Volume 29, Number 2—February 2023
Research
Early Introduction and Community Transmission of SARS-CoV-2 Omicron Variant, New York, New York, USA
Figure 4
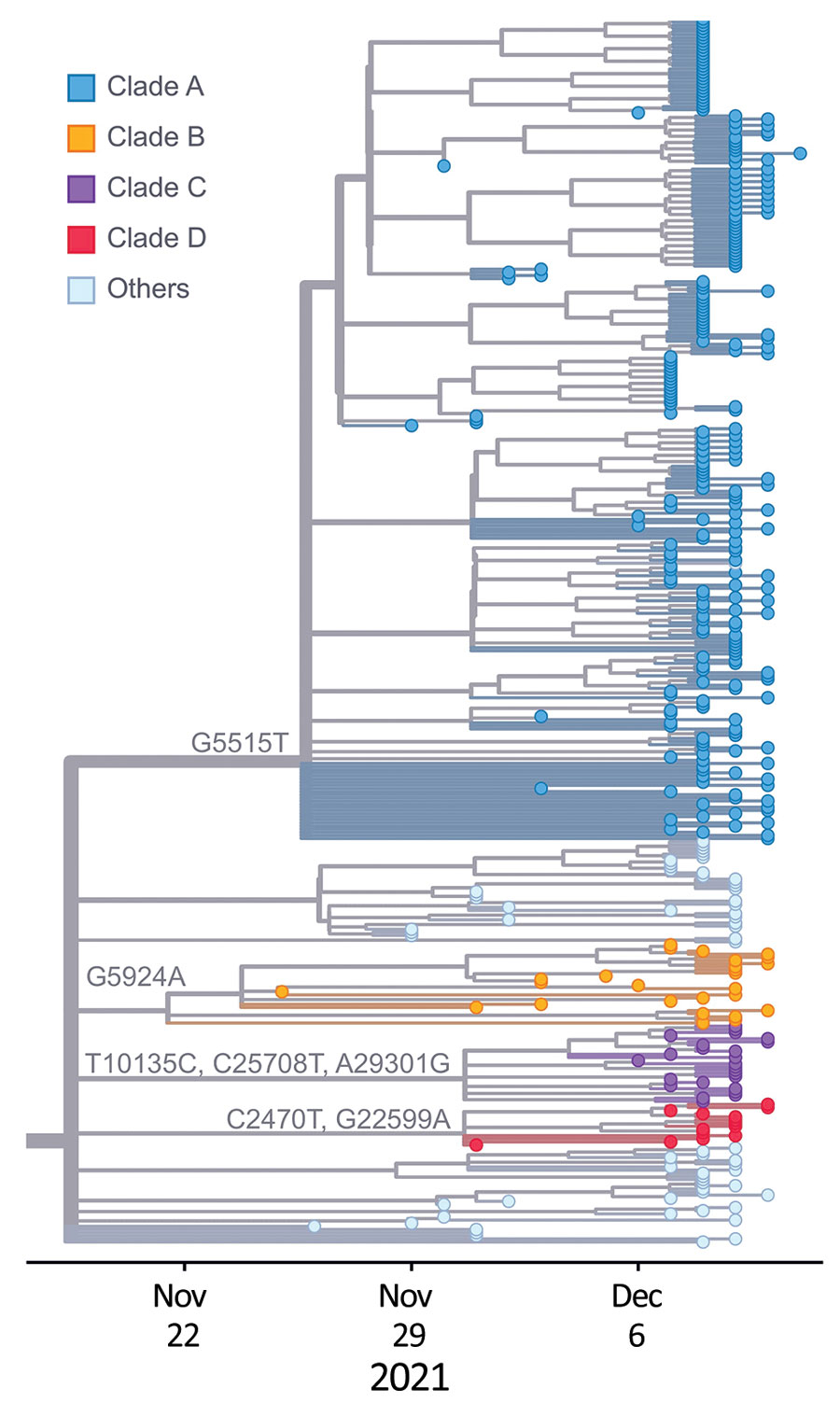
Figure 4. Phylogeny of 392 SARS-CoV-2 Omicron variant virus isolates from New York, New York, USA, November 25–December 11, 2021. Colored dots represent isolates from this study by clade. Substitution locations are indicated.
1These authors contributed equally to this article.
Page created: January 11, 2023
Page updated: January 21, 2023
Page reviewed: January 21, 2023
The conclusions, findings, and opinions expressed by authors contributing to this journal do not necessarily reflect the official position of the U.S. Department of Health and Human Services, the Public Health Service, the Centers for Disease Control and Prevention, or the authors' affiliated institutions. Use of trade names is for identification only and does not imply endorsement by any of the groups named above.