Volume 29, Number 2—February 2023
Research
Influence of Landscape Patterns on Exposure to Lassa Fever Virus, Guinea
Cite This Article
Citation for Media
Abstract
Lassa fever virus (LASV) is the causative agent of Lassa fever, a disease endemic in West Africa. Exploring the relationships between environmental factors and LASV transmission across ecologically diverse regions can provide crucial information for the design of appropriate interventions and disease monitoring. We investigated LASV exposure in 2 ecologically diverse regions of Guinea. Our results showed that exposure to LASV was heterogenous between and within sites. LASV IgG seropositivity was 11.9% (95% CI 9.7%–14.5%) in a coastal study site in Basse-Guinée, but it was 59.6% (95% CI 55.5%–63.5%) in a forested study site located in Guinée Forestière. Seropositivity increased with age in the coastal site. We also found significant associations between exposure risk for LASV and landscape fragmentation in coastal and forested regions. Our study highlights the potential link between environmental change and LASV emergence and the urgent need for research on land management practices that reduce disease risks.
Lassa virus (LASV), the cause of Lassa fever in humans, is an enveloped and negative-sense, single-stranded RNA virus belonging to the Arenaviridae family (1). Initial clinical manifestations, such as fever, weakness, headache, respiratory and gastrointestinal symptoms, and conjunctivitis, are often nonspecific and could indicate of several other febrile illnesses (2). The mortality rate is unclear because of inconsistent reporting practices and lack of diagnostic services in most endemic areas but is estimated to be 5%–20% in hospitalized patients (2,3).
The Natal multimammate rat (Mastomys natalensis) is considered the main natural reservoir of LASV. It is a commensal rodent and agricultural pest that aggregates in human dwellings and surrounding fields (4). Both zoonotic and nonzoonotic transmission mechanisms have been described (5). Human LASV infections most commonly occur through infected rodent excreta, contaminated food, and inhalation of aerosols from rodent urine or droppings (6,7). Person-to-person transmission may also occur through exposure to contaminated bodily fluids (5,8).
The distribution of the Natal multimammate rat is a major risk factor for LASV exposure. The presence of the rat increases in houses during the dry season because of restricted food supply, which enhances the risk for humans encountering rodents or being in contact with excreta (9). Climatic variability leading to changes in human agriculture, food storage practices, and land cover can affect rodent population dynamics (10). Recent studies have also identified additional rodent reservoirs (e.g., African wood mouse and Guinea multimammate mouse), indicating the potential for a wider ecologic niche for the virus (11,12).
LASV is endemic in West Africa, particularly in Nigeria, Sierra Leone, Liberia, and Guinea (13,14). Estimates on annual human cases of LASV infections in West Africa range from 100,000 to 300,000, with 5,000–10,000 estimated deaths per year and 80% asymptomatic infections. Further epidemiologic studies are needed because these figures have been extrapolated from 1 longitudinal study conducted in Sierra Leone in 1987 (15). Characterizing the distribution and transmission intensity of LASV in endemic areas is essential for designing effective surveillance and disease prevention and control programs.
Exposure to LASV may lead to IgG seroconversion (16). Analysis of age-stratified serologic responses is a powerful tool to estimate exposure to LASV and to reconstruct historical patterns of transmission and distribution of previously unreported cases (17–19). Measurement of LASV antibodies has been used to detect circulation of LASV across Guinea in previous studies conducted in the 1990s and early 2000s (6,20,21). However, the current incidence and distribution of LASV in Guinea is largely unknown.
Our study aimed to characterize LASV exposure in human populations in 2 ecologically diverse regions of Guinea: the Basse-Guinée (coastal site); and the Guinée Forestière region, which includes the town of Guéckédou and Macenta Prefecture (forested site). We describe LASV-specific IgG levels within these populations, estimate transmission intensity by using age-stratified antibody responses, and explore associations with demographic and environmental risk factors, including fragmentation (the breaking apart of an organism’s preferred habitat into smaller patches).
Ethics Considerations
The part of the study conducted in Basse-Guinée (coastal site) was approved by the London School of Hygiene and Tropical Medicine Ethics Committee (reference no. 17429) and the Guinean Ministry of Health. For the part conducted in the town of Guéckédou and Macenta Prefecture (forested site), ethics approval for research projects on human volunteers was obtained from the Republic of Guinea’s National Ethics Committee for Health Research on March 16 (approval no. 012/CNERS/17).
