Volume 27, Number 6—June 2021
Dispatch
Brucellosis Outbreak Traced to Commercially Sold Camel Milk through Whole-Genome Sequencing, Israel
Cite This Article
Citation for Media
Abstract
Brucellosis, a neglected zoonotic disease acquired from contaminated food products, remains a public health concern worldwide. We describe an outbreak in which commercially sold camel milk containing Brucella melitensis was distributed across Israel. Whole-genome sequencing linked patients infected with B. melitensis to wholesale camel milk and unregulated livestock trade.
Brucellosis, caused by bacteria of the Brucella genus, is a neglected zoonotic disease that affects marginalized populations worldwide (1). Human transmission occurs mainly through consumption of unpasteurized, contaminated dairy products from infected, domesticated animals. In Israel, brucellosis primarily affects Arab populations, especially the seminomadic Bedouin tribal communities in southern Israel (2).
Dromedary camels, which are capable of asymptomatic carriage of Brucella, are valuable domesticated animals in Bedouin culture, and unpasteurized camel milk has gained international popularity because of its alleged medicinal properties. Few reported brucellosis outbreak investigations involving camel milk have described families infected by milk from privately owned camels and have used serologic testing to confirm diagnoses (3,4). The standard tool for molecular typing of Brucella has been multilocus variable-number tandem-repeat analysis or multilocus sequence typing (MLST). However, whole-genome sequencing (WGS) is increasingly used for the study of genomic epidemiology of B. melitensis (5).
We report an outbreak of human brucellosis with unique epidemiologic characteristics, which originated from commercial, single-brand, unpasteurized camel milk; infection was diagnosed in 19 patients with a common exposure history over 4 months. We demonstrate the utility of WGS for brucellosis outbreak investigations.
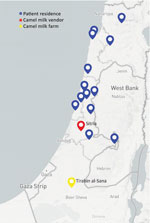
From July–November 2016, the Israeli Ministry of Health noted an increase in brucellosis cases in non-Arab patients in central and northern Israel, raising suspicion of a common source (Figure 1). An epidemiologic investigation noted patients were exposed to the same brand of camel milk. A total of 20 isolates were obtained from 19 patients across Israel (nos. 1–20; Table). Patients from shared households included 2 pairs of siblings and 1 married couple. We studied 2 isolates (nos. 1, 2) from an infant from whom B. melitensis was isolated from blood and cerebrospinal fluid. We also included 1 person (isolate no. 12) who consumed camel milk of an unknown brand.
The suspected vendor obtained milk from a Bedouin camel farm in southern Israel (Figure 1). Field investigation of the farm revealed 32 female and 2 male camels. A total of 4 female camels had positive serologic test results for Brucella, but none were available for further testing. We sampled 6 bottles of camel milk obtained from a natural food store carrying the suspected brand and recovered a few colonies of B. melitensis from 3 of the bottles.
Clinical isolates were submitted to the National Brucellosis Reference Laboratory (Kimron Veterinary Institute, Beit Dagan, Israel) for confirmation, and milk samples were cultured at the same laboratory. In total, 10 B. melitensis human isolates from the outbreak and 3 camel milk isolates (nos. 21–23) from the implicated source were available for sequencing. An additional 4 epidemiologically unrelated isolates were sequenced and used as outliers, including 1 isolate from camel milk (no. 24), 2 isolates from patients with B. melitensis with no camel milk exposure (nos. 26, 27), and 1 isolate from an unrelated patient with B. melitensis acquired after consuming camel milk (no. 25). Finally, we used a B. melitensis reference genome sequence from the National Center for Biotechnology Information Sequence Read Archive database (no. 28).
DNA was extracted from Brucella isolates by heat killing (80 C°, 10 min) and by using the DNeasy Blood & Tissue kit (QIAGEN, https://www.qiagen.com). Genomic libraries were prepared with a Nextera Flex kit (Illumina, https://www.illumina.com) and subjected to paired-end sequencing by using the Illumina Miseq or Nextseq platforms. Sequences have been deposited in the European Nucleotide Archive (BioProject PRJEB43660).
