Volume 28, Number 12—December 2022
Dispatch
Highly Diverse Arenaviruses in Neotropical Bats, Brazil
Cite This Article
Citation for Media
Abstract
We detected arenavirus RNA in 1.6% of 1,047 bats in Brazil that were sampled during 2007–2011. We identified Tacaribe virus in 2 Artibeus sp. bats and a new arenavirus species in Carollia perspicillata bats that we named Tietê mammarenavirus. Our results suggest that bats are an underrecognized arenavirus reservoir.
Bats are prominent hosts of zoonotic RNA viruses because of immunologic, physiologic, and ecologic factors (1). The Arenaviridae family comprises 4 genera: Reptarenavirus and Hartmanivirus, whose members infect reptiles; Antennavirus, whose members infect fish; and Mammarenavirus, whose members infect mammals. Mammarenaviruses can be separated into globally distributed lymphocytic choriomeningitis–Lassa virus serocomplex and New World arenaviruses (NWAs) (2). The NWAs Junin, Machupo, Sabia, Chapare, and Guanarito cause viral hemorrhagic fever and must be handled under Biosafety Level 4 conditions (2).
All highly pathogenic arenaviruses known thus far are hosted by and transmitted to humans from persistently infected rodents (2). Only Tacaribe virus (TCRV; Tacaribe mammarenavirus) has been identified in bats (3,4). Although TCRV is not considered a human pathogen, anecdotal evidence exists for potential laboratory acquired infection that causes influenza-like symptoms (5,6). In addition, TCRV is phylogenetically related to pathogenic arenaviruses that cause viral hemorrhagic fever; viral properties associated with severe disease, such as evasion of immune responses and cellular tropism, might be conserved in TCRV and genetically related animal arenaviruses (7).
Associations between TCRV and Artibeus spp. bats are supported only by limited epidemiologic data, including a single virus isolation and serologic evidence (3,4), considerable illness of bats during experimental infection (5), and isolation of TCRV from mosquitoes and ticks that primarily feed on rodents and rarely on bats (3,6). Limited genetic data exist for TCRV; a single genomic sequence was obtained from a bat-derived isolate generated in the 1950s from Trinidad that has been extensively passaged in mice and cell cultures and another from a recent tick-derived isolate (3,4,8).
We investigated diverse specimens from 1,047 adult bats belonging to 32 species collected from southeastern Brazil (Appendix). We analyzed a total of 3,670 different tissue specimens, including spleens (n = 893), lungs (n = 889), intestines (n = 973), and livers (n = 915), for arenavirus RNA by using reverse transcription PCR (RT-PCR) (9) modified to promote NWA amplification (Appendix Table 1, Figure 1).
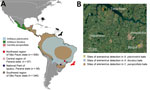
We detected arenavirus RNA in 4 Artibeus lituratus, 1 A. planirostris, and 12 Carollia perspicillata bats; the overall detection rate was 1.62% (95% CI 0.95%–2.59%). Arenavirus-positive bats were collected during 2007–2011 from 3 sampling sites located in both forest and urban areas within a 60-km radius (Figure 1), suggesting arenavirus maintenance in bat populations in this region. All 3 arenavirus-positive bat species are abundant in tropical environments and well-adapted to urban landscapes, indicating potential for dispersion and spillover to humans and other animals.
Most arenavirus-positive bats were collected in 2 forest fragments in 2007 (Tables 1, 2; Figure 1), where most bat species positive for arenavirus RNA were sampled. Whether high detection rates at those sites correspond to epizootics or sampling bias remains unknown.
All arenavirus-positive animals appeared healthy, suggesting limited negative effects of arenavirus infection on bat hosts. This observation was similar in rodent arenavirus hosts (10) and consistent with high TCRV seroprevalence in a serologic survey (4) but different from experimental TCRV infections (5), likely because of different routes and high doses used for infecting bats in laboratory settings. High seroprevalence and low arenavirus detection rates suggest that arenaviruses do not infect bats persistently, which is distinct from results for rodent arenavirus infections (11). Lack of persistence is important for public health because it indicates potential limitations of arenavirus shedding by bat hosts whose lifespan is <8–12 years (12).
