Volume 26, Number 12—December 2020
Dispatch
Range Expansion of Bombali Virus in Mops condylurus Bats, Kenya, 2019
Cite This Article
Citation for Media
Abstract
Previously identified only in Sierra Leone, Guinea, and southeastern Kenya, Bombali virus–infected Mops condylurus bats were recently found »750 km away in western Kenya. This finding supports the role of M. condylurus bats as hosts and the potential for Bombali virus circulation across the bats’ range in sub-Saharan Africa.
Bombali virus (BOMV) is the sixth and most recently identified virus of the genus Ebolavirus (1), first detected in Sierra Leone in oral and rectal swab samples from 2 species of insectivorous bats, Mops condylurus and Chaerephon pumilus (2). Since then, BOMV has been found in the tissues and excreta of M. condylurus bats in southeastern Kenya (3) and Guinea (4). To explore the role of M. condylurus bats as hosts for BOMV and the geographic distribution of the virus, we trapped bats in western Kenya, screened tissues for BOMV, and conducted next-generation sequencing on positive samples.
Bats were trapped in mist nets at 4 sites in Busia County: 2 house roosts, 1 orchard, and 1 cave. A total of 182 bats were captured, including 113 M. condylurus and 18 C. pumilus (Table 1). Similarly, at the original location in the Taita Hills, bats were trapped at a bridge site where an infected bat had previously been identified (3), at 4 additional building roosts, and over a water hole. From these sites, 396 bats were captured, including 177 M. condylurus and 219 C. pumilus (Table 1). Captured bats were euthanized with terminal isoflurane anesthesia followed by cervical dislocation. We collected mouth swab samples, fecal and blood samples, and major organs (kidney, spleen, liver, intestine, lung, and brain) and stored them in RNAlater (Invitrogen, https://www.thermofisher.com) as described previously (3).
Samples were stored at −20°C for up to 10 days in Kenya before being shipped to Helsinki, Finland, where they were stored at −70°C before processing in a Biosafety Level 3 laboratory. Tissue samples were treated with TRIzol (Invitrogen) for virus inactivation, and RNA was extracted according to the manufacturer’s instructions. Because previous studies have identified the highest BOMV viral loads in bat lungs (3,4), we initially conducted reverse transcription PCR (RT-PCR) on pooled lung samples from 3 bats (same species, collection date, and location) by using the BOMV-specific RT-PCR protocol described earlier (2). Samples in positive pools were then screened individually, and other sample types (other organs, saliva, and excreta) from these bats were also tested.
We conducted next-generation sequencing on positive lung samples. Before sequencing, we applied a multiplex PCR protocol for amplification. The primers for the entire BOMV coding region were designed by using the PrimalScheme tool (5) with a target amplicon of 500 bp with 50-bp overlap. Complementary DNA was synthesized from RNA samples positive by RT-PCR by using SuperScript III enzyme (Invitrogen) and random hexamers; PCR was conducted by using a Q5 PCR kit (New England Biolabs, https://www.neb.com) (5). The PCR products were purified by using AMPure XP magnetic beads (Beckman Coulter, https://www.beckmancoulter.com), and sequencing libraries were prepared by using a Nextera XT kit (Illumina) according to the manufacturer’s instructions. Sequencing was conducted with the Illumina MiSeq Reagent Kit v2, a sequencing kit with 150-bp paired-end reads. The raw sequence reads were trimmed by using Trimmomatic (Q-score >30, read length >50 bp) and assembled to the reference sequence (MK340750) by using the Burrows-Wheeler Aligner–Maximal Exact Match algorithm implemented in SAMTools version 1 ([6], H. Li et al. unpub. data, https://arxiv.org/abs/1303.3997?upload=1).
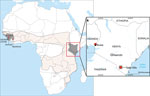
We identified 3 BOMV RNA–positive M. condylurus bats (Z153, Z178, and X030). All other bats were negative. Two of the BOMV-positive bats were captured in Busia (Z153 and Z178, both adult males) in 2 distinct trapping locations (both house roosts) »7 km apart (Figure 1). One BOMV-positive bat (X030, gravid female) was captured in the Taita Hills at the location previously reported to have a BOMV-infected bat (3). Among the bat samples, BOMV RNA was present in lung (3/3), spleen (2/3), mouth (1/3), liver (1/3), and fecal (1/2) samples but absent from all kidney, intestine, and brain samples (Table 2). Of note, BOMV RNA was not consistently detected from the mouth swab or fecal samples of tissue-positive individuals. Viral loads were quantified as previously described (3); the highest viral loads were detected in the lungs (both male bats) and the spleen (female bat).
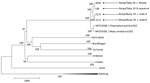
By sequencing, we obtained 2 full genomes (from X030 and Z153, GenBank accession nos. MW056492 and MW056493) and 1 partial genome (from Z178, GenBank accession no. MW056494). Phylogenetic analysis showed 99% nt identity between the complete sequences from Busia and Taita (200-nt difference) and 97% nt identity with the prototype strain from Sierra Leone (Figure 2). The virus sequence obtained from a bat at the Taita Hills in this study was almost identical to the sequence from a bat at the same site in 2018 (4-nt difference).
We identified 2 BOMV-positive M. condylurus bats from separate roost sites in western Kenya, expanding the known virus geographic range in East Africa. This new location in Busia County, on the Kenya–Uganda border, is »750 km from the previously reported site. We also identified a second BOMV-positive M. condylurus bat at the original site near the Taita Hills in southeastern Kenya, indicating sustained local persistence but limited prevalence.
