Volume 4, Number 2—June 1998
Dispatch
Molecular Characterization of a Novel Rickettsia Species from Ixodes scapularis in Texas
Cite This Article
Citation for Media
Abstract
A novel Rickettsia species of undetermined pathogenicity was detected in Ixodes scapularis. DNA sequencing showed the highest nucleotide sequence similarities with R. australis for the 17 kDa gene, R. helvetica for gltA, and R. montana for rompA. The new organism, provisionally designated as genotype Cooleyi, is highly divergent in three conserved genes from recognized Rickettsia species.
Rickettsia spp., obligately intracellular, gram-negative, vector-transmitted bacteria, are divided antigenically into the typhus group (TG) and the spotted fever group (SFG). In nature, TG rickettsiae inhabit either flea (Rickettsia typhi and R. felis), louse (R. prowazekii), or tick (R. canada) arthropod hosts. Except for R. akari, whose invertebrate host is a mite, all SFG rickettsiae reside in tick hosts. SFG rickettsiae, excluding R. akari, are maintained in nature by a cycle involving both transovarial and transstadial passage of the microorganism in the tick and, for some species, amplification in small mammal hosts. Ticks infected with pathogenic rickettsiae (i.e., R. rickettsii, R. conorii, R. sibirica, R. australis, R. japonica, R. africae, and R. honei) that feed on humans, who are dead-end hosts, may transmit an SFG rickettsiosis (i.e., Rocky Mountain spotted fever [RMSF], boutonneuse fever, North Asian tick typhus, Queensland tick typhus, Japanese spotted fever, African tick bite fever, or Flinders Island spotted fever, respectively).
In the United States, Dermacentor andersoni and D. variabilis are the classic vectors of R. rickettsii to humans; other recognized vectors include Rhipicephalus sanguineus for R. rickettsii in Mexico and for R. conorii in Eurasia and Africa, Amblyomma cajennense for R. rickettsii in Brazil, and Ixodes holocyclus for R. australis in Australia. Other rickettsiae, such as R. parkeri and R. amblyommii, have been isolated from human-biting ticks; however, these rickettsiae have not been proven pathogenic for humans. The presence of apparently nonpathogenic rickettsiae whose vector host and geographic distribution overlap with pathogenic rickettsiae may interfere with the transmission of virulent rickettsiae to humans, as may be the case with R. peacockii (nonpathogenic) (1,2) and R. rickettsii in D. andersoni in the Bitterroot Valley of Montana.
The black-legged tick, I. scapularis, is of major public health importance because it feeds indiscriminately on numerous vertebrate hosts, including humans, and transmits Borrelia burgdorferi, Babesia microti, and human granulocytotropic ehrlichia to humans. Rickettsiae have been detected in I. scapularis (3-8); however, the ability of black-legged ticks to serve as vectors of rickettsiae to humans or to other small mammals important in maintaining and spreading rickettsiae in nature has not been examined. Because rickettsiae are regularly detected by immunofluorescence staining in I. scapularis specimens sent to the Texas Department of Health (5,7) and because the range of I. scapularis and RMSF cases overlaps in Texas (9), we characterized the immunofluorescence-detected microbial agent in this tick collected from Anderson County in eastern Texas.
We examined the field-collected I. scapularis, preserved in alcohol, to determine species, sex, and stage of development. Midgut tissues were removed and smeared on two glass slides. One tissue smear was screened for rickettsiae by the Gimenez staining technique. The smear was further analyzed by using a fluorescein isothiocyanate–labeled rabbit antibody against R. rickettsii that cross-reacts with many other SFG rickettsiae (10).
DNA from the tick was extracted with phenol:chloroform (1:1) by the standard method (11) after proteinase K and sodium dodecyl sulfate digestion of homogenized tick tissue. The DNA was resuspended in 50 μl of sterile water.