We detected arenavirus RNA in multiple organs at similar concentrations, including spleens (mean, 1.2 × 107 RNA copies/mg) and lungs (mean, 6.4 × 106 RNA copies/mg) (p = 0.53 by Mann-Whitney U test) (Table 2), suggesting systemic infection similar to that observed in experimentally infected bats (5). We observed the highest arenavirus RNA concentration in the single arenavirus-positive intestine specimen, followed by the spleen, lung, liver, and kidney in that animal (Table 2). High arenavirus RNA concentrations in intestines are consistent with virus shedding through the enteric route, which has been observed during experimental infections with TCRV (5). Although rodents shed arenaviruses primarily through urine and saliva, shedding also occurs in feces (2). Determining differences in arenavirus transmission routes between bats and rodents will require further investigation. We were unsuccessful isolating bat arenaviruses from organ homogenates despite repeated attempts (Appendix), likely because of tissue degradation under tropical conditions.
We performed phylogenetic analysis of the partial sequence for the arenavirus RNA-dependent RNA polymerase gene obtained from RT-PCR screening. We found 2 NWA clades in bats from Brazil: 1 clade for both Artibeus spp. and 1 clade for C. perspicillata bats (GenBank accession nos. ON648806–16) (Figure 2, panel A). We obtained complete arenavirus coding sequences from 1 A. planirostris and 3 C. perspicillata bats (GenBank accession nos. ON648817–24) by using Illumina-based deep sequencing (Illumina, https://www.illumina.com); genome organization was identical to other mammarenaviruses. Both arenaviruses formed a well-supported monophyletic clade with TCRV in sister relationship to Junin and Machupo viruses (Figure 2, panel B) and Ocozocoautla de Espinosa virus that was possibly responsible for a hemorrhagic fever outbreak in Mexico (Figure 2, panel C) (13). These results highlight the genetic relationship of those bat-associated arenaviruses with highly pathogenic NWAs (Appendix Table 2). Identical topology in phylogenetic reconstructions argued against potential reassortment (Figure 2, panels B, C), and homogeneous sequence distances and recombination analyses along the genome did not indicate recombination events (Appendix Figure 2).
The A. planirostris bat was infected with a previously unknown TCRV strain (Appendix Table 2) that had an amino acid identity of 93.8%–95.5% with other TCRV sequences, depending on the protein analyzed. The arenaviruses from C. perspicillata bats formed a separate species in clade B of the TCRV serogroup (Figure 2, panels B, C). Species assignment relied on taxonomic criteria (14) that included exclusive detection in a distinct host, nucleotide sequence identity of <80% in the small segment, and 88.6%–90% amino acid identity in the nucleocapsid protein compared with TCRV and pairwise sequence comparison (https://www.ncbi.nlm.nih.gov/sutils/pasc/viridty.cgi?textpage=overview) results for large and small segments (Appendix Figure 3). The 5′ and 3′ ends of large and small genomic segments obtained from the newly identified arenavirus from C. perspicillata bats were nearly identical to TCRV, consistent with a close genetic relationship between those NWAs (Appendix Table 3, Figure 4). We propose that the arenavirus sequenced from C. perspicillata bats should be named Tietê virus (species Tiête mammarenavirus) and abbreviated as TETV; the name comes from the main river located <4 km from the capture site (Figure 1).
Arenavirus genetic diversity is hypothesized to result from a complex macro-evolutionary pattern that includes both co-evolution and host switching in the Muridae family of rodents. In South America, arenaviruses might have co-evolved with rodents in the Sigmodontinae subfamily, with the exception of TCRV (10). Further investigation will be required to determine whether bat arenaviruses evolved from an ancestral host switch involving rodents, which would be consistent with the genetic relationship between TCRV or Tietê virus and rodent-derived Ocozocoautla de Espinosa virus, or whether bats and arenaviruses co-evolved. Of note, bats play an essential role in ecosystems, and stigmatization of bats as sources of zoonotic viruses is unwarranted.
In summary, the epidemiology, genealogy, and zoonotic potential of bat arenaviruses deserve further investigation. Our results suggest that bats are an underrecognized arenavirus reservoir.
Dr. Góes is a postdoctoral researcher at the Institute of Virology at Charité-Universitätsmedizin, Berlin, Germany. His research focuses on pathogen discovery, virus epidemiology, and evolution of emerging viruses.
Acknowledgments
Authorization for bat captures was provided by the Instituto Brasileiro do Meio Ambiente e dos Recursos Naturais (approval nos. 12751-2, 12751-3, 21748-1, and 27346-2), Instituto Ambiental do Paraná (approval no. 235/10), ethics committee of the Universidade Estadual Paulista (UNESP) (approval no. 009/2012), and the ethics committee of the Institute of Biomedical Science from the University of São Paulo (approval no. 56–18–03/2014).