Study Sites
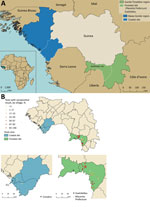
Guinea is administratively classified into regions on the basis of broad ecologic characteristics, including the Basse-Guinée and Guinée Forestière regions (22). The samples used in this study were collected during 3 separate studies in 2 geographically diverse regions of Guinea: Basse-Guinée (coastal site); and Guinée Forestière, including Macenta Prefecture and the town of Guéckédou (forested site) (22). Basse-Guinée (population 2,749,909, area 60,059 km2) is the coastal region of Guinea, extending for 30–40 miles inland from the Atlantic Ocean to the coastal plains. Basse-Guinée is mainly covered by mangroves and cultivated areas. The rainy season lasts 3 months per year in this region (Figure 1, panel A). Macenta Prefecture (population 297,779, area 8,600 km2) and the town of Guéckédou (population 346,908) are located to the southeast of the country, close to the border with Liberia and Sierra Leone, within Guinée Forestière. In addition to primary and secondary forests (umbrella and palm trees), which mainly cover this area, mountains and savannah can also be found. The rainy season lasts 9 months per year in this region (23–25) (Figure 1, panel A).
Study Participants
We obtained written informed consent from participants. Consent forms stated that blood samples would be tested for antibodies specific to Ebola virus and other pathogens, including LASV.
Samples from Basse-Guinée were collected in 2016 as part of a clinical trial evaluating the safety and immunogenicity of the vesicular stomatitis virus–based Ebola virus vaccine in contacts of Ebola survivors. That trial was performed in partnership with Guinea’s Ministry of Health, the World Health Organization, and Public Health England. For this study site, we randomly selected 702 samples taken before vaccination and tested them for the presence of LASV-specific IgG. Data recorded included the person’s age, sex, and village of residence at the time of vaccination. The selected samples originated from 7 prefectures in Basse-Guinée, encompassing 24 subprefectures and 29 villages or neighborhoods.
We collected samples from Macenta Prefecture during February–December 2017 as part of a seroepidemiologic study on Ebola virus that included both affected and nonaffected villages during the 2013–2016 Ebola virus disease (EVD) outbreak (26). In total, 517 participants living in 7 subprefectures encompassing 44 villages were enrolled. Volunteers were recruited upon agreement. The volunteers were from the general population of the village, including bush meat hunters, their family members, local healthcare workers. Volunteers <16 years of age, pregnant persons, travelers or visitors, and persons with chronic disease were excluded from the original study. In total, we tested 516 samples from the Macenta Prefecture cohort for the presence of LASV-specific IgG. One sample was not analyzed because not enough serum remained to run the assay. For our analysis, we included 68 samples from Guéckédou, collected in 2018 in the context of a longitudinal study of EVD survivors, in the Macenta Prefecture population because both Guéckédou and Macenta Prefecture are located within Guinée Forestière.
Blood Sample Collection and Processing
We collected samples (5–20 mL) of human peripheral blood into nonanticoagulant Vacutainer blood collection tubes (Becton, Dickinson and Company, https://www.bd.com). Blood clotted within 2 hours, and we transported samples to the laboratory at 4°C. We centrifuged samples at 2,000 × g for 10 min at room temperature and aliquoted serum into 2-mL microtube tubes (Starsted, https://www.sarstedt.com) in an air-purifying class II microbiologic safety cabinet (Envair, https://envairtechnology.com). We shipped aliquots of serum samples at –20°C; we processed and stored samples at –80°C at Public Health England’s Porton Down laboratories.
