Volume 23, Number 3—March 2017
Research Letter
Potentially Zoonotic Bartonella in Bats from France and Spain
Cite This Article
Citation for Media
Abstract
We detected Bartonella in 11 of 109 insectivorous bats from France and 1 of 26 bats from Spain. These genetic variants are closely related to bat-associated Bartonella described in Finland and the United Kingdom and to B. mayotimonensis, the agent of a human endocarditis case in the United States.
Bartonellae have been identified in bats sampled in locations around the world where diverse chiropteran host species can interact with numerous Bartonella variants and potential arthropod vectors (1–3). Many Bartonella species are zoonotic, potentially affecting human and bat health (4). Bartonella spp. in bat populations of Europe are of particular interest because some variants described in Finland and the United Kingdom are closely related to Bartonella mayotimonensis, a species detected in the resected aortic valve of a 59-year-old endocarditis patient in the United States (5,6). To determine if potentially zoonotic bat-associated bartonellae are circulating elsewhere in Europe, we tested insectivorous bats from France and Spain for the presence of Bartonella spp.
We performed necropsies on 26 bats from Spain and 109 from France to collect heart tissue for Bartonella spp. diagnostics (Technical Appendix Table 1). Bats from Spain were originally collected during active surveillance for rabies at the Unidad de Aislamiento y Detección Virus, Instituto de Salud Carlos III, Madrid, Spain. Of the bats from France, 97 were originally submitted for passive rabies surveillance to the Agence Nationale de Sécurité Sanitaire de l’Alimentation,de l’Environnement et du Travail (ANSES), Laboratoire de la Rage et de la Faune Sauvage de Nancy in Malzéville, France. We sampled the remaining 12 animals from a rehabilitation center at the Musée d’Histoire Naturelle in Bourges, France. All bats collected for Bartonella diagnostics tested negative for rabies. Spatial coordinates were recorded for all bats at the point of sampling before submission for centralized laboratory testing (online Technical Appendix Figure). Whenever possible, we identified bat species and sex. Methods were approved by the University of California, Davis (Davis, CA, USA), Institutional Animal Care and Use Committee (protocol 17669).
We used sampling records to determine the genus and species for 118 of the 135 bats; identification data were not available for 17 of 26 bats from Spain. The 118 identified bats belonged to 8 genera and at least 13 different species: 70 Pipistrellus spp. (31 P. pipistrellus, 24 P. nathusii, 6 P. kuhlii, 4 P. pigmaeus, and 5 Pipistrellus spp.); 15 Nyctalus noctula; 7 Eptesicus serotinus; 11 Myotis spp. (4 M. mystacinus, 3 M. daubentonii, 1 M. bechsteinii, 1 M. myotis, 1 M. nattereri, 1 M emarginatus); 6 Plecotus spp. (4 P. austriacus and 2 P. auritus); 6 Tadarida teniotis; 2 Barbastella barbastellus; and 1 Vespertilio murinus.
We used NucleoSpin Blood QuickPure kits (Machery-Nagel, Düren, Germany) to extract DNA from 25 mg of macerated heart tissue according to the manufacturer’s instructions. We used tissue spiked with B. henselae as a positive control for DNA extraction. We screened samples by PCR targeting the citrate synthase gene (gltA). Primers CSH1f (GCGAATGAAGCGTGCCTAAA) and BhCS1137.n (AATGCAAAAAGAACAGTAAACA) amplified an ≈350-bp fragment suitable for distinguishing Bartonella species (7). PCR thermal cycler parameters were set at 10 min at 95°C, followed by 40 cycles of 30 s at 94°C, 1 min at 57°C, 2 min at 72°C, 5 mins at 75°C, and infinite hold at 4°C. We verified amplicon sizes by gel electrophoresis, and Service de Séquençage de Eurofins (Paris, France) generated sequence data from PCR products.
We used OpenEpi version 3.01 (http://www.openepi.com/) to calculate descriptive statistics and CIs for prevalence data. We constructed phylogenetic trees by using the MrBayes plugin in Geneious version 8.1.7 with a Markov Chain Monte Carlo value of 1,100,000 and 100,000 burn-in length (8). We used the ggplot2 package in R (https://www.r-project.org/) to create spatial maps.
We detected Bartonella DNA in 12 (8.9%) of 135 bat heart tissue samples (Technical Appendix Table 2); 11 of the tissues were from bats from France, and 1 was from an unidentified bat captured in Torreferrusa, Catalonia, Spain. The 11 Bartonella-positive bats from France belonged to only 4 of the 13 sampled species: N. noctula (2/15 bats [13.3%, 95% CI 1.7%–40.5%]), P. nathusii (6/24 bats [25%, 95% CI 9.8%–46.7%]), M. daubentonii (2/3 bats [66.6%, 95% CI 9.4%–99.1%]), and M. mystacinus (1/4 bats [25%, 95% CI 0.6%–80.6%]).
