Volume 24, Number 11—November 2018
Dispatch
Detection and Characterization of Human Pegivirus 2, Vietnam
Figure
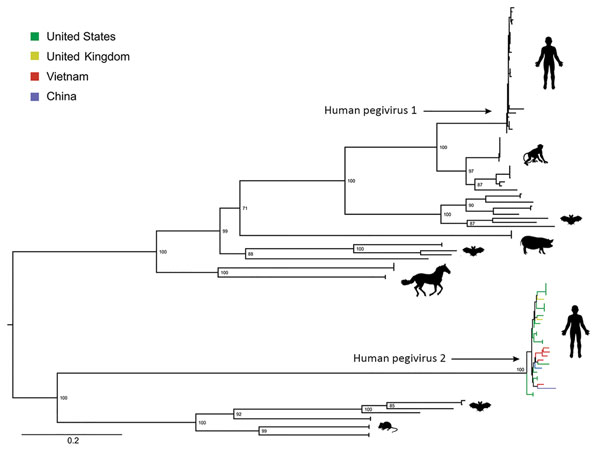
Figure. Maximum-likelihood phylogenetic tree of amino acid sequences of coding sequences of human pegivirus 2 strains from Vietnam compared with global strains and other pegiviruses. We used the general matrix with empirical amino acid frequencies, a gamma distribution of 4 rates, and invariant sites, as suggested by IQ TREE (http://www.iqtree.org), to reconstruct the phylogenetic trees. We assessed support for individual nodes using a bootstrap procedure of 10,000 replicates. Scale bar indicates amino acid substitutions per site.
1Members of the Southeast Asia Infectious Disease Clinical Research Network are listed at the end of this article.
Page created: October 17, 2018
Page updated: October 17, 2018
Page reviewed: October 17, 2018
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.