Samples Analysis by LASV-Specific ELISA
LASV exposure, which is defined as the contact with the virus, can be assessed by the presence of LASV-specific antibodies in the blood (27). We used the Blackbox LASV IgG Kit from Bernhard Nocht Institute for Tropical Medicine in Germany (https://www.bnitm.de) for qualitative serologic detection of acute or past LASV infection (28,29). We used the kit according to the manufacturer’s instructions; all reagents were provided, including positive and negative controls. We ran each sample as single replicate. After washing them with a saline solution, we transferred 25 μL of diluted biotinylated recombinant LASV nucleoprotein onto the ELISA plate and added 25 μL of diluted sample before plate incubation for 24 h at 4°C. After a wash step, we added 50 μL of diluted streptavidin horseradish peroxidase and incubated the plate for 1 h at 4°C. Finally, we developed the plate by using 100 μL/well of 3,3′,5,5′-tetramethylbenzidine peroxidase substrate and stopped the chemical reaction by using 100 μL/well of 1 mol/L H2SO4. We obtained optical density (OD) readings at wavelengths 450–620 nm (OD450–620) by using SpectraMax plate reader with SoftMaxPro 9 software (Molecular Devices, https://www.moleculardevices.com). OD is a term used to describe the propagation of the light induced by the reaction. We interpreted results as valid if the following criteria were met: OD450–620 of negative control <0.10 and OD450–620 of positive control >1.00. We used average absolute OD450–620 absorbance value for negative control samples ODNeg,AV to calculate the assay cutoff ODCO according to the formula ODNeg,AV + 0.150 = ODCO. Index values (IVs) are values comparing the ODs of the sample and the cutoff. We calculated IV for the tested samples (IVsample) by using the formula IVsample = OD450–620 (sample)/ODCO. Results classification for IV was carried out as follows: IVsample = >1.00 = positive; 0.90 <IVsample <1.00 = equivocal; IVsample <0.90 = negative.
Environmental and Spatial Covariates
To assess the effect of land cover and other environmental covariates on LASV exposure, we assembled a dataset of remote sensing–derived land cover maps from 2017 at 100 m resolution (30). We reclassified land cover into the following standardized categories, which are mutually exclusive: closed forest (canopy cover >70% of surface area), open forest (canopy cover 15%–70% of surface area), shrubs (woody perennial plants with persistent stems <5 m), vegetation (plants without persistent stems and canopy cover <10% of surface area), and urban or built up areas.
Village populations had been recorded during field surveys. Because villages were only geolocated to centroids, we extracted environmental data within buffer radii of 500 m, 1 km, 2 km, 5 km, and 10 km from the village centroid to represent the possible distributions of the village and lands used (31). For all buffer radii, we extracted the proportions of different land cover types and the mean canopy cover. To evaluate the importance of landscape configuration (spatial arrangement of different elements of the landscape) (32), we also extracted class-level metrics, including perimeter area ratio, shape index (patch perimeter divided by the minimum possible patch perimeter), and fractal dimension index (an index of shape complexity) (33). Those indices are commonly used as measures of landscape fragmentation, with higher values indicating greater levels of fragmentation (34,35).
Models and Statistical Analyses
Because assays were not validated for quantitative antibody responses, we fit all models by using classified values of seropositivity. To characterize transmission intensity in both sites, we integrated data on age-specific frequency of seropositivity. We fit reversible catalytic models to age-specific seroprevalence data by using maximum-likelihood methods (17). Those models generate age-specific seroprevalence curves, enabling the calculation of a seroconversion rate (SCR), which representing the force of infection for the community, a measure of transmission intensity (number of infections per person per year). SCR is the rate at which seronegative persons become seropositive; although it is a measure of transmission intensity, SCR is not incidence because it does not capture seropositive persons who are repeatedly exposed. Those models are widely used to characterize transmission of endemic diseases such as malaria (36). We used profile likelihood plots to explore possible historical changes in transmission intensity represented by the SCR and selected final models by using likelihood ratio tests (37).
We additionally assessed whether fine-scale patterns of seroconversion were associated with landscape characteristics. Because environmental variables can be associated with risk at different spatial scales, we used a data-driven approach to identify environmental risk factors (38). Because our environmental dataset was high-dimensional and highly correlated, we first performed mixed-effects logistic regression models for each environmental variable at each spatial scale (126 in total). We excluded from subsequent analysis any environmental variables with p values >0.2. We used Pearson correlation to identify highly correlated variables (Pearson correlation coefficient >0.75). For these highly correlated variables (e.g., environmental factors at different spatial scales), we identified a single variable for inclusion based on the Akaike information criteria. Because sampling frames were different between sites, we ran models separately for both sites, using the primary outcome as the binary classification for person-level seropositivity. We fitted models with village as a random effect to account for observations within villages lacking independence. We first assessed associations with key demographic factors, including sex, continuous age, and age category. For the forested site model, we included only environmental predictors because demographic factors (age and sex) were not associated with exposure risks. For the coastal site, we included age category in all models in addition to environmental predictors. When assessing for inclusion in the multivariate analysis, we considered univariate results to be statistically significant at p<0.2 to avoid exclusion of variables that alone lack significance but contribute in the presence of other variables. We conducted final multivariate model selection by using the dredge function in the MuMIn package (39), selecting the best model according to the Akaike information criteria and using model averaging approaches to obtain average regression coefficients for the best fitting models. The code for environmental data extraction and model fitting is available online (https://github.com/kfornace/LFV_environmental).
