Volume 27, Number 12—December 2021
Research Letter
SARS-CoV-2 B.1.1.7 Variant Infection in Malayan Tigers, Virginia, USA
Figure
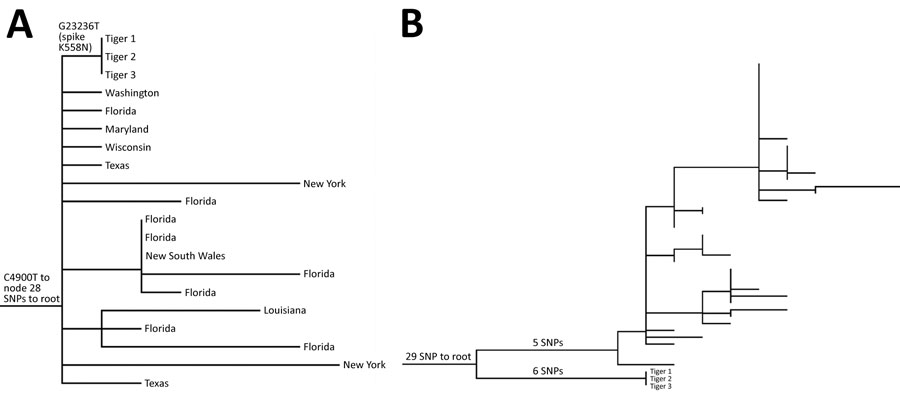
Figure. Maximum-likelihood phylogenetic trees of severe acute respiratory syndrome coronavirus 2 from 3 Malayan tigers, Virginia, USA. Tiger samples are numbered in order of symptom onset. A) Subset of phylogenetic tree showing parent (G23236T) and grandparent (C4900T) nodes of the tiger sequences, with tips labeled as states of origin in the United States or Australia. B) Phylogenetic tree showing that other B.1.1.7 viruses detected in Virginia that contain the K558N mutation are not epidemiologically related to the sequences detected in tigers 1, 2, and 3. SNP, single-nucleotide polymorphism.
Page created: September 01, 2021
Page updated: November 19, 2021
Page reviewed: November 19, 2021
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.