We used WGS to analyze a total of 18 isolates: 10 human isolates, 3 camel milk isolates, 4 outlier isolates, and 1 reference sequence. Raw sequences underwent quality control using fastQC version 0.11.8 (6). We assembled sequence reads that passed QC by the pipeline shovill version 1.0.4 with Trimmomatic version 0.39, SPAdes version 3.13.1, and Pilon version 1.23 with the parameters “–trim –opts –sc” (7). We implemented in silico MLST with MLST version 2.10 per the pubMLST Brucella spp. Scheme (8,9). We conducted core genome single-nucleotide polymorphism (SNP) analysis of the WGS assemblies by using ekidna version 0.3.2 (10). The sequence from isolate 2 was selected for reference with complete assembly of 3,297,563 bases (Appendix Table 1). We masked resulting core genome SNPs for recombination events using Gubbins version 2.3.4 (11) and visualized the final SNP alignment as a minimum-spanning tree created by PHYLOViZ2 (12).
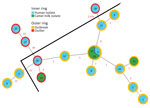
We assigned all isolates to sequence type 8 by using in silico MLST. Core genome SNP analysis found 2,443 SNP sites with the sequence from isolate 2 for reference. The phylogenetic tree demonstrated 1 main cluster including 8 human outbreak isolates and all 3 isolates from camel milk (Figure 2). Two human outbreak isolates (nos. 4, 9) did not belong to the main cluster, suggesting >1 clone might have caused the outbreak. Isolate 12 clustered with outbreak isolates, confirming that the case-patient who consumed an unknown brand of camel milk was part of the outbreak. Outlier sequences (nos. 25, 27, 28) were genetically distinct from outbreak isolates, expectedly. However, outliers (nos. 26, 24) clustered with outbreak isolates, suggesting an unrecognized epidemiologic link between Bedouin camel farming and outbreak source.
We describe an outbreak of brucellosis unique in its patient population, source of infection, chain of transmission, and use of WGS. Prior outbreak investigations of brucellosis have seldom used WGS to link the outbreak with samples from the implicated source. Here, WGS exposed genetic linkage between bottled camel milk and 8 isolates from 7 patients, providing conclusive evidence of this common source. Isolates obtained from the same person (nos. 1, 2) and that person’s sibling (no. 3) clustered tightly, providing internal validity to the discriminative power of WGS. WGS also associated isolate no. 12 (unknown camel milk brand) with the outbreak.
Two isolates (nos. 4, 9) from cases clustered apart from the outbreak strains. Because 4 female camels at the implicated farm had positive Brucella serologic test results, they might have harbored several different strains of B. melitensis, which could explain these results. We acquired all bottles of camel milk sampled during a single day, representing just 1 batch, and other batches were not tested. Therefore, the presence of additional clones implicated in this outbreak could not be ascertained.
Surprisingly, isolates 24 and 26 clustered with the outbreak strains, suggesting an unrecognized epidemiologic chain of transmission. This finding could reflect patterns of unregulated animal trade in Israel in which domesticated animals, including camels, are trafficked from Hebron (origin of isolate 26) throughout the Negev region to Bedouin communities (in this case Rahat [origin of isolate 24], the largest Bedouin city). Clustering of these outliers suggests that the outbreak might have been more widespread than documented.
In contrast with traditional patterns of brucellosis cases and case clusters, which occur mainly in small communities where residents consume locally produced unpasteurized dairy products, this outbreak resulted from web-based commercial sales of an unregulated food product. This sales channel enabled the spread of B. melitensis throughout Israel, similar to a recent United States report on commercially sold contaminated raw cow milk (13). Therefore, modern consumer trends in small agriculture and food production may result in new food safety risks.