In this study, we assessed LASV IgG seropositivity in 1,286 persons by using serum samples collected in 2 sites in Guinea during 2016–2018. In the coastal site, we analyzed 702 samples (Table; Figure 1, panel A; Appendix Table 1). In the forested site, we analyzed 516 samples from Macenta Prefecture and 68 from the town of Guéckédou (Table; Figure 1, panel A; Appendix Table 2).
We observed that exposure to LASV was highly heterogenous both between and within each study site (Figure 1, panel B). The LASV IgG seropositivity was 11.9% (95% CI 9.7%–14.5%) in the coastal site (Figure 1, panel B; Appendix Table 1), whereas it was 59.6% (95% CI 55.5%–63.5%) in the forested site (Figure 1, panel B; Appendix Table 2).
Demographic characteristics of exposed persons varied by study site. In the coastal site, seropositivity in women was 9.0% (95% CI 5.8%–13.1%) and in men was 13.5% (95% CI 11.0%–17.0%). For the forested site, seropositivity in women was 61.8% (95% CI 56.0%–67.7%) and in men was 58.0% (95% CI 52.5%–63.3%). Sex was not significantly associated with seropositivity in either site (coastal site, odds ratio 1.25 [95% CI 0.69–2.27], p = 0.46; forested site, odds ratio 0.90 [95% CI 0.61–1.33], p = 0.59).
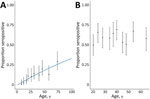
Age patterns of seropositivity differed markedly between sites. In the coastal site, exposure was strongly associated with age, and seropositivity increased with age (Figure 2, panel A). In that population, reversible catalytic models estimated the force of infection (SCR) as 0.0041 (95% CI 0.0033–0.0050). We observed no changes in historical transmission intensity (p = 0.328). In contrast, in the forested site, we identified no clear age patterns, and neither continuous nor categorical age were associated with person-level seropositivity (Figure 2, panel B). Although age ranges for included persons were different between both sites (Table), we detected substantial differences in exposure risks in similar age groups between sites. For example, persons 17–20 years of age had an estimated seropositivity of 5.4% (95% CI 1.5%–13.3%) in the coastal site, compared with 50.0% in the forested site (95% CI 37.2%–62.8%). The high proportions of seropositivity in the youngest age groups and lack of association with age precluded fitting reversible catalytic models for the forested region.
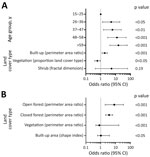
Despite clear differences in seropositivity between sites, we identified strong associations between exposure and environmental factors for both sites (Figure 3). As characterized by perimeter area ratios, fragmentation was strongly positively associated with exposure risk for both sites (Figure 4; Appendix Tables 3, 4). In the coastal site, odds of exposure increased with the perimeter area ratio of built-up areas but were lower where the proportion of vegetation in the landscape was higher. The fractal dimension index measuring fragmentation of shrubland was not significantly associated with exposure risk but improved overall model fit (Figure 4, panel A; Appendix Table 3). In the forested site, higher perimeter area ratios of both open and closed forests and the shape index of built areas, all indicating higher fragmentation levels, were positively associated with risk, whereas for vegetation a high perimeter area ratio was protective (Figure 4, panel B; Appendix Table 4). Across both sites, significant variables were identified at different spatial scales.
This study aimed to estimate LASV transmission intensity from serologic data in Guinea and analyze environmental risk factors. We found that LASV seropositivity was significantly greater in the forested site than in the coastal site. In 1993, Lukashevich et al. (20) also found high LASV nucleoprotein–specific IgG positivity among inhabitants of tropical forest (Guéckédou, Yomou, and Lola prefectures) and Guinea savannah (Faranah and Kindia prefectures) (25%–55%) and a lower positivity among inhabitants of mountainous areas (Pita, Labe, and Mali prefectures) and coastal areas (Boffa and Boké prefectures) (4%–7%).