A volume of 10 μl of the polymerase chain reaction (PCR) product was separated by electrophoresis on 6% polyacrylamide gels. After electrophoresis, gels were stained with ethidium bromide, and amplicons were visualized with UV transillumination. Each PCR product was sequenced after DNA purification by a QIAquick Spin PCR Purification Kit (Qiagen, Santa Clarita, CA). Perkin Elmer Ready Reactions (Foster City, CA) and AmpliTaq FS with a Perkin Elmer ABI Prism 377 Automated DNA Sequencer (Foster City, CA) were used for sequencing.
Nucleotide and deduced amino acid sequences were aligned by using the PILEUP program (12). Phylogenetic analyses were conducted by using the PAUP parsimony algorithm (13), and the NEIGHBOR and FITCH distance matrix programs were implemented in the PHYLIP package (14). Parsimony analysis was implemented by using the heuristic algorithm and both ordered (transition:transversion = 3:1) and unordered characters. Because nucleotide sequences generally differed by less than 20%, distance methods used the Jukes-Cantor formula (15). Sequences were added at random with 10 replications. Bootstrap resampling with 200 replicates was used to place confidence values on groupings within trees.
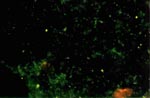
Rickettsiae were visualized in a Gimenez-stained tick smear. A duplicate slide contained rickettsiae stained by an SFG-specific fluorescein isothiocyanate–labeled antibody (Figure 1).
PCR amplification was performed on the fluorescent antibody-positive tick, and the DNA sequences of the Rickettsia-specific 17 kDa gene, gltA, and rompA were determined. Comparison of the highly conserved 17 kDa gene sequences showed the highest level of nucleotide sequence similarity (94.0% identical) between the I. scapularis rickettsia and R. australis (Figure 2A). Comparison of the I. scapularis rickettsia gltA partial nucleotide sequence showed the highest nucleotide sequence similarity (95.6% identical) with R. helvetica (Figure 2B). Comparison of the amplified 532 bp-product of rompA over a length of 441 nucleotides with all GenBank's rompA sequences showed the highest nucleotide sequence similarity (87.8% identical) with the established species R. montana (Figure 2C). A portion of rompB was not amplified for the rickettsia in this tick. The expected PCR product of 650 bp was obtained from the positive control, R. rickettsii DNA.
All three genes (17 kDa, gltA, and rompA) analyzed from this rickettsia had distinct sequences. Indeed, comparison of 17 kDa, gltA, and rompA partial nucleotide sequences showed at least 7.5%, 4.4%, and 12% differences, respectively, with the closest rickettsial species for each of the three genes analyzed. Both parsimony and distance phylogenetic methods indicated that the most closely related rickettsia was different at each of the three genes. However, bootstrap values for grouping the I. scapularis rickettsia with other rickettsiae were low, with the exception of its pairing with R. montana for the rompA gene (bootstrap value 99%, Figure 2C), perhaps because only relatively short partial sequences of the three genes were used for analysis. Otherwise, the three phylogenetic methods generally yielded trees with similar branching patterns; minor differences were observed within the SFG and in the position of the R. akari, R. australis, and R. felis groups for the gltA gene (Figure 2B). Figure 2A and B generally support the monophyletic nature of SFG. The few exceptions were the grouping of R. australis and R. akari with R. felis at the gltA gene (Figure 2B). This grouping of R. australis and R. akari with TG rickettsiae was supported by a bootstrap value of only 47%. In contrast, TG sequences never made up a monophyletic group when either gltA or 17 kDa sequences were analyzed by any of the phylogenetic methods. However, groupings placing TG sequences with sequences for other organisms such as R. bellii, AB Bacterium, or I. scapularis rickettsia were only weakly supported by bootstrap analyses.
When R. helvetica sequence information becomes available and is analyzed, it will be interesting to view the placement of the I. scapularis rickettsia and R. helvetica within the 17 kDa and rompA phylogeny. The closest rickettsial species at the 17 kDa, R. australis, and at the gltA gene, R. helvetica, are both Ixodes-associated rickettsiae. That a segment of the rompB was not amplified when the BG-12 primer pair was used is not surprising; the rompB segments of R. akari, R. bellii, R. helvetica, R. massiliae, and the I. scapularis rickettsia identified recently in Minnesota were also not amplified when this primer pair was used (8,18). Indeed, the highest sequence similarity of the gltA gene was with R. helvetica, which occurs in Swiss I. ricinus ticks (19).
