Volume 6, Number 5—October 2000
Perspective
Comparative Genomics and Understanding of Microbial Biology
Cite This Article
Citation for Media
Abstract
The sequences of close to 30 microbial genomes have been completed during the past 5 years, and the sequences of more than 100 genomes should be completed in the next 2 to 4 years. Soon, completed microbial genome sequences will represent a collection of >200,000 predicted coding sequences. While analysis of a single genome provides tremendous biological insights on any given organism, comparative analysis of multiple genomes provides substantially more information on the physiology and evolution of microbial species and expands our ability to better assign putative function to predicted coding sequences.
Perhaps no other field of research has been more energized by the application of genomic technology than the field of microbiology. Five years ago, The Institute for Genomic Research published the first complete genome sequence for a free-living organism, Haemophilus influenzae (1). Since then, 27 more microbial genome sequences (2-28) and 3 lower eukaryotic chromosome sequences (29-31) have been published, and at least three times that many sequencing projects are under way. Several important human pathogens are included: Helicobacter pylori (7, 19), Borrelia burgdorferi (12), Treponema pallidum (16), Mycobacterium tuberculosis (15), Rickettsia prowazekii (18), and Chlamydia species (17, 20); the simplest known free-living organism, Mycoplasma genitalium (2); the model organisms, Escherichia coli (8) and Bacillus subtilis (10); Aquifex aeolicus (13) and Thermotoga maritima (21), two thermophilic bacterial species that may represent some of the deepest branching members of the bacterial lineage; five representatives of the archaeal domain (3, 9, 11, 14, 28); and the first eukaryote, Saccaromyces cerevisiae (6).
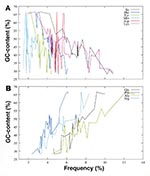
Genomic analyses show a tremendous variability not only in prokaryotic genome size but also in guanine plus cytosine (GC) content, from a low of 29% for B. burgdorferi (12) to a high of 68% for M. tuberculosis (15). The more than twofold difference in GC content affects the codon use and amino acid composition of species. For example, glycine, alanine, proline, and arginine, represented by GC-rich codons, are found at a much higher frequency in the predicted open reading frames from GC-rich genomes (Figure 1). Similarly, isoleucine, phenylalanine, tyrosine, methionine, and aspartic acid, represented by adenine plus thymine (A+T)-rich codons, are found at a higher frequency in the predicted open reading frames from AT-rich genomes. Genome organization also varies among microbial species, from single circular chromosomes to the most unusual situation seen with B. burgdorferi, whose genome is composed of an ~1 Mbp (million base pairs) linear chromosome and 21 linear and circular extrachromosomal elements.
Results from the completed prokaryotic genome sequences show that almost half of predicted coding regions identified are of unknown biological function (Table). More unexpectedly, approximately one-quarter of the predicted coding sequences in each species are unique, with no appreciable sequence similarity to any other known protein sequence. These data indicate large areas of microbial biology yet to be understood and suggest that in the microbial world the idea of a model organism may not be valid.
Functions can be assigned to coding regions by making generalizations about proteins. The number of genes involved in certain functions (transcription and translation, for example) is quite similar, even when genome size differs by fivefold or more (Figure 2). This suggests that a basic complement of proteins is required for certain cellular processes. In contrast, the number of proteins in other function categories--such as biosynthesis of amino acids, energy metabolism, transporters, and regulatory functions--can vary and often increases with genome size (Figure 2). A substantial proportion of the larger microbial genomes represent paralogous genes, that is, genes related by duplication rather than by vertical descent. With few exceptions, the number of total genes that are members of paralogous gene families increases from approximately 12% to 15% in genomes of 1 Mbp up to approximately 50% in genomes of >3 Mbp. For a given organism, as genome size increases so do functional diversity and biochemical complexity. The one exception with regard to gene duplications is B. burgdorferi, where nearly half the genes in its 1.5-Mbp genome are paralogs. Most paralogous genes in B. burgdorferi are plasmid-encoded genes, many of which are putative lipoproteins. The reason for the large number of paralogous genes in this organism is not known.