We found that sex was not significantly associated with seropositivity in either site. This result was consistent with previous studies conducted in Guinea and Sierra Leone (15,20).
Our findings showed a significant increase in LASV IgG seropositivity with age in the coastal site but not in the forested site. Because no persons <16 years of age were included in our study, additional serologic studies of children would be needed to assess the age of exposure and how this is affected by environment. Although previous studies did not detect any correlation between antibody prevalence and age in these regions (20), differences in age patterns of seropositivity between regions were observed in other studies in West Africa. A study conducted in a forested region in 2000 found that children <10 and adults 20–29 years of age had the highest LASV IgG prevalence (6). A study conducted in Sierra Leone found that antibody prevalence increased with age, reaching a peak at 20–50 years of age and decreasing thereafter (15). A similar trend was reported in a 2015 study conducted in Nigeria (40). Age, often associated with differences in occupation or daily activities, may play a role in exposure (e.g., bushmeat hunters are mainly young to middle-age men and may have more contact with rodents). However, our results in the forested site suggest that in some areas, age-independent factors such as environmental features may have a greater effect on exposure risk.
All models showed an association between landscape patterns and exposure risk for both study sites (Appendix Tables 3–6). Associations between landscape fragmentation and exposure risk to zoonotic diseases also have been observed for malaria (41,42) and Ebola virus (43). Fragmentation may affect disease transmission by increasing spatial overlap between humans and wildlife populations at habitat edges (41). In our study, the fragmentation and the proportion of forests appear to affect exposure risk but their exact role in the complex dynamics of LASV circulation remains to be determined. Because the distribution of Natal multimammate rats is the leading LASV risk factor (9), the link between landscape patterns and the distribution of rodent reservoirs should be further explored. Of interest, one study found that the proportion of LASV-infected Natal multimammate rats was higher in forested regions than in coastal regions (13).
One limitation of our study is that, although the Blackbox IgG ELISA was shown to be sensitive (90%) and very specific (99%) (28), it was not tested against other arenaviruses because of a lack of suitable serum panels. We cannot fully exclude the potential circulation of related or yet undescribed arenaviruses in specific regions (44) and a partial cross-reactivity of antibodies against other arenaviruses. In addition, because the samples were collected as part of a vaccine evaluation and a seroepidemiologic study on Ebola virus, the sampling procedure was not standardized between the studies. Because the sampling approaches were not designed to be representative of the wider populations across Guinea, the estimates cannot be used to generalize about population-level disease risks.
In addition, behavioral exposure risks (e.g., occupation, medical history, hunting or foraging habits, location of previous residence, methods for food storage, and rodent trapping around the house) could not be evaluated because we did not collect person-level questionnaire data. Future studies could assess the number of persons in the household, the quality of the building, and proximity to rodent burrows. This study measured only IgG in serum samples collected at a single timepoint. As such, our data do not enable us to capture the potential effect of changes in seasonal rodent activity, vegetation patterns surrounding the villages, or the fluctuation of LASV persistence in rodents linked to climate variation. Because persons tend to move frequently across wide regions, either to find work during harvest season or to hunt (45) or in response to EVD hotspots, they may have been exposed to LASV in a different location from their current residence.
Because samples were not collected uniformly from younger age groups and a high proportion of persons from the forested region were exposed by 18 years of age, transmission intensity could not be directly compared between sites using reversible catalytic models. Although population-based epidemiologic studies are required to characterize wider LASV exposure and risk factors, the differences between these populations surveyed highlight the potential heterogeneity of LASV transmission intensity in Guinea.
In conclusion, LASV dynamics result from complex interactions between several human and environmental factors. Our study compared the LASV-specific IgG seropositivity in coastal and forested regions in Guinea and described associations between exposure to LASV and landscape patterns. Enhanced knowledge of regions with a potentially increased risk for LASV exposure can help improve epidemiologic surveillance; devise a global approach to optimize the health of humans, animals, and the environment; and highlight potential trial sites to assess new LASV vaccines and therapeutics.
Dr. Longet is a postdoctoral researcher in immunology at the University of Oxford. She specializes in infectious diseases and vaccines.