Because I. scapularis readily feeds upon humans in the northeastern United States and occasionally in Texas, this rickettsia, genetically distinct from R. rickettsii, may cause an unrecognized rickettsiosis manifesting as a clinical disease different from the severe illness caused by R. rickettsii. In fact, I. scapularis has been associated epidemiologically with a fatal illness in Texas considered to be RMSF (5). Elliott and colleagues reported a fatal case of SFG rickettsiosis in a patient who jogged with a friend at a state park. Ticks collected at that site were only I. scapularis, some infected with an SFG rickettsia. An I. scapularis tick removed from the scalp of the patient's friend during an interview was later found to contain a rickettsia. In addition, rickettsiae of lower virulence may exist in Texas. A serologic study by Taylor and others suggested that 9% to 21% of sixth-grade students in Burleson and Mansfield Counties of Texas had prior immune stimulation by a rickettsial agent (7). None of the 32 children with serologic evidence of prior rickettsial infection had ever been treated for or diagnosed with RMSF. This study implies that these children were infected with a rickettsia of low virulence, or contrarily, that these children experienced asymptomatic immune stimulation with rickettsial antigens.
On the other hand, if not pathogenic for humans, the I. scapularis rickettsia could interfere with the transmission of R. rickettsii by other ticks to animals by inducing cross-protective immunity or by an interference mechanism such as that of R. peacockii (1,2). Noda et al. have proposed that a rickettsia found in laboratory-reared I. scapularis from Minnesota is an endosymbiont of the tick because the microorganism appears to be localized only to the ovaries (8). The potential pathogenicity of a rickettsia confined to the ovaries raises the question of whether or not it could escape from this intracellular niche to become infectious. Also, naturally occurring dual infections of rickettsiae in a tick have never been established. If this rickettsia is restricted to the ovaries, this tick may be precluded from serving as a vector of rickettsiosis under natural conditions. However, because the black-legged tick is a three-host tick, it may potentially serve as a vector of pathogenic rickettsiae to a wide range of susceptible hosts, including mammals, birds, and reptiles, from which other tick vectors could become infected after a rickettsemic blood meal. SFG rickettsiae have been detected in 10.2% to 16.7% of I. scapularis ticks in Texas (5,7), 5.6% in Connecticut (4), and 11.1% to 66.7% in South Carolina (3). While the numbers of ticks in these collections have been low, the prevalence of SFG rickettsiae in these ticks is among the highest recorded. Because rickettsiae have been regularly detected in unengorged black-legged ticks from different geographic locations (Texas, South Carolina, and Connecticut), we believe that I. scapularis is infected with SFG rickettsiae and that infection in these ticks likely represents more than ingestion in blood meals from rickettsemic animals.
Because for each of the three genes analyzed phylogenetically a robust relationship with other known Rickettsia species does not appear to exist, we believe that this rickettsia is genetically unique, and therefore, we provisionally designate this agent genotype Cooleyi after Dr. Robert A. Cooley, who served the Montana State Board of Entomology from 1899 to 1944. Isolation of this new rickettsial agent, characterization of its pathogenicity as the potential etiologic agent of an emerging infectious disease, and investigation of its role in its ecologic niche are subjects of interest for future study.
Adrian Billings is a Ph.D. candidate in the experimental pathology graduate program, University of Texas Medical Branch. His research interests include the epizootiology and pathogenesis of tick-borne zoonoses.
Acknowledgment
The authors thank Josie Ramirez and Kay Kantowski for their expert secretarial assistance. This research was supported by grants from the National Institutes of Health (AI 21242 and AI 31431-05A1).
References
- Neibylski ML, Schrumpf ME, Burgdorfer W, Fischer ER, Gage KL, Schwan TG. Rickettsia peacockii sp. nov., a new species infecting wood ticks, Dermacentor andersoni, in Western Montana. Int J Syst Bacteriol. 1997;47:446–52. DOIPubMedGoogle Scholar
- Burgdorfer W, Hayes SF, Marvos AJ. Nonpathogenic rickettsiae in Dermacentor andersoni: a limiting factor for the distribution of Rickettsia rickettsii. In: Burgdorfer W, Anacker RL, editors. Rickettsiae and rickettsial diseases. New York: Academic Press, Inc.; 1981; p. 585.
