Volume 27, Number 6—June 2021
Dispatch
Molecular Characterization and Antimicrobial Resistance in Neisseria gonorrhoeae, Nunavut Region of Inuit Nunangat, Canada, 2018–2019
Figure 2
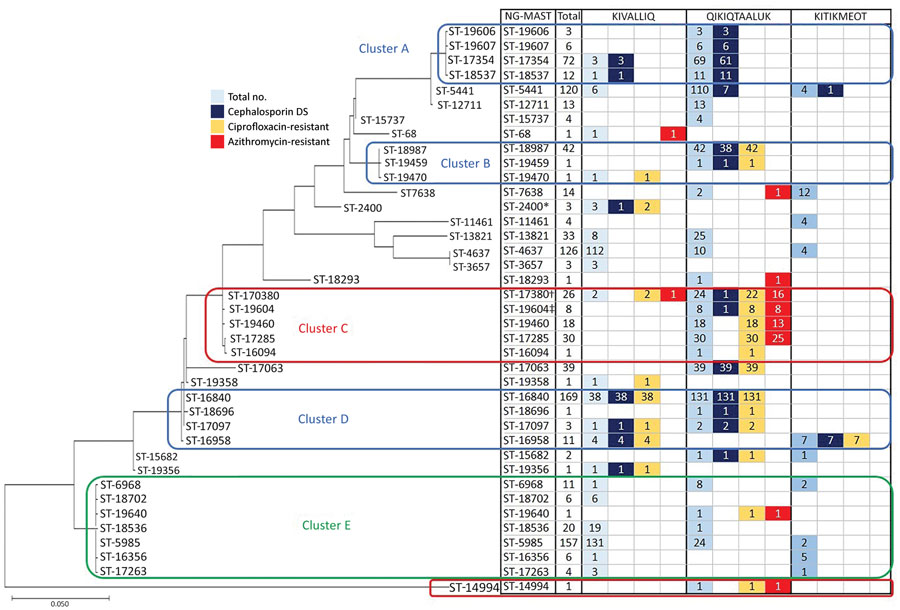
Figure 2. Genetic relationship of Neisseria gonorrhoeae multiantigen sequence typing sequence types (STs) of gonorrhea-positive nucleic acid amplification specimens with prevalent and predicted nonsusceptible SNP assay results (n = 975) from a study of antimicrobial resistance in N. gonorrhoeae in the Nunavut region of Inuit Nunangat, Canada, 2018–2019. Only prevalent STs or STs whose samples predicted decreased susceptibility to cephalosporins or resistance to ciprofloxacin or azithromycin were included. The evolutionary history was inferred by using the maximum-likelihood method and Tamura-Nei model (13). The tree with the highest log likelihood (–4321.41) is shown. Initial trees for the heuristic search were obtained automatically by applying neighbor-joining and BioNJ algorithms to a matrix of pairwise distances estimated using the maximum composite likelihood approach, and then selecting the topology with superior log likelihood value. The tree is drawn to scale, with branch lengths measured in the number of substitutions per site. This analysis involved 39 nucleotide sequences. Codon positions included were 1st+2nd+3rd+Noncoding; a total of 917 positions were in the final dataset. Evolutionary analyses were conducted in MEGA X (14). Clusters were identified as STs with ≤5 base pair differences between them. *One of the ST2400 samples was predicted to be cephalosporin decreased susceptibility (not cephalosporin/DS). †The sample that was predicted to be cephalosporin/DS was not azithromycin resistant. ‡The sample that was cephalosporin/DS was also azithromycin resistant. Cephalosporin/DS, cephalosporin intermediate/decreased susceptibility.
References
- Choudhri Y, Miller J, Sandhu J, Leon A, Aho J. Gonorrhea in Canada, 2010-2015. Can Commun Dis Rep. 2018;44:37–42. DOIPubMedGoogle Scholar
- Unemo M, Seifert HS, Hook EW III, Hawkes S, Ndowa F, Dillon JR. Gonorrhoea. Nat Rev Dis Primers. 2019;5:79. DOIPubMedGoogle Scholar
- World Health Organization, Department of Reproductive Health and Research. Global action plan to control the spread and impact of antimicrobial resistance in Neisseria gonorrhoeae. 2012 [cited 2021 Apr 21]. https://www.who.int/reproductivehealth/publications/rtis/9789241503501/en
- Public Health Agency of Canada, National Microbiology Laboratory. 2020. National surveillance of antimicrobial susceptibilities of Neisseria gonorrhoeae annual summary 2018 [cited 2021 Apr 21]. https://www.canada.ca/en/public-health/services/publications/drugs-health-products/national-surveillance-antimicrobial-susceptibilities-neisseria-gonorrhoeae-annual-summary-2018.html
- Statistics Canada. Population and dwelling count highlight tables, 2016 census [cited 2021 Apr 21]. https://www12.statcan.gc.ca/census-recensement/2016/dp-pd/hlt-fst/pd-pl/Table.cfm?Lang=Eng&T=101&SR=1&S=3&O=D#tPopDwell
- Nunavut Tunngavik Incorporated. Annual report on the state of Inuit culture and society, 2007–8 [cited 2021 Apr 21]. https://www.tunngavik.com/publications/annual-report-on-the-state-of-inuit-culture-and-society-2007-2008
- Kanatami IT. National Inuit strategy on research, 2018 [cited 2021 Apr 21]. https://www.itk.ca/wp-content/uploads/2018/04/ITK_NISR-Report_English_low_res.pdf
- Martin IM, Ison CA, Aanensen DM, Fenton KA, Spratt BG. Rapid sequence-based identification of gonococcal transmission clusters in a large metropolitan area. J Infect Dis. 2004;189:1497–505. DOIPubMedGoogle Scholar
- Peterson S, Martin I, Demczuk W, Barairo N, Naidu P, Lefebvre B, et al. Multiplex real-time polymerase chain reaction assays for the prediction of cephalosporin, ciprofloxacin, and azithromycin antimicrobial susceptibility of positive Neisseria gonorrhoeae nucleic acid amplification test samples. J Antimicrob Chemother. 2020;75:3485–90. DOIPubMedGoogle Scholar
- Trembizki E, Buckley C, Donovan B, Chen M, Guy R, Kaldor J, et al. Direct real-time PCR-based detection of Neisseria gonorrhoeae 23S rRNA mutations associated with azithromycin resistance. J Antimicrob Chemother. 2015;70:3244–9.PubMedGoogle Scholar
- Tamura K, Nei M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol Biol Evol. 1993;10:512–26.PubMedGoogle Scholar
- Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018;35:1547–9. DOIPubMedGoogle Scholar
- Gonorrhea public health protocol. Nunavut communicable disease and surveillance manual. Iqaluit (Canada): Nunavut Department of Health. 2018.
- Didelot X, Dordel J, Whittles LK, Collins C, Bilek N, Bishop CJ, et al. Genomic analysis and comparison of two gonorrhea outbreaks. MBio. 2016;7:e00525–16. DOIPubMedGoogle Scholar
- Ota KV, Fisman DN, Tamari IE, Smieja M, Ng L-K, Jones KE, et al. Incidence and treatment outcomes of pharyngeal Neisseria gonorrhoeae and Chlamydia trachomatis infections in men who have sex with men: a 13-year retrospective cohort study. Clin Infect Dis. 2009;48:1237–43. DOIPubMedGoogle Scholar