
Letter
Generalized Cowpox Virus Infection in a Patient with HIV, Germany, 2012
Figure
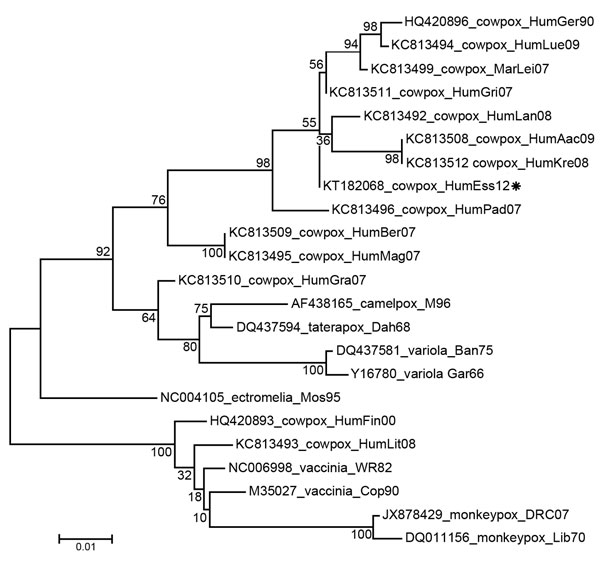
Figure. Evolutionary relationships of cowpox virus isolated from a 35-year-old man with HIV infection treated in the intensive care unit at the University of Duisburg–Essen, Essen, Germany (KT182068_HumEss12, asterisk), other human isolates of cowpox virus, and other orthopoxviruses. The evolutionary history was inferred by using the maximum-likelihood method. The percentage of trees in which the associated taxa clustered together is shown next to the branch nodes. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. The analysis involved 23 nt sequences (GenBank accession numbers are indicated). All positions containing gaps and missing data were eliminated; the final dataset had 772 positions. Evolutionary analyses were conducted in MEGA6 (http://www.megasoftware.net). Aac, Aachen; Ban, Bangladesh; Ber, Berlin; Cop, Copenhagen; Dah, Dahomey; DRC, Democratic Republic of Congo; Ess, Essen; Fin, Finland; Gar, Garcia; Ger, Germering; Gra, Graz; Gri, Grimmen; Hum, human; Kre, Krefeld; Lan, Landau; Lei, Leipzig; Lib, Liberia; Lit, Lithuania; Lue, Luedenscheid; Mag, Magdeburg; Mar, Patagonian mara (Dolichotis patagonum); Mos, Moscow; Pad, Paderborn; WR, Western Reserve. Scale bar indicates the number of nucleotide changes per site.