The advantages of WGS in this outbreak investigation included the capacity to analyze both related and outlier isolates. This capability enabled us to resolve complex linkages between cases, suggesting unequivocal, temporal relationships that depict outbreak directionality and can aid future preventative measures.
In conclusion, we describe a unique B. melitensis outbreak linking human cases with commercially sold, unregulated camel milk. These results demonstrate that WGS can be a powerful tool for investigating transmission of brucellosis, a neglected zoonotic disease, in a world of modernized commercialism.
Dr. Svetlana Bardenstein is a veterinary doctor specializing in clinical microbiology and currently heading the World Organisation for Animal Health/Food and Agriculture Organization of the United Nations Brucella Reference Laboratory in Israel. Her main interest is the epidemiology of brucellosis in Israel.
Acknowledgments
We would like to thank all personnel of the Ministry of Health and Ministry of Agriculture and Rural Development who contributed to the investigation of the outbreak and incident management.
This work was supported by a grant from the Ministry of Agriculture and Rural Development (#33-08-0009).
References
- Pappas G, Papadimitriou P, Akritidis N, Christou L, Tsianos EV. The new global map of human brucellosis. Lancet Infect Dis. 2006;6:91–9. DOIPubMedGoogle Scholar
- Anis E, Leventhal A, Grotto I, Gandacu D, Warshavsky B, Shimshony A, et al. Recent trends in human brucellosis in Israel. Isr Med Assoc J. 2011;13:359–62.PubMedGoogle Scholar
- Shimol SB, Dukhan L, Belmaker I, Bardenstein S, Sibirsky D, Barrett C, et al. Human brucellosis outbreak acquired through camel milk ingestion in southern Israel. Isr Med Assoc J. 2012;14:475–8.PubMedGoogle Scholar
- Garcell HG, Garcia EG, Pueyo PV, Martín IR, Arias AV, Alfonso Serrano RN. Outbreaks of brucellosis related to the consumption of unpasteurized camel milk. J Infect Public Health. 2016;9:523–7. DOIPubMedGoogle Scholar
- Tan K-K, Tan Y-C, Chang L-Y, Lee KW, Nore SS, Yee W-Y, et al. Full genome SNP-based phylogenetic analysis reveals the origin and global spread of Brucella melitensis. BMC Genomics. 2015;16:93. DOIPubMedGoogle Scholar
- Andrews S. FastQC: a quality control tool for high throughput sequence data. 2018 [cited 2021 Apr 4]. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
- Seeman T, Kwong J, Gladman S, Goncalves da Silva A. Shovil. Faster SPAdes assembly of Illumina reads. [cited 2021 Apr 4]. https://github.com/tseemann/shovill
- Jolley KA, Bray JE, Maiden MCJ. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018;3:124. DOIPubMedGoogle Scholar
- Croucher NJ, Page AJ, Connor TR, Delaney AJ, Keane JA, Bentley SD, et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 2015;43:e15–15. DOIPubMedGoogle Scholar
- Nascimento M, Sousa A, Ramirez M, Francisco AP, Carriço JA, Vaz C. PHYLOViZ 2.0: providing scalable data integration and visualization for multiple phylogenetic inference methods. Bioinformatics. 2017;33:128–9. DOIPubMedGoogle Scholar
- Gruber JF, Newman A, Egan C, Campbell C, Garafalo K, Wolfgang DR, et al. Notes from the field: Brucella abortus RB51 infections associated with consumption of raw milk from Pennsylvania—2017 and 2018. MMWR Morb Mortal Wkly Rep. 2020;69:482–3. DOIPubMedGoogle Scholar
Figures
Cite This ArticleOriginal Publication Date: May 10, 2021
Table of Contents – Volume 27, Number 6—June 2021
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Jacob Moran-Gilad, Dept. of Health Systems Management, Faculty of Health Sciences, Ben Gurion University of the Negev, POB 653, Beer Sheva 8410501, Israel
Top