Acknowledgment
This work was funded by the US Food and Drug Administration (grant no. HHSF223201510104C) and the CGIAR Research Program on Agriculture for Nutrition and Health (grant no. A4NH). S.G. was supported by grants from the German Research Foundation (grant nos. GU883/5-1 and GU883/5-2). K.F. is supported by a Sir Henry Dale Fellowship jointly funded by the Wellcome Trust and the Royal Society (grant no. 221963/Z/20/Z).
References
- Hensley LE, Smith MA, Geisbert JB, Fritz EA, Daddario-DiCaprio KM, Larsen T, et al. Pathogenesis of Lassa fever in cynomolgus macaques. Virol J. 2011;8:205. DOIPubMedGoogle Scholar
- Richmond JK, Baglole DJ. Lassa fever: epidemiology, clinical features, and social consequences. BMJ. 2003;327:1271–5. DOIPubMedGoogle Scholar
- McCormick JB, Walker DH, King IJ, Webb PA, Elliott LH, Whitfield SG, et al. Lassa virus hepatitis: a study of fatal Lassa fever in humans. Am J Trop Med Hyg. 1986;35:401–7. DOIPubMedGoogle Scholar
- Monath TP, Newhouse VF, Kemp GE, Setzer HW, Cacciapuoti A. Lassa virus isolation from Mastomys natalensis rodents during an epidemic in Sierra Leone. Science. 1974;185:263–5. DOIPubMedGoogle Scholar
- Fisher-Hoch SP, Tomori O, Nasidi A, Perez-Oronoz GI, Fakile Y, Hutwagner L, et al. Review of cases of nosocomial Lassa fever in Nigeria: the high price of poor medical practice. BMJ. 1995;311:857–9. DOIPubMedGoogle Scholar
- Kernéis S, Koivogui L, Magassouba N, Koulemou K, Lewis R, Aplogan A, et al. Prevalence and risk factors of Lassa seropositivity in inhabitants of the forest region of Guinea: a cross-sectional study. PLoS Negl Trop Dis. 2009;3:
e548 . DOIPubMedGoogle Scholar - Stephenson EH, Larson EW, Dominik JW. Effect of environmental factors on aerosol-induced Lassa virus infection. J Med Virol. 1984;14:295–303. DOIPubMedGoogle Scholar
- Ajayi NA, Nwigwe CG, Azuogu BN, Onyire BN, Nwonwu EU, Ogbonnaya LU, et al. Containing a Lassa fever epidemic in a resource-limited setting: outbreak description and lessons learned from Abakaliki, Nigeria (January-March 2012). Int J Infect Dis. 2013;17:e1011–6. DOIPubMedGoogle Scholar
- Fichet-Calvet E, Lecompte E, Koivogui L, Soropogui B, Doré A, Kourouma F, et al. Fluctuation of abundance and Lassa virus prevalence in Mastomys natalensis in Guinea, West Africa. Vector Borne Zoonotic Dis. 2007;7:119–28. DOIPubMedGoogle Scholar
- Redding DW, Gibb R, Dan-Nwafor CC, Ilori EA, Yashe RU, Oladele SH, et al. Geographical drivers and climate-linked dynamics of Lassa fever in Nigeria. Nat Commun. 2021;12:5759. DOIPubMedGoogle Scholar
- Olayemi A, Oyeyiola A, Obadare A, Igbokwe J, Adesina AS, Onwe F, et al. Widespread arenavirus occurrence and seroprevalence in small mammals, Nigeria. Parasit Vectors. 2018;11:416. DOIPubMedGoogle Scholar
- Olayemi A, Cadar D, Magassouba N, Obadare A, Kourouma F, Oyeyiola A, et al. New hosts of the Lassa virus. Sci Rep. 2016;6:25280. DOIPubMedGoogle Scholar
- Lecompte E, Fichet-Calvet E, Daffis S, Koulémou K, Sylla O, Kourouma F, et al. Mastomys natalensis and Lassa fever, West Africa. Emerg Infect Dis. 2006;12:1971–4. DOIPubMedGoogle Scholar
- Ogbu O, Ajuluchukwu E, Uneke CJ. Lassa fever in West African sub-region: an overview. J Vector Borne Dis. 2007;44:1–11.PubMedGoogle Scholar
- McCormick JB, Webb PA, Krebs JW, Johnson KM, Smith ES. A prospective study of the epidemiology and ecology of Lassa fever. J Infect Dis. 1987;155:437–44. DOIPubMedGoogle Scholar