Lack of evidence of infection in bats of other species, including C. pumilus, indicates a dominant host role for M. condylurus compared with other species. Although no evidence of human infection with BOMV has yet been reported (3), the close phylogenetic relationship of BOMV to pathogenic human ebolaviruses necessitates prudence. Because M. condylurus bats are one of the several synanthropic species of bat often found roosting in human-made structures (e.g., dwellings, schools, offices, and bridges in rural and urban areas [8,9]), the potential for human exposure to M. condylurus bats and the pathogens they carry is likely to be higher than exposure to bats of many other species. However, study of M. condylurus bats has been relatively limited, and other aspects of human exposure risk, such as bat movement range, are inferred from closely related species; M. condylurus bats are thought to travel only short distances in line with their feeding activities (8).
Our results expand the known distribution of BOMV and increase support for a role of M. condylurus bats as hosts. Although BOMV had been previously reported in 3 distinct locations, most evidence came from Sierra Leone and adjacent Guinea (2,4). Kenya is located on the opposite side of Africa, where only 1 BOMV-positive bat had been identified (3). These new findings from 2 disparate locations in Kenya demonstrate the established presence of BOMV in East Africa and the potential for BOMV circulation across the M. condylurus bat range in sub-Saharan Africa (10). The low virus prevalence observed in M. condylurus bats (1.7% in Busia, 0.6% in Taita), however, is below that for reservoir hosts in other bat pathogen systems (e.g., Marburg virus [11]). Therefore, questions remain as to how the virus is maintained within and transmitted among bat colonies and whether bats of other taxa are involved.
Dr. Kareinen is a veterinarian and a PhD student at the University of Helsinki. His research interests include the maintenance and circulation of zoonotic viruses in bat hosts.
Acknowledgments
We thank Ali Mirazimi and Samir Abdurahman for their assistance with analyzing the mouth swab samples and Ruut Uusitalo for preparing the map image.
This research was supported by the Finnish Cultural Foundation, the Jenny and Antti Wihuri Foundation, the Academy of Finland (grant no. 318726), Helsinki University Hospital Funds, and the Jane and Aatos Erkko Foundation. Bat trapping and sample collections were conducted under permits from the National Commission for Science, Technology and Innovation (permit no. NACOSTI/P/18/76501/22243) and the Kenya Wildlife Service (permit no. KWS/BRM/500). Sample import to Finland was approved by the Finnish Food Safety Authority (EVIRA; 4250/0460/2016 and 2809/0460/2018).
References
- Olival KJ, Hayman DT. Filoviruses in bats: current knowledge and future directions. Viruses. 2014;6:1759–88. DOIPubMedGoogle Scholar
- Goldstein T, Anthony SJ, Gbakima A, Bird BH, Bangura J, Tremeau-Bravard A, et al. The discovery of Bombali virus adds further support for bats as hosts of ebolaviruses. Nat Microbiol. 2018;3:1084–9. DOIPubMedGoogle Scholar
- Forbes KM, Webala PW, Jääskeläinen AJ, Abdurahman S, Ogola J, Masika MM, et al. Bombali virus in Mops condylurus bat, Kenya. Emerg Infect Dis. 2019;25:955–7. DOIPubMedGoogle Scholar
- Karan LS, Makenov MT, Korneev MG, Sacko N, Boumbaly S, Yakovlev SA, et al. Bombali virus in Mops condylurus bats, Guinea. Emerg Infect Dis. 2019;25:1774–5. DOIPubMedGoogle Scholar
- Quick J, Grubaugh ND, Pullan ST, Claro IM, Smith AD, Gangavarapu K, et al. Multiplex PCR method for MinION and Illumina sequencing of Zika and other virus genomes directly from clinical samples. Nat Protoc. 2017;12:1261–76. DOIPubMedGoogle Scholar
- Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–20. DOIPubMedGoogle Scholar
- Kumar S, Stecher G, Tamura K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol. 2016;33:1870–4. DOIPubMedGoogle Scholar
- Noer CL, Dabelsteen T, Bohmann K, Monadjem A. Molossid bats in an African agro-ecosystem select sugarcane fields as foraging habitat. Afr Zool. 2012;47:1–11. DOIGoogle Scholar
- Bronrier GN, Maloney SK, Buffenstein R. Survival tactics within thermally-challenging roosts: heat tolerance and cold sensitivity in the Angolan free-tailed bat, Mops condylurus. S Afr Zool. 1999;34:1–10.
- Happold M, Happold D. Tadarida condylura Angolan free-tailed bat. In: Happold M, Happold D, editors. Mammals of Africa. Vol. 4. London: Bloomsbury Publishing; 2013. p. 505–7.
- Amman BR, Carroll SA, Reed ZD, Sealy TK, Balinandi S, Swanepoel R, et al. Seasonal pulses of Marburg virus circulation in juvenile Rousettus aegyptiacus bats coincide with periods of increased risk of human infection. PLoS Pathog. 2012;8:
e1002877 . DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleOriginal Publication Date: November 16, 2020
Table of Contents – Volume 26, Number 12—December 2020
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Tarja Sironen, University of Helsinki, Department of Medicine, Haartmaninkatu 3, FI-00290, Helsinki, Finland
Top