Volume 22, Number 4—April 2016
Letter
Enterovirus A71 Genogroups C and E in Children with Acute Flaccid Paralysis, West Africa
Cite This Article
Citation for Media
To the Editor: Human enterovirus 71 (EV-A71) of the species enterovirus A, genus Enterovirus, and family Picornaviridae is a serious public health threat because it can cause large outbreaks of hand, foot and mouth disease (HFMD). In addition, EV-A71 has been associated with severe and sometimes fatal neurologic complications that affect mostly infants and children and that range from aseptic meningitis and encephalitis to poliomyelitis-like acute flaccid paralysis (AFP) (1). EV-A71 has been classified into 7 genogroups, A–G, on the basis of the diversity of the nucleotide sequences of the viral protein 1 (VP1) capsid (1,2). Since 1997, increasing epidemic activity of genogroups B and C has been reported in the Asia-Pacific region and has caused large HFMD outbreaks with high rates of illness and death. Subgenogroup C2 was identified as the main causal agent associated with cases of fatal encephalitis in the devastating HFMD outbreaks of Taiwan in 1998 and Australia in 1999 (1). Since those outbreaks, C2 has been frequently reported in countries in Asia, Europe, North America, and South America and has been associated with neurologic complications and fatal infection (1). Although genogroups E and F were recently discovered in Africa, the epidemiology of EV-A71 has been largely unexplored in this continent. Only 4 AFP cases associated with EV-A71 infection have been reported in Africa (2–5), and the only outbreak caused by EV-A71 occurred in Kenya in 2000 among a small number of HIV-infected orphans and was attributed to genogroup C (6).
To investigate the circulation and genetic diversity of EV-A71 in West Africa, we retrospectively analyzed 236 nonpolio enterovirus (NPEV) isolates obtained through routine poliomyelitis surveillance activities at the World Health Organization’s Regional Polio Laboratory in Senegal during 2013–2014. Following WHO guidelines (7), the laboratory received and processed stool specimens from 1,600 children with AFP from various West Africa countries. NPEV was found in isolates from most countries except Cape Verde (0/5 specimens): Gambia (8/64), Guinea-Bissau (8/49), Guinea (42/355), Mauritania (20/108), Niger (95/596), and Senegal (63/423). NPEV was initially characterized by amplification of the VP1 capsid protein coding region by using reverse transcription PCR and partial sequencing, as described (8). After molecular typing of all NPEV isolates, we identified 4 new EV-A71 isolates from 4 patients, 1 each from Guinea, Mauritania, Niger, and Senegal (Technical Appendix Table 1). BLAST analysis (http://www.ncbi.nlm.nih.gov/) of the partial VP1 nucleotide sequences showed that the isolates from this study shared 96%–97% nt sequence identity with those of other EV-A71 isolates deposited in GenBank.
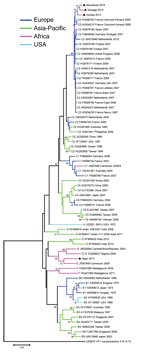
The complete VP1 nucleotide sequences of the 4 EV-A71 isolates were determined by using reverse transcription PCR (GenBank accession nos. KT818793–KT818796; Technical Appendix Table 2). Sequences were aligned by using ClustalW (http://www.clustal.org). Phylogenetic investigation indicated that the EV-A71 strains detected in 3 of the 4 isolates (from Senegal, Guinea, and Mauritania) clustered within subgenogroup C2. The most closely related EV-A71 strains are not those previously reported from other Africa countries (Madagascar, Cameroon, Nigeria, and Central African Republic) but are strains found in isolates from France, Finland, United Kingdom, Spain, Germany and Netherlands, all isolated during 2006–2010 (Figure). The C2 clusters from Africa and Europe showed an average of 97.1% nt identity and 99.8% aa similarity. Within the Africa C2 cluster, the nucleotide and amino acid alignments displayed a substantial proportion of conserved positions: 862/891 (96.7%) and 297/297 (100%), respectively. Because most enteroviruses evolve at the rate of ≈1% nt substitutions per year in the VP1 region, the sequence divergence of ≈3% suggests that these enteroviruses had been circulating in the region for ≈3 years before they were detected. However, we cannot exclude the possibility that these isolates originated from multiple importation events from abroad, especially from countries in Europe. The fourth isolate (from Niger) clustered within genogroup E, which, before this study, included only 2 complete VP1 sequences from isolates from Central African Republic and Cameroon and an additional partial VP1 sequence from an isolate from Nigeria.
