Volume 23, Number 11—November 2017
Research Letter
Unrecognized Dengue Virus Infections in Children, Western Kenya, 2014–2015
Cite This Article
Citation for Media
Abstract
We detected a cluster of dengue virus infections in children in Kenya during July 2014–June 2015. Most cases were serotype 1, but we detected all 4 serotypes, including co-infections with 2 serotypes. Our findings implicate dengue as a cause of febrile illness in this population and highlight a need for robust arbovirus surveillance.
Due to lack of national surveillance programs, the extent of infection with dengue viruses (DENV) among children is largely unknown in much of sub-Saharan Africa (1). Uncovering this hidden burden is critical for making informed public health decisions that affect populations that are most vulnerable to vectorborne disease. To address this knowledge gap, in January 2014, we initiated a 4.5-year study of the transmission and extent of arbovirus infection in children in Kenya. Although the study continues through June 2018, we identified a cluster of DENV infections among febrile children in western Kenya, prompting this report to raise awareness of DENV as a cause of acute febrile illness (AFI) among children in this country.
We describe results from a cohort of children with AFI who came to 1 of 2 regional health centers that serve the communities of Chulaimbo (a rural village) and Kisumu (an urban city), both located in western Kenya. The cohort is part of a larger ongoing parent study that enrolls additional child cohorts at other study sites or with other study designs and collects vector and environmental data. Details of the parent study are beyond the scope of this report and will be described elsewhere. The study is being conducted under the supervision of the institutional review board of Stanford University (Stanford, CA, USA; IRB-31488) and the scientific and ethics review unit of the Kenya Medical Research Institute (Kisumu, Kenya; SSC 2611).
For this study, we enrolled febrile children 1–17 years of age. At the enrollment and 1-month follow-up visits, we collected information on demographics and risk factors, performed a physical examination, and obtained blood samples. We tested the blood samples for DENV RNA by reverse transcription PCR (RT-PCR) (2).
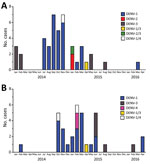
We enrolled 1,258 children with AFI during January 2014–April 2016. We tested blood samples from 1,104 study participants (87.8%) by RT-PCR. Of those, 82 (7.4%) were positive for DENV RNA: 58 (70%) were serotype 1 (DENV-1), 2 (2.4%) were serotype 2 (DENV-2), 13 (15.9%) were serotype 3 (DENV-3), and 2 (2.4%) were serotype 4 (DENV-4) (Figure). We also detected co-infection with 2 serotypes in 6 participants: 2 children had DENV-1 and -3; 3 children had DENV-1 and -4; and 1 child had DENV-2 and -3.
Most DENV cases occurred during July 2014–June 2015. The likelihood of testing positive for DENV by PCR did not differ by age or sex. The median age of DENV PCR-positive participants was 3.5 years (IQR 2.0–5.2 years); 43.9% of participants were female. We did observe a higher likelihood of testing positive by PCR for children from the rural site, Chulaimbo (9.6%), compared with those from the urban site, Kisumu (5.8%) (p = 0.017 by Fisher exact test; odds ratio 1.97, 95% CI 1.14–3.43, adjusted for age and sex). Continued surveillance is needed to investigate whether DENV is endemic to these areas and if there are differences in regional endemicity.
Although reports of DENV infection in Kenya remain sparse, several sero-epidemiologic studies have produced evidence of DENV transmission by demonstrating serum DENV IgG in patients of all ages (3–6). In 2013, an outbreak of DENV-2 in Mombasa was detected, in part, by RT-PCR (7). More recently, DENV infections were described in 43 adult patients with fever who resided in Mtwapa, on the coast of Kenya near Mombasa; these cases occurred February 2014–January 2015 (8), which overlapped with the period in which we observed the DENV cases we report in this study. All of the DENV PCR-positive subjects from the 2016 study were infected with DENV-2, which is consistent with the DENV serotype observed in the 2013 Mombasa outbreak (7). In contrast, most DENV PCR-positive subjects from our study were infected with DENV-1, which suggests regional variation in DENV strain predominance.