- Bond N, Schieffelin JS, Moses LM, Bennett AJ, Bausch DG. A historical look at the first reported cases of Lassa fever: IgG antibodies 40 years after acute infection. Am J Trop Med Hyg. 2013;88:241–4. DOIPubMedGoogle Scholar
- Drakeley CJ, Corran PH, Coleman PG, Tongren JE, McDonald SL, Carneiro I, et al. Estimating medium- and long-term trends in malaria transmission by using serological markers of malaria exposure. Proc Natl Acad Sci U S A. 2005;102:5108–13. DOIPubMedGoogle Scholar
- Greenhouse B, Daily J, Guinovart C, Goncalves B, Beeson J, Bell D, et al.; Malaria Serology Convening. Priority use cases for antibody-detecting assays of recent malaria exposure as tools to achieve and sustain malaria elimination. Gates Open Res. 2019;3:131. DOIPubMedGoogle Scholar
- Arnold BF, Scobie HM, Priest JW, Lammie PJ. Integrated serologic surveillance of population immunity and disease transmission. Emerg Infect Dis. 2018;24:1188–94. DOIPubMedGoogle Scholar
- Lukashevich IS, Clegg JC, Sidibe K. Lassa virus activity in Guinea: distribution of human antiviral antibody defined using enzyme-linked immunosorbent assay with recombinant antigen. J Med Virol. 1993;40:210–7. DOIPubMedGoogle Scholar
- Bausch DG, Demby AH, Coulibaly M, Kanu J, Goba A, Bah A, et al.; Epidemiology of Human Disease and Clinical Observations. Lassa fever in Guinea: I. Epidemiology of human disease and clinical observations. Vector Borne Zoonotic Dis. 2001;1:269–81. DOIPubMedGoogle Scholar
- L’Institut Guinéen d’Études et de Sondages. Histoire politique de la Guinée [cited 2022 Oct 18]. https://www.iges-guinee.com/histoire-politique-de-la-guinee
- République de Guinée. Institut National de la Statistique [cited 2022 Oct 18]. https://www.stat-guinee.org
- Wey J, Guillaume J, Kolie D, Souma Y, Cisse S, Traoré S. Guinée Forestière: de l’etude de systemes de productions a base riz pluvial à la conception de systemes d’agriculture durable. 2001 [cited 2022 Oct 18]. https://agritrop.cirad.fr/485415/1/document_485415.pdf
- Diallo MB. Présentation générale du pays et méthodologie de l'enquête: caractéristiques de la Guinée [cited 2022 Oct 18]. https://dhsprogram.com/pubs/pdf/FR109/01Chapitre01.pdf
- Thom R, Tipton T, Strecker T, Hall Y, Akoi Bore J, Maes P, et al. Longitudinal antibody and T cell responses in Ebola virus disease survivors and contacts: an observational cohort study. Lancet Infect Dis. 2021;21:507–16. DOIPubMedGoogle Scholar
- Ogunro BN, Olugasa BO, Kayode A, Ishola OO, Kolawole ON, Odigie EA, et al. Detection of antibody and antigen for Lassa virus nucleoprotein in monkeys from southern Nigeria. J Epidemiol Glob Health. 2019;9:125–7. DOIPubMedGoogle Scholar
- Gabriel M, Adomeh DI, Ehimuan J, Oyakhilome J, Omomoh EO, Ighodalo Y, et al. Development and evaluation of antibody-capture immunoassays for detection of Lassa virus nucleoprotein-specific immunoglobulin M and G. PLoS Negl Trop Dis. 2018;12:
e0006361 . DOIPubMedGoogle Scholar - Schmitz H, Gabriel M, Emmerich P. Specific detection of antibodies to different flaviviruses using a new immune complex ELISA. Med Microbiol Immunol (Berl). 2011;200:233–9. DOIPubMedGoogle Scholar
- Buchhorn M, Smets B, Bertels L, De Roo B, Lesiv M, Tsendbazar N-E, et al. Copernicus Global Land Service: land cover 100m: collection 3: epoch 2019: globe. 2020 Sep 8 [cited 2022 May 27]. https://zenodo.org/record/3939050
- Brock PM, Fornace KM, Grigg MJ, Anstey NM, William T, Cox J, et al. Predictive analysis across spatial scales links zoonotic malaria to deforestation. Proc Biol Sci. 2019;286:
20182351 . DOIPubMedGoogle Scholar - Thierry H, Rose E, Rogers H. Landscape configuration and frugivore identity affect seed rain during restoration. Oikos. 2021 Oct 1 [cited 2022 May 27]. https://onlinelibrary.wiley.com/doi/full/10.1111/oik.08323
- McGarigal K, Cushman S, Catherine Neel MEE. Spatial pattern analysis program for categorical maps. 2002 [cited 2022 May 27]. https://github.com/kmcgarigal/Fragstats
- Flowers B, Huang K-T, Aldana G. Analysis of the habitat fragmentation of ecosystems in Belize using landscape metrics. Sustainability. 2020;12:3024. DOIGoogle Scholar
- Liu Z, He C, Wu J. The relationship between habitat loss and fragmentation during urbanization: an empirical evaluation from 16 world cities. PLoS One. 2016;11:
e0154613 . DOIPubMedGoogle Scholar - Cook J, Reid H, Iavro J, Kuwahata M, Taleo G, Clements A, et al. Using serological measures to monitor changes in malaria transmission in Vanuatu. Malar J. 2010;9:169. DOIPubMedGoogle Scholar
- Stewart L, Gosling R, Griffin J, Gesase S, Campo J, Hashim R, et al. Rapid assessment of malaria transmission using age-specific sero-conversion rates. PLoS One. 2009;4:
e6083 . DOIPubMedGoogle Scholar - Fornace KM, Brock PM, Abidin TR, Grignard L, Herman LS, Chua TH, et al. Environmental risk factors and exposure to the zoonotic malaria parasite Plasmodium knowlesi across northern Sabah, Malaysia: a population-based cross-sectional survey. Lancet Planet Health. 2019;3:e179–86. DOIPubMedGoogle Scholar
- Bartoń K. Package “MuMIn”. 2022 Oct 12 [cited 2022 Oct 18]. https://cran.r-project.org/web/packages/MuMIn/MuMIn.pdf
- Tobin E, Asogun D, Akpede N, Adomeh D, Odia I, Gunther S. Lassa fever in Nigeria: insights into seroprevalence and risk factors in rural Edo State: a pilot study. J Med Trop. 2015;17:51–5. DOIGoogle Scholar
- Fornace KM, Abidin TR, Alexander N, Brock P, Grigg MJ, Murphy A, et al. Association between landscape factors and spatial patterns of Plasmodium knowlesi infections in Sabah, Malaysia. Emerg Infect Dis. 2016;22:201–8. DOIPubMedGoogle Scholar
- Stefani A, Dusfour I, Corrêa AP, Cruz MC, Dessay N, Galardo AK, et al. Land cover, land use and malaria in the Amazon: a systematic literature review of studies using remotely sensed data. Malar J. 2013;12:192. DOIPubMedGoogle Scholar
- Olivero J, Fa JE, Real R, Márquez AL, Farfán MA, Vargas JM, et al. Recent loss of closed forests is associated with Ebola virus disease outbreaks. Sci Rep. 2017;7:14291. DOIPubMedGoogle Scholar
- Coulibaly-N’Golo D, Allali B, Kouassi SK, Fichet-Calvet E, Becker-Ziaja B, Rieger T, et al. Novel arenavirus sequences in Hylomyscus sp. and Mus (Nannomys) setulosus from Côte d’Ivoire: implications for evolution of arenaviruses in Africa. PLoS One. 2011;6:
e20893 . DOIPubMedGoogle Scholar - Mari Saez A, Cherif Haidara M, Camara A, Kourouma F, Sage M, Magassouba N, et al. Rodent control to fight Lassa fever: Evaluation and lessons learned from a 4-year study in Upper Guinea. PLoS Negl Trop Dis. 2018;12:
e0006829 . DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleOriginal Publication Date: January 10, 2023
1These first authors contributed equally to this article.
2These senior authors contributed equally to this article.
Table of Contents – Volume 29, Number 2—February 2023
| EID Search Options |
|---|
|
|
|
|
|
|



Please use the form below to submit correspondence to the authors or contact them at the following address:
Miles W. Carroll and Stephanie Longet, Pandemic Sciences Institute, Nuffield Department of Medicine University of Oxford, Oxford, OX3 7BN, UK
Top