Volume 25, Number 5—May 2019
CME ACTIVITY - Research
Novel Sequence Type in Bacillus cereus Strains Associated with Nosocomial Infections and Bacteremia, Japan
Figure 3
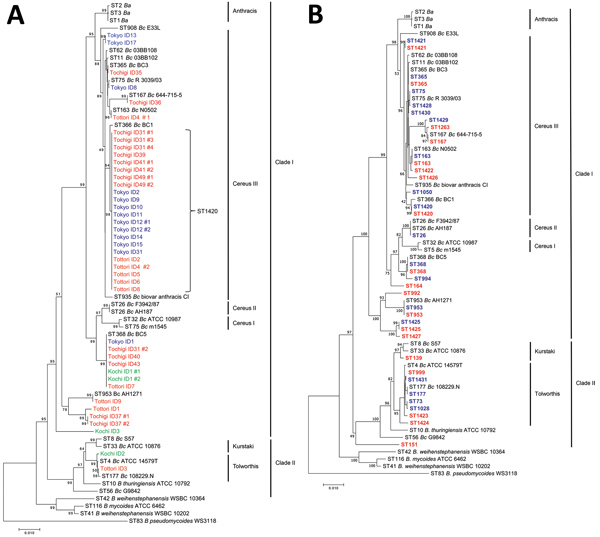
Figure 3. Multilocus sequence typing (MLST)–based phylogenetic trees of strains and STs of Bacillus cereus isolates, Japan. Reference sequences were obtained from the MLST database (https://pubmlst.org). Definitions of clades and lineage names followed those of Priest et al. (9). A) Phylogenetic tree of isolates from patients with bacteremia. Blue indicates Tokyo strains, red indicates Tochigi stains, orange indicates Tottori strains, and green indicates Kochi strains. B) Phylogenetic tree of STs detected in Tokyo and Tochigi strains. Blue indicates STs detected in Tokyo strains, and red indicates STs detected in Tochigi strains. Scale bars indicates nucleotide substitutions per site. Ba, B. anthracis; Bc, B. cereus; ID, identification; ST, sequence type.
References
- Bottone EJ. Bacillus cereus, a volatile human pathogen. Clin Microbiol Rev. 2010;23:382–98. DOIPubMedGoogle Scholar
- Schaefer G, Campbell W, Jenks J, Beesley C, Katsivas T, Hoffmaster A, et al. Persistent Bacillus cereus bacteremia in 3 persons who inject drugs, San Diego, California, USA. Emerg Infect Dis. 2016;22:1621–3. DOIPubMedGoogle Scholar
- Thomas BS, Bankowski MJ, Lau WK. Native valve Bacillus cereus endocarditis in a non-intravenous-drug-abusing patient. J Clin Microbiol. 2012;50:519–21. DOIPubMedGoogle Scholar
- Marley EF, Saini NK, Venkatraman C, Orenstein JM. Fatal Bacillus cereus meningoencephalitis in an adult with acute myelogenous leukemia. South Med J. 1995;88:969–72. DOIPubMedGoogle Scholar
- Hoffmaster AR, Ravel J, Rasko DA, Chapman GD, Chute MD, Marston CK, et al. Identification of anthrax toxin genes in a Bacillus cereus associated with an illness resembling inhalation anthrax. Proc Natl Acad Sci U S A. 2004;101:8449–54. DOIPubMedGoogle Scholar
- Bryce EA, Smith JA, Tweeddale M, Andruschak BJ, Maxwell MR. Dissemination of Bacillus cereus in an intensive care unit. Infect Control Hosp Epidemiol. 1993;14:459–62. DOIPubMedGoogle Scholar
- Hernaiz C, Picardo A, Alos JI, Gomez-Garces JL. Nosocomial bacteremia and catheter infection by Bacillus cereus in an immunocompetent patient. Clin Microbiol Infect. 2003;9:973–5. DOIPubMedGoogle Scholar
- Barrie D, Hoffman PN, Wilson JA, Kramer JM. Contamination of hospital linen by Bacillus cereus. Epidemiol Infect. 1994;113:297–306. DOIPubMedGoogle Scholar
- Priest FG, Barker M, Baillie LW, Holmes EC, Maiden MC. Population structure and evolution of the Bacillus cereus group. J Bacteriol. 2004;186:7959–70. DOIPubMedGoogle Scholar
- Zwick ME, Joseph SJ, Didelot X, Chen PE, Bishop-Lilly KA, Stewart AC, et al. Genomic characterization of the Bacillus cereus sensu lato species: backdrop to the evolution of Bacillus anthracis. Genome Res. 2012;22:1512–24. DOIPubMedGoogle Scholar