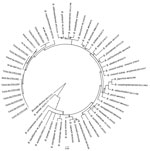
All 12 Bartonella variants (GenBank accession nos. KY041981–KY041992) clustered closely with zoonotic B. mayotimonensis (Figure). Two sequences obtained from M. daubentonii bats sampled in Lorraine (GenBank accession no. KY041985) and Upper Normandy (GenBank accession no. KY041989), France, shared 100% nt identity with Bartonella strains previously isolated from bats of the same species in Finland and the United Kingdom (5,9). None of the Bartonella variants were closely related to Candidatus Bartonella naantaliensis or Candidatus Bartonella hemsundetiensis, which were also described in bats sampled in Finland (5,10). The absence of variants resembling these bartonellae from northern Europe suggests a spatial heterogeneity in the distribution of Bartonella spp. across bat populations and selective adaptations to specific host reservoirs.
Further research is needed to better evaluate the prevalence of zoonotic Bartonella species in western Europe and to determine if B. mayotimonensis, the agent of a US case of human endocarditis, is present across a broader range than currently documented. Future studies should consider specifically focusing on Nyctalus, Pipistrellus, and Myotis bat species, from which we most frequently detected variants similar to B. mayotimonensis.
Dr. Stuckey works in the Department of Population Health and Reproduction at the University of California Davis School of Veterinary Medicine. His research interests include disease ecology and epidemiology of zoonoses.
Acknowledgments
We thank Laurent Arthur, Michèle Lemaire, Alvaro Aguilar Setién, Noël Tordo, Janet Foley, Deana Clifford, Rickie Kasten, Philip Kass, Daniel Greenia, Tristan Burgess, José M. Berciano, Annick Suzor-Weiner, Nadia Haddad, Martine Monteil, Elisabeth Petit, Thibaud Dugat, and Jean-Philippe Buffet for help with the study design, bat sampling, laboratory diagnostics, manuscript preparation, and general guidance. We also thank the Société Française pour l’Étude et la Protection des Mammifères (SFEPM) group for their participation in the passive bat rabies surveillance in France.
M.J.S. was funded by a Chateaubriand STEM (Science, Technology, Engineering & Mathematics) Fellowship (French Ministry of Foreign Affairs) with matching funds from Mérial, Lyon, France.
References
- McKee CD, Hayman DT, Kosoy MY, Webb CT. Phylogenetic and geographic patterns of Bartonella host shifts among bat species. Infect Genet Evol. 2016;44:382–94. DOIPubMedGoogle Scholar
- Lei BR, Olival KJ. Contrasting patterns in mammal–bacteria coevolution: Bartonella and Leptospira in bats and rodents. PLoS Negl Trop Dis. 2014;8:e2738 Pubmed DOIGoogle Scholar
- Judson SD, Frank HK, Hadly EA. Bartonellae are prevalent and diverse in Costa Rican bats and bat flies. Zoonoses Public Health. 2015;62:609–17. DOIPubMedGoogle Scholar
- Mühldorfer K. Bats and bacterial pathogens: a review. Zoonoses Public Health. 2013;60:93–103. DOIPubMedGoogle Scholar
- Veikkolainen V, Vesterinen EJ, Lilley TM, Pulliainen AT. Bats as reservoir hosts of human bacterial pathogen, Bartonella mayotimonensis. Emerg Infect Dis. 2014;20:960–7. DOIPubMedGoogle Scholar
- Lin EY, Tsigrelis C, Baddour LM, Lepidi H, Rolain JM, Patel R, et al. Candidatus Bartonella mayotimonensis and endocarditis. Emerg Infect Dis. 2010;16:500–3. DOIPubMedGoogle Scholar
- Kamani J, Baneth G, Mitchell M, Mumcuoglu KY, Gutiérrez R, Harrus S. Bartonella species in bats (Chiroptera) and bat flies (Nycteribiidae) from Nigeria, West Africa. Vector Borne Zoonotic Dis. 2014;14:625–32. DOIPubMedGoogle Scholar
- Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012;28:1647–9.
- Concannon R, Wynn-Owen K, Simpson VR, Birtles RJ. Molecular characterization of haemoparasites infecting bats (Microchiroptera) in Cornwall, UK. Parasitology. 2005;131:489–96. DOIPubMedGoogle Scholar
- Lilley TM, Veikkolainen V, Pulliainen AT. Molecular detection of Candidatus Bartonella hemsundetiensis in bats. Vector Borne Zoonotic Dis. 2015;15:706–8. DOIPubMedGoogle Scholar
Figure
Cite This ArticleTable of Contents – Volume 23, Number 3—March 2017
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Address for correspondence:Bruno B. Chomel, VM3B Room 1020, 1089 Veterinary Medicine Dr, University of California, Davis, CA 95616, USA
Top