Volume 28, Number 7—July 2022
Research Letter
Natural Reassortment of Eurasian Avian-Like Swine H1N1 and Avian H9N2 Influenza Viruses in Pigs, China
Figure
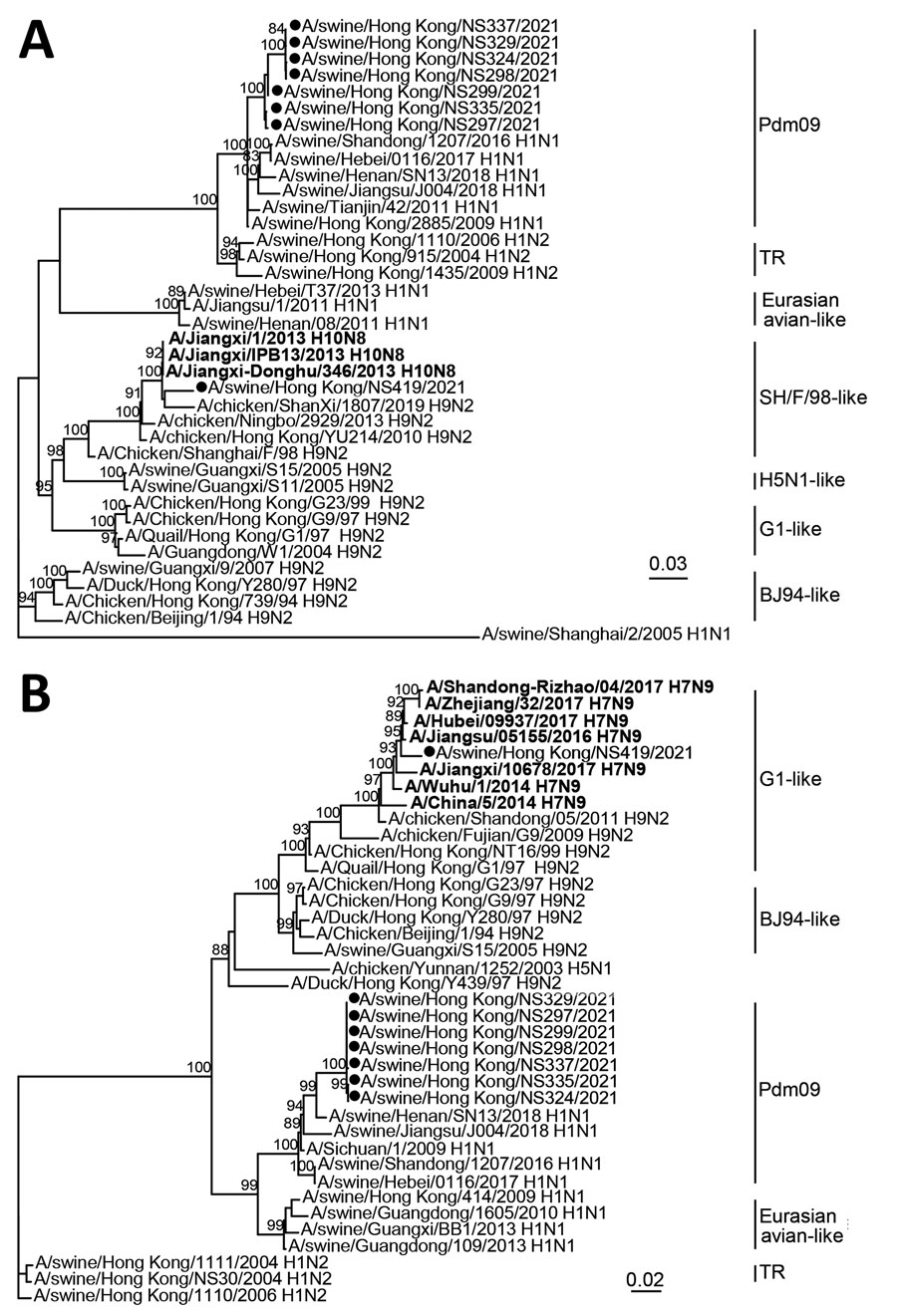
Figure. Phylogenetic tree of polymerase basic 1 (A) and matrix (B) gene sequences of swine influenza viruses from China and reference sequences. Bold indicates human H7N9 and H10N8 sequences. Viral sequences generated in this study (black circles) and those downloaded from public domains (Appendix Table) were aligned by using Muscle version 3.8 (http://www.drive5.com/muscle). Phylogenetic trees were constructed by IQ-TREE 1.6.12 (http://www.iqtree.org) by using the generalized time reversible plus gamma model. Major animal viral lineages are as shown. Bootstrap values ≥80% are shown. Scale bar indicates estimated genetic distance.
1These authors contributed equally to this article.
Page created: June 10, 2022
Page updated: June 18, 2022
Page reviewed: June 18, 2022
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.