- Hoffmaster AR, Novak RT, Marston CK, Gee JE, Helsel L, Pruckler JM, et al. Genetic diversity of clinical isolates of Bacillus cereus using multilocus sequence typing. BMC Microbiol. 2008;8:191. DOIPubMedGoogle Scholar
- Zhang J, van Hung P, Hayashi M, Yoshida S, Ohkusu K, Ezaki T. DnaJ sequences of Bacillus cereus strains isolated from outbreaks of hospital infection are highly similar to Bacillus anthracis. Diagn Microbiol Infect Dis. 2011;70:307–15. DOIPubMedGoogle Scholar
- Dohmae S, Okubo T, Higuchi W, Takano T, Isobe H, Baranovich T, et al. Bacillus cereus nosocomial infection from reused towels in Japan. J Hosp Infect. 2008;69:361–7. DOIPubMedGoogle Scholar
- Sasahara T, Hayashi S, Morisawa Y, Sakihama T, Yoshimura A, Hirai Y. Bacillus cereus bacteremia outbreak due to contaminated hospital linens. Eur J Clin Microbiol Infect Dis. 2011;30:219–26. DOIPubMedGoogle Scholar
- McDougal LK, Steward CD, Killgore GE, Chaitram JM, McAllister SK, Tenover FC. Pulsed-field gel electrophoresis typing of oxacillin-resistant Staphylococcus aureus isolates from the United States: establishing a national database. J Clin Microbiol. 2003;41:5113–20. DOIPubMedGoogle Scholar
- Fluit AC, Terlingen AM, Andriessen L, Ikawaty R, van Mansfeld R, Top J, et al. Evaluation of the DiversiLab system for detection of hospital outbreaks of infections by different bacterial species. J Clin Microbiol. 2010;48:3979–89. DOIPubMedGoogle Scholar
- Kajitani R, Toshimoto K, Noguchi H, Toyoda A, Ogura Y, Okuno M, et al. Efficient de novo assembly of highly heterozygous genomes from whole-genome shotgun short reads. Genome Res. 2014;24:1384–95. DOIPubMedGoogle Scholar
- Larsen MV, Cosentino S, Rasmussen S, Friis C, Hasman H, Marvig RL, et al. Multilocus sequence typing of total-genome-sequenced bacteria. J Clin Microbiol. 2012;50:1355–61. DOIPubMedGoogle Scholar
- Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol. 2016;33:1870–4. DOIPubMedGoogle Scholar
- Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4:406–25.PubMedGoogle Scholar
- Ogawa H, Fujikura D, Ohnuma M, Ohnishi N, Hang’ombe BM, Mimuro H, et al. A novel multiplex PCR discriminates Bacillus anthracis and its genetically related strains from other Bacillus cereus group species. PLoS One. 2015;10:e0122004. DOIPubMedGoogle Scholar
- Ramisse V, Patra G, Garrigue H, Guesdon JL, Mock M. Identification and characterization of Bacillus anthracis by multiplex PCR analysis of sequences on plasmids pXO1 and pXO2 and chromosomal DNA. FEMS Microbiol Lett. 1996;145:9–16. DOIPubMedGoogle Scholar
- Ramisse V, Patra G, Vaissaire J, Mock M. The Ba813 chromosomal DNA sequence effectively traces the whole Bacillus anthracis community. J Appl Microbiol. 1999;87:224–8. DOIPubMedGoogle Scholar
- Vassileva M, Torii K, Oshimoto M, Okamoto A, Agata N, Yamada K, et al. Phylogenetic analysis of Bacillus cereus isolates from severe systemic infections using multilocus sequence typing scheme. Microbiol Immunol. 2006;50:743–9. DOIPubMedGoogle Scholar
1Current affiliation: Asahikawa Medical University, Asahikawa, Japan.
2Current affiliation: Japanese Foundation for Cancer Research, Tokyo, Japan