The study of transport proteins in prokaryotic species elucidates the relationship between genome size and biological complexity. The ability to discriminate and transport appropriate compounds is an essential function of cell membranes and their resident proteins. The fidelity of these transport reactions is particularly critical at the cytoplasmic membrane of prokayotes since this is the primary barrier that separates the physiologic reactions of the cytosol from the external environment. Many bacterial pathogens face astounding chemical and biological challenges from their host environment (e.g., the extreme acidity of the gastrointestinal tract challenges H. pylori ). In each host-pathogen relationship, the microbial membrane system contributes to the cell's strategy for energy production and carbon fixation while maintaining ionic homeostasis so that the enzymatic activities of the cytosol can proceed. In addition, all species encode proteins to expel toxic ions (particularly metals) and metabolites.
With complete genome sequences, evaluating the quantity and contribution of solute traffic across the membrane boundaries of pathogenic organisms is now possible. Comparisons between 11 sequenced bacterial pathogens (Table) indicate that approximately 6% of each genome encodes proteins (holoenzymes and subunits) involved in solute transport. This percentage is likely an underestimate since many of the gene products annotated as hypothetical proteins have hydropathy profiles reflective of known transporters. Genome size and the number of transport systems are directly related; the greatest number, 53, is annotated in the M. tuberculosis genome, and the smallest, 12, is found in the sequence of M. genitalium. Bacterial pathogens are heterotrophs; therefore, most of their import systems are used for the uptake of organic compounds (carbohydrates, organic alcohols, acids, amino acids, peptides, and amines). M. tuberculosis is the exception; it has 18 annotated transporters for organic substrates and 34 for the movement of ions. In this genome, there are nine copies of a P-type ATPase with a predicted substrate specificity for divalent cations. Whether this reflects a physiologic specialization allowing M. tuberculosis to be more resilient in its host environment is unknown.
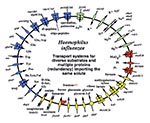
Underlying these general trends are some unique genomic solutions to niche selection and species survival. Two pathogens, H. influenzae and M. pneumoniae, both infect the respiratory tract, yet their strategies for acquiring solutes are distinct (Figure 3). In H. influenzae, the genes encoding transporters show a marked diversification. For example, in systems for amino acid uptake there are transporters for 13 different amino acids as well as proteins for the import of small peptides. These uptake systems work with several metabolic pathways for de novo amino acid synthesis. H. influenzae therefore employs a battery of redundant processes that allow it to optimize survival.
By contrast, the M. pneumoniae genome encodes only three transporters with substrate specificity for amino acids. This species, which may have evolved through reductive evolution from a gram-positive ancestor, has discarded all of the genes' encoding enzymes for amino acid biosynthesis (32). Instead of presenting a diverse group of porters for amino acid import, M. pneumoniae presents transport proteins with relatively broad substrate specificity: an oligopeptide transporter; a system for the aromatic residues tryptophan, tyrosine, and phenylalanine; and the spermidine/putrescine porter. M. pneumoniae uses a generalist strategy of maintaining proteins that are more versatile because of their broad substrate range. These same principles, diversification and redundancy, are repeated in the transport systems for carbohydrates (Figure 3).
In each analyzed genome, transport capacity appears to regulate the metabolic potential of that organism and dictates the range of tissues where a species can reside. Global analysis of transporters within a genome leads to several conclusions of practical consequence. First, culturing of pathogenic organisms is essential for understanding their physiology and for evaluating therapeutic agents. For species such as T. pallidum that have not yet successfully been grown in vitro, transporter analysis provides a clear starting point for the development of a defined culture medium based on information about the range of substrates a given cell can import and metabolize (16). Second, knowledge of transport processes and metabolic pathways they sustain provides novel solutions to the development of antimicrobial agents. An integrated view of cellular biochemistry enables selection of the pathway(s) essential for cell viability. Third, comparisons between genomes elucidate the diverse survival strategies found in pathogens with distinct evolutionary histories.
