Volume 29, Number 7—July 2023
Dispatch
Candida vulturna Outbreak Caused by Cluster of Multidrug-Resistant Strains, China
Figure 1
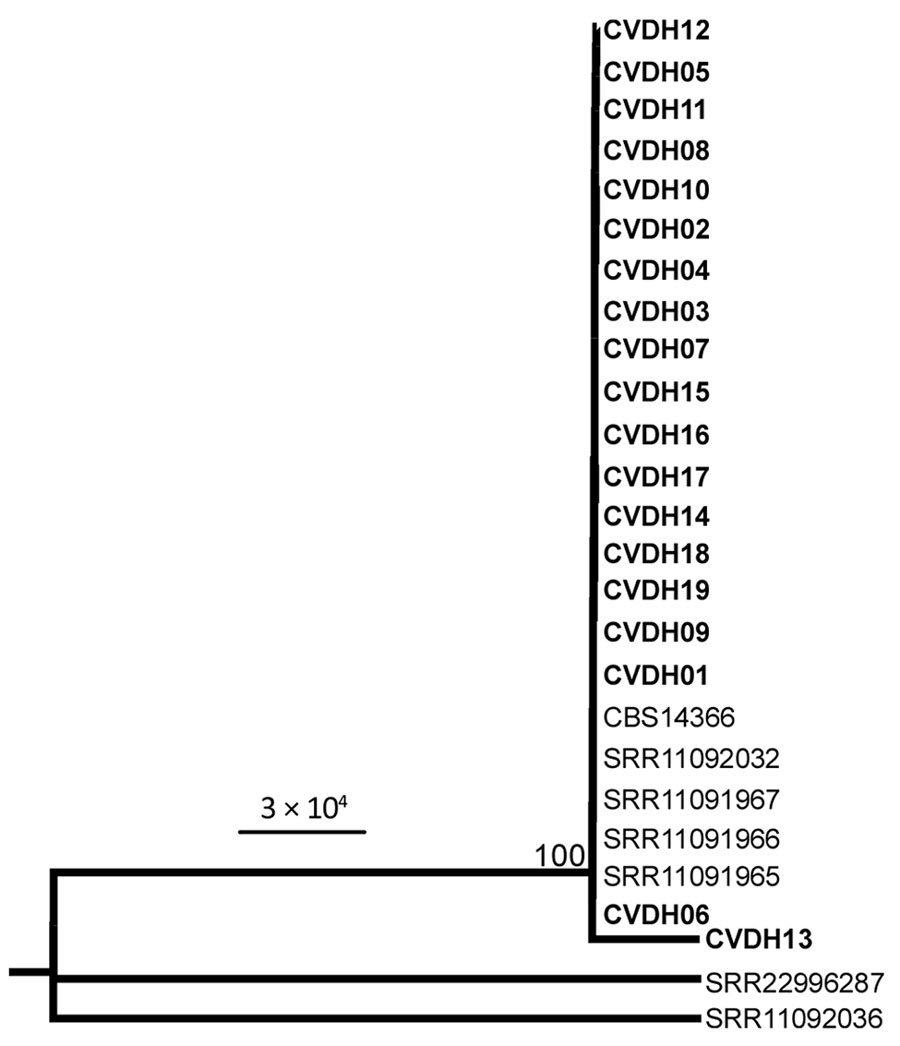
Figure 1. Maximum-likelihood phylogeny analysis of Candida vulturna strains from 19 infected patients in Shanxi Province, China, January 1, 2019–October 26, 2022, based on multilocus sequence typing (MLST). Eight genes (AAT1, ACC1, ADP1, ALA1, ERG11, RPB1, RPB2, and ZWF1) were concatenated and used for phylogenetic analyses. The tree was generated using the program RAxML (https://cme.h-its.org/exelixis/web/software/raxml). The general time reversible model, gamma distribution, 1,000 bootstraps, and midpoint root were adopted. Bold text indicates strains isolated in this study; reference strain data from whole-genome sequencing is from the National Center for Biotechnology Information gene database (accession nos. SRR11091965–67, SRR11092032, SRR11092036, SRR22996287). Sequences for strain CBS14366 were retrieved from its genomic assembly (GenBank accession no. GCA_026585945.1). Strains CVDH01-CVDH19 were isolated from patients of C. vulturna infection (cases C1–C19; Table; Appendix Figure 1). Scale bar indicates substitutions per site.
1These first authors contributed equally to this article.
2These senior authors contributed equally to this article.