Volume 25, Number 7—July 2019
Research Letter
Outbreak of African Swine Fever, Vietnam, 2019
Figure
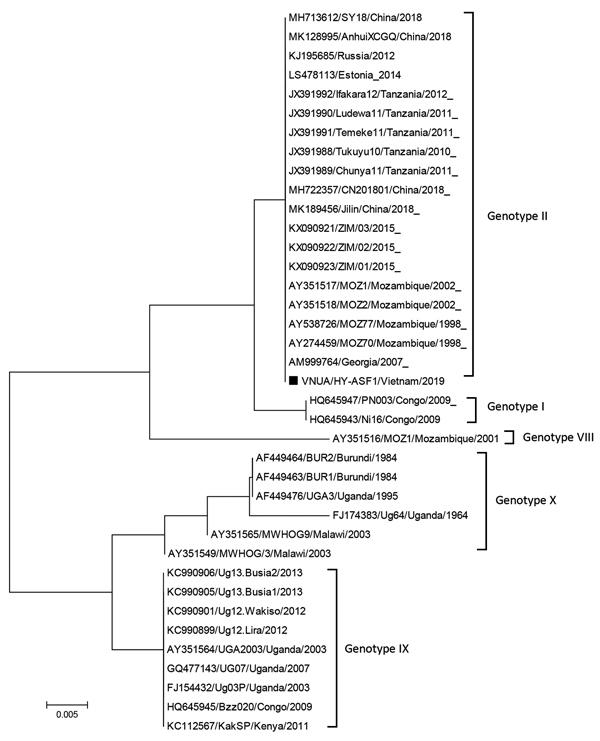
Figure. Phylogenetic analysis of major capsid protein gene (p72) of African swine fever virus isolated during outbreak in Vietnam in 2019 (VNUA HY-ASF1; black square) and reference isolates. The phylogenetic tree was constructed by using the neighbor-joining method in MEGA7 (http://www.megasoftware.net). Bootstrap values were calculated with 1,000 replicates. GenBank accession numbers, strain name, country, and year of collection are indicated. Scale bars indicate nucleotide substitutions per site.
1These authors contributed equally to this article.
Page created: June 17, 2019
Page updated: June 17, 2019
Page reviewed: June 17, 2019
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.