In addition to transport proteins, other membrane proteins in human pathogens play important roles in cell adhesion and as potential antigenic targets. Perhaps not surprisingly, in most human pathogens whose genome sequencing has been completed, mechanisms for generating antigenic variation on the cell surface have been proposed as a result of genome analysis. The following mechanisms for generating antigenic variation have been described: slipped strand mispairing within DNA sequence repeats found in 5-intergenic regions and coding sequences as described for H. influenzae (1), H. pylori (7), and M. tuberculosis (15); recombination between homologous genes encoding OSPs, as described for M. genitalium (2), M. pneumoniae (5), and T. pallidum (16); and clonal variability in surface-expressed proteins, as described for Plasmodium falciparum (29) and possibly B. burgdorferi (12). Studies of clinical isolates of some species have demonstrated phenotypic variation in the relevant cell surface proteins (33), suggesting that (at least for human pathogens) evolution of antigenic proteins occurs in real time, as cell populations divide.
The Institute for Genomic Research has recently launched the Comprehensive Microbial Resource (CMR), a database designed to facilitate comparative genomic studies on organisms whose genome sequencing has been completed. CMR (http://www.tigr.org) includes the sequence and annotation of each of the completed genomes and associated information (such as taxon and Gram stain pattern) about the organisms, the structure and composition of their DNA molecules (such as plasmid vs. chromosome and GC content), and many attributes of the protein sequences predicted from the DNA sequence (such as pI and molecular weight). With CMR, a user can query all the genomes at once or any subset of them, as well as make complex queries based on any properties of the organism or genome. CMR can be used to mine the completed genomes in ways not possible with single genome databases, furthering the progress of comparative genomics.
Studies of complete genomes have provided an unprecedented window into the evolution of life on this planet. For example, analysis of bacterial, archaeal, and eukaryotic genomes has confirmed the uniqueness of the archaeal lineage. Comparative studies of genome sequences have also revealed that lateral gene transfer has been very common over evolutionary time, occurring between both close and distant relatives (21). While the value of genome sequences in studies of evolution has been widely applauded, evolutionary analysis, which can provide great insight into genome sequences, is less well appreciated.
In any comparative biological study, an evolutionary perspective allows one to focus not only on characterizing the similarities and differences between species but also on explaining how and why those similarities and differences may have arisen (34). One area in genome analysis where an evolutionary perspective is useful is in distinguishing similarities due to homology (i.e., common ancestry) from those due to convergence (i.e., a separate origin). An example of the uses of distinguishing convergence from homology is the study of ribosomal RNA (rRNA) genes, which have been cloned from thousands of species; comparisons of these gene sequences are used extensively in evolutionary studies of these species. In early studies of rRNA sequences, most thermophiles were noted to have rRNA genes with high GC content relative to mesophiles. Since the rRNA genes in these thermophiles were similar in sequence and not just GC content, many of the thermophiles (e. g., the bacterial genera Aquifex and Thermotoga) were considered closely related. However, recent studies show that these genera are not closely related and the similarities in their rRNA genes are due to convergence (35). The most likely theory is that, to be stable at high temperatures, rRNAs need high GC contents, and therefore, even unrelated thermophiles will have similar sequences because many positions in the rRNA gene will converge to G or C (36). Finding this convergence explains the selective constraints on rRNA genes and shows that these genes may not be the best markers for evolutionary studies of species.
A highly practical use of evolutionary analysis in genome studies is predicting the function of genes (37). Predictions of gene function, a key step in the annotation of genomes, help researchers decide what types of experiments might be useful for a particular species or even a particular gene. Predictions are frequently made by assigning the uncharacterized gene the annotated function of the gene it is most similar to (similarity is measured by a database searching program such as BLAST). However, such predictions are frequently inaccurate because the annotated function may not be the best match (which would lead to error propagation if only the best match were used) and sequence similarity is not the best predictor of function. Several studies have shown that information about the evolutionary relationships of the uncharacterized gene can greatly improve predictions of function. For example, many gene families have undergone gene duplication. Since gene duplication is frequently accompanied by divergence of function, identifying the duplicate lineage (or orthology group) of a particular gene can greatly improve predictions of the gene's function. One orthology identification method is a clustering system developed by Tatusov et al. (38). This method (COG , for clusters of orthologous groups) classifies groups of genes by levels of sequence similarity. Although rapid and accurate in many cases, a clustering method such as COG does not always accurately infer the evolutionary history of genes. For this reason, and because orthologs do not always have the same function, we have developed a phylogenetic-tree-based function prediction method. This method involves inferring the evolutionary relationships of genes and then overlaying onto this history any experimentally determined functions of the genes. For uncharacterized genes, predictions are made according to their position in the tree relative to genes with known functions and according to evolutionary events (such as gene duplications) that may identify groups of genes with similar functions (39). Whatever method is used, information about the evolution of a gene can greatly improve function predictions.