- Loving SM, Smith AB, DiSalvo AF, Burgdorfer WB. Distribution and prevalence of spotted fever group rickettsiae in ticks from South Carolina, with an epidemiological survey of persons bitten by infected ticks. Am J Trop Med Hyg. 1978;27:1255–60.PubMedGoogle Scholar
- Magnarelli LA, Anderson JF, Philip RN, Burgdorfer W. Antibodies to spotted fever-group rickettsiae in dogs and prevalence of infected ticks in southern Connecticut. Am J Vet Res. 1982;43:656–9.PubMedGoogle Scholar
- Elliott LB, Fournier PV, Teltow GJ. Rickettsia in Texas. Ann N Y Acad Sci. 1990;590:221–6. DOIPubMedGoogle Scholar
- Magnarelli LA, Andreadis TG, Stafford KC, Holland CJ. Rickettsiae and Borrelia burgdorferi in ixodid ticks. J Clin Microbiol. 1991;29:2798–804.PubMedGoogle Scholar
- Taylor JB, Tanner WB, Rawlings JA, Buck J, Elliott LB, Dewlett HJ, Serological evidence of subclinical Rocky Mountain spotted fever in Texas. J Infect Dis. 1985;151:367–8.PubMedGoogle Scholar
- Noda H, Munderloh UG, Kurtii TJ. Endosymbionts of ticks and their relationship to Wolbachia spp. and tick-borne pathogens of human and animals. Appl Environ Microbiol. 1997;63:3926–32.PubMedGoogle Scholar
- Hebert AG, Tzianabos T, Gamble WC, Chappell WA. Development and characterization of high-titered, group-specific fluorescent-antibody reagents for direct identification of rickettsiae in clinical specimens. J Clin Microbiol. 1980;11:503–7.PubMedGoogle Scholar
- Sambrook J, Fritsch EF, Maniatis T. Molecular cloning. A Laboratory Manual. Cold Spring Harbor (NY): Cold Spring Harbor Laboratory Press; 1989.
- Devereux J, Haeberli P, Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984;12:387–95. DOIPubMedGoogle Scholar
- Swofford DL. PAUP: phylogenetic analysis using parsimony [computer program]. Version 3.0. Champaign (IL): Illinois Natural History Survey; 1991.
- Felsenstein J. PHYLIP (Phylogeny Inference Package [computer program]). Version 3.5p. Seattle: Department of Genetics, University of Washington; 1993.
- Jukes TH, Cantor CR. Evolution of protein molecules. In: Munro HN, editor. Mammalian protein metabolism. New York: Academic Press; 1969; p. 21-132.
- Webb L, Carl M, Malloy DC, Dasch GA, Azad AF. Detection of murine typhus infection in fleas by using the polymerase chain reaction. J Clin Microbiol. 1990;28:530–4.PubMedGoogle Scholar
- Regnery RL, Spruill CL, Plikaytis BD. Genotypic identification of rickettsiae and estimation of intraspecies sequence divergence for portions of two rickettsial genes. J Bacteriol. 1991;173:1576–89.PubMedGoogle Scholar
- Eremeeva M, Yu X, Raoult D. Differentiation among spotted fever group rickettsiae species by analysis of restriction fragment length polymorphism of PCR-amplified DNA. J Clin Microbiol. 1994;32:803–10.PubMedGoogle Scholar
- Beati L, Peter O, Burgdorfer W, Aeschlimann A, Raoult D. Confirmation that Rickettsia helvetica sp. nov. is a distinct species of the spotted fever group of rickettsiae. Int J Syst Bacteriol. 1993;43:521–6. DOIPubMedGoogle Scholar
Figures
Cite This ArticleTable of Contents – Volume 4, Number 2—June 1998
| EID Search Options |
|---|
|
|
|
|
|
|