This work was supported by the National Council for Scientific and Technological Development–CNPq (grant agreement nos. 152365/2019-2 and 203084/2019-5), São Paulo Research Foundation (FAPESP) (grant agreement nos. 2013/11.006-0 and 2014/15090-8), and Human Fronter Science Program (grant agreement no. RPG0013/2018).
References
- Guth S, Mollentze N, Renault K, Streicker DG, Visher E, Boots M, et al. Bats host the most virulent-but not the most dangerous-zoonotic viruses. Proc Natl Acad Sci U S A. 2022;119:
e2113628119 . DOIPubMedGoogle Scholar - Charrel RN, de Lamballerie X. Zoonotic aspects of arenavirus infections. Vet Microbiol. 2010;140:213–20. DOIPubMedGoogle Scholar
- Downs WG, Anderson CR, Spence L, Aitken THG, Greenhall AH. Tacaribe virus, a new agent isolated from Artibeus bats and mosquitoes in Trinidad, West Indies. Am J Trop Med Hyg. 1963;12:640–6. DOIPubMedGoogle Scholar
- Malmlov A, Seetahal J, Carrington C, Ramkisson V, Foster J, Miazgowicz KL, et al. Serological evidence of arenavirus circulation among fruit bats in Trinidad. PLoS One. 2017;12:
e0185308 . DOIPubMedGoogle Scholar - Cogswell-Hawkinson A, Bowen R, James S, Gardiner D, Calisher CH, Adams R, et al. Tacaribe virus causes fatal infection of an ostensible reservoir host, the Jamaican fruit bat. J Virol. 2012;86:5791–9. DOIPubMedGoogle Scholar
- Sayler KA, Barbet AF, Chamberlain C, Clapp WL, Alleman R, Loeb JC, et al. Isolation of Tacaribe virus, a Caribbean arenavirus, from host-seeking Amblyomma americanum ticks in Florida. PLoS One. 2014;9:
e115769 . DOIPubMedGoogle Scholar - Moreno H, Möller R, Fedeli C, Gerold G, Kunz S. Comparison of the innate immune responses to pathogenic and nonpathogenic clade B New World arenaviruses. J Virol. 2019;93:e00148–19. DOIPubMedGoogle Scholar
- Holzerland J, Leske A, Fénéant L, Garcin D, Kolakofsky D, Groseth A. Complete genome sequence of Tacaribe virus. Arch Virol. 2020;165:1899–903. DOIPubMedGoogle Scholar
- Vieth S, Drosten C, Lenz O, Vincent M, Omilabu S, Hass M, et al. RT-PCR assay for detection of Lassa virus and related Old World arenaviruses targeting the L gene. Trans R Soc Trop Med Hyg. 2007;101:1253–64. DOIPubMedGoogle Scholar
- Grande-Pérez A, Martin V, Moreno H, de la Torre JC. Arenavirus quasispecies and their biological implications. Curr Top Microbiol Immunol. 2016;392:231–76. DOIPubMedGoogle Scholar
- Hoffmann C, Wurr S, Pallasch E, Bockholt S, Rieger T, Günther S, et al. Experimental Morogoro virus infection in its natural host, Mastomys natalensis. Viruses. 2021;13:851. DOIPubMedGoogle Scholar
- Jones ML. Longevity of captive mammals. Zool Garten N. F. Jena. 1982;52;113–28 [cited 2022 Nov 10]. http://www.rhinoresourcecenter.com/pdf_files/125/1256468598.pdf
- Cajimat MNB, Milazzo ML, Bradley RD, Fulhorst CF. Ocozocoautla de espinosa virus and hemorrhagic fever, Mexico. Emerg Infect Dis. 2012;18:401–5. DOIPubMedGoogle Scholar
- Radoshitzky SR, Bào Y, Buchmeier MJ, Charrel RN, Clawson AN, Clegg CS, et al. Past, present, and future of arenavirus taxonomy. Arch Virol. 2015;160:1851–74. DOIPubMedGoogle Scholar
Figures
Tables
Cite This ArticleOriginal Publication Date: November 15, 2022
Table of Contents – Volume 28, Number 12—December 2022
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Jan Felix Drexler, Helmut-Ruska-Haus, Institute of Virology, Campus Charité Mitte, Charitéplatz 1, 10098 Berlin, Germany
Top