DENV viremia during acute infection is known to be transient (9); thus, our definition of DENV cases, based on RT-PCR detection of RNA, probably underestimated the true incidence of DENV infection. Data from testing of serum samples for DENV IgM and IgG seroconversion were not available as we wrote this report but would provide confirmation of our PCR results by a second method of detecting DENV infection. IgM testing, although less specific than PCR (10), may identify additional cases of DENV infection and could alter the descriptive statistics we present by enhancing our sensitivity for detecting DENV infection.
Improving efforts to detect DENV infections will raise awareness of DENV and increase the likelihood that healthcare providers will suspect it in patients with AFI. Correct diagnoses improve the ability of public health ministries to detect and react to outbreaks of DENV or other arboviral illnesses, rather than contributing to cycles of missed opportunities for preventive interventions.
Dr. Vu is an instructor of pediatrics at Stanford University School of Medicine, conducting arbovirus epidemiology research in the laboratory of A.D.L. His primary research interests center on host responses to dengue and malaria infections.
Acknowledgments
We thank our study participants.
This project was made possible by National Institutes of Health grant R01-AI102918 to A.D.L. Additionally, support was provided by a grant from the Stanford Child Health Research Institute & Lucille Packard Foundation for Children’s Health to D.M.V. This work was conducted with support from a KL2 Mentored Career Development Award to D.M.V. from Spectrum, supported by the Stanford Clinical and Translational Science Award (NIH KL2 TR 001083 and UL TR 001085).
References
- Amarasinghe A, Kuritsk JN, Letson GW, Margolis HS. Dengue virus infection in Africa. Emerg Infect Dis. 2011;17:1349–54.PubMedGoogle Scholar
- Lanciotti RS, Calisher CH, Gubler DJ, Chang GJ, Vorndam AV. Rapid detection and typing of dengue viruses from clinical samples by using reverse transcriptase-polymerase chain reaction. J Clin Microbiol. 1992;30:545–51.PubMedGoogle Scholar
- Johnson BK, Ocheng D, Gichogo A, Okiro M, Libondo D, Kinyanjui P, et al. Epidemic dengue fever caused by dengue type 2 virus in Kenya: preliminary results of human virological and serological studies. East Afr Med J. 1982;59:781–4.PubMedGoogle Scholar
- Mease LE, Coldren RL, Musila LA, Prosser T, Ogolla F, Ofula VO, et al. Seroprevalence and distribution of arboviral infections among rural Kenyan adults: a cross-sectional study. Virol J. 2011;8:371.PubMedGoogle Scholar
- Sutherland LJ, Cash AA, Huang YJ, Sang RC, Malhotra I, Moormann AM, et al. Serologic evidence of arboviral infections among humans in Kenya. Am J Trop Med Hyg. 2011;85:158–61.PubMedGoogle Scholar
- Vu DM, Banda T, Teng CY, Heimbaugh C, Muchiri EM, Mungai PL, et al. Dengue and West Nile virus transmission in children and adults in coastal Kenya. Am J Trop Med Hyg. 2017;96:141–3.PubMedGoogle Scholar
- Ellis EM, Neatherlin JC, Delorey M, Ochieng M, Mohamed AH, Mogeni DO, et al. A household serosurvey to estimate the magnitude of a dengue outbreak in Mombasa, Kenya, 2013. PLoS Negl Trop Dis. 2015;9:e0003733.PubMedGoogle Scholar
- Ngoi CN, Price MA, Fields B, Bonventure J, Ochieng C, Mwashigadi G, et al. Dengue and chikungunya virus infections among young febrile adults evaluated for acute HIV-1 infection in coastal Kenya. PLoS One. 2016;11:e0167508.PubMedGoogle Scholar
- Murgue B, Roche C, Chungue E, Deparis X. Prospective study of the duration and magnitude of viraemia in children hospitalised during the 1996-1997 dengue-2 outbreak in French Polynesia. J Med Virol. 2000;60:432–8.PubMedGoogle Scholar
- de Oliveira Poersch C, Pavoni DP, Queiroz MH, de Borba L, Goldenberg S, dos Santos CN, et al. Dengue virus infections: comparison of methods for diagnosing the acute disease. J Clin Virol. 2005;32:272–7.PubMedGoogle Scholar
Figure
Cite This ArticleTable of Contents – Volume 23, Number 11—November 2017
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
David M. Vu, Stanford University School of Medicine, Pediatric Infectious Diseases, 300 Pasteur Dr, Rm G312, Stanford, CA 94305, USA
Top