Comparison of complete VP1 amino acid sequences of all EV-A71 strains considered for the phylogenetic analysis showed that 107 (36%) of 297 aa sites were variable. None of the residues found in the variable sites of the Africa strains in our study corresponded to residues previously associated with genogroup C neurovirulent phenotypes (A170V, N31D, L97R, G145E and D164E) (9,10). The Niger E isolate showed specific residues (L24M, A170T) that differed from those of other genogroup E isolates.
Our findings highlight the presence of EV-A71 with a high degree of genetic diversity in patients with AFP in West Africa. Future studies about the burden of disease, epidemiologic features, and evolution of EV-A71 in this region of Africa are needed to implement appropriate public health measures.
Acknowledgments
We are grateful to Romain Volle and Florence Colbère-Garapin for critical review of the manuscript and to Hiroyuki Shimizu for fruitful discussions.
This work was supported by the Pasteur Institute’s Transverse Research Programs PTR-276 and PTR-484.
References
- Solomon T, Lewthwaite P, Perera D, Cardosa MJ, McMinn P, Ooi MH. Virology, epidemiology, pathogenesis, and control of enterovirus 71. Lancet Infect Dis. 2010;10:778–90. DOIPubMedGoogle Scholar
- Bessaud M, Razafindratsimandresy R, Nougairede A, Joffret ML, Deshpande JM, Dubot-Peres A, Molecular comparison and evolutionary analyses of VP1 nucleotide sequences of new African human enterovirus 71 isolates reveal a wide genetic diversity. PLoS One. 2014;9:e90624. DOIPubMedGoogle Scholar
- Bessaud M, Pillet S, Ibrahim W, Joffret ML, Pozzetto B, Delpeyroux F, Molecular characterization of human enteroviruses in the Central African Republic: uncovering wide diversity and identification of a new human enterovirus A71 genogroup. J Clin Microbiol. 2012;50:1650–8. DOIPubMedGoogle Scholar
- Oyero OG, Adu FD. Non-polio enteroviruses serotypes circulating in Nigeria. Afr J Med Sci. 2010;39(Suppl):201–8.PubMedGoogle Scholar
- Sadeuh-Mba SA, Bessaud M, Massenet D, Joffret ML, Endegue MC, Njouom R, High frequency and diversity of species C enteroviruses in Cameroon and neighboring countries. J Clin Microbiol. 2013;51:759–70. DOIPubMedGoogle Scholar
- Chakraborty R, Iturriza-Gomara M, Musoke R, Palakudy T, D’Agostino A, Gray J. An epidemic of enterovirus 71 infection among HIV-1-infected orphans in Nairobi. AIDS. 2004;18:1968–70. DOIPubMedGoogle Scholar
- World Health Organization. Polio laboratory manual. 4th ed. 2004 Aug 31 [cited 2015 Jan 30]. http://apps.who.int/iris/bitstream/10665/68762/1/WHO_IVB_04.10.pdf
- Bessaud M, Jegouic S, Joffret ML, Barge C, Balanant J, Gouandjika-Vasilache I, Characterization of the genome of human enteroviruses: design of generic primers for amplification and sequencing of different regions of the viral genome. J Virol Methods. 2008;149:277–84. DOIPubMedGoogle Scholar
- Zhang B, Wu X, Huang K, Li L, Zheng L, Wan C, The variations of VP1 protein might be associated with nervous system symptoms caused by enterovirus 71 infection. BMC Infect Dis. 2014;14:243. DOIPubMedGoogle Scholar
- Kataoka C, Suzuki T, Kotani O, Iwata-Yoshikawa N, Nagata N, Ami Y, The role of VP1 amino acid residue 145 of enterovirus 71 in viral fitness and pathogenesis in a cynomolgus monkey model. PLoS Pathog. 2015;11:e1005033. DOIPubMedGoogle Scholar
Figure
Cite This ArticleRelated Links
Table of Contents – Volume 22, Number 4—April 2016
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Maria D. Fernandez-Garcia, Pôle de Virologie, Institut Pasteur, 36 Avenue Pasteur, BP220, Dakar, Senegal
Top