Characterizing the evolutionary history of a particular gene is useful for other reasons. Identifying gene duplication events can provide insight into the mechanisms of gene duplication between genomes (e. g., proximity, age). Comparisons of the evolutionary history of different gene families can be used to infer recombination patterns within species as well as lateral gene transfers between species. While the likelihood of extensive gene transfers between species has thrown our concepts of the evolutionary history of species into disarray (21), identifying particular gene transfer events can be of great practical use. For example, there is a good correlation between regions of genomes responsible for pathogenicity and regions that have undergone lateral gene transfer (40). In analysis of eukaryotic genomes, identifying genes in the nucleus that have been transferred from the organellar genomes can best be done by phylogenetic analysis. Genes derived from the mitochondrial genome should branch most closely with genes from alpha-Proteobacteria, and genes derived from the chloroplast genome should branch most closely with cyanobacterial genes. In most cases, nuclear genes derived from these organelles still encode proteins that function in the organelles.
Evolutionary analysis is also very important for inferring gene loss. For example, we have used phylogenomic analysis to show that the mismatch repair genes MutS and MutL have been lost separately in multiple pathogenic species (e. g., H. pylori, M. tuberculosis, M. genitalium, and M. pneumoniae) (37). Several studies have shown that defects in mismatch repair increase pathogenicity, probably because these defects increase the mutation rate, which allows faster evolutionary response to immune systems and other host defenses. With more and more completed genome sequences, finding any other genes that may have been consistently lost in pathogenic species or strains will be possible. Identifying gene loss can also be useful in making function predictions for genes or species. For example, genes with a conserved association with each other might be lost as a unit--if one is lost, there is probably not much reason for the others to persist. The correlated presence and absence of genes constitute the basis of the phylogenetic profiles method of Pellegrini et al. (41), a very important tool in predicting functions.
The study of the evolutionary relationship of the M. tuberculosis complex has been greatly enhanced by the availability of two complete sequences from different strains (15 and www.tigr.org) and most sequences from the M. bovis genome (http://www.sanger.ac.uk). The H37Rv laboratory strain of M. tuberculosis was first isolated in 1905 and has been passed for many decades; substantial differences have been demonstrated between recent clinical isolates and genomes of laboratory strains with long histories of passage. A highly infectious clinical isolate of M. tuberculosis, CDC1551, was involved in a recent cluster of tuberculosis cases in the United States (42). Whole genome analysis of single nucleotide polymorphisms, insertions and deletions, and gene duplications provides comparisons that were previously unobtainable. Studies examining a limited set of M. tuberculosis genes from various strains suggest a limited sequence diversity between strains and in the complex, with a nucleotide polymorphism rate of approximately 1 in 10,000 bp (43). Detailed comparison of strains H37Rv and CDC1551 indicates a higher frequency of polymorphism, approximately 1 in 3,000 bp, with approximately half the polymorphism occurring in the intergenic regions. In other words, 50% of the polymorphisms are in 10% of the genome. While this rate is higher than that suggested (43), it still represents a lower nucleotide diversity than found in limited comparisons from other pathogens.
Examination of insertion and deletion events and gene duplication between species and strains allows insight into the evolutionary relationship of the M. tuberculosis complex. For example, a phospholipase C region, present in CDC1551 and absent in H37Rv, is also present in M. bovis. The simplest explanation for this is that the common ancestor of M. tuberculosis and M. bovis contained this region, and the region was subsequently deleted in the H37Rv lineage.
Membrane lipid proteins are identified by a unique signature sequence that is the target for a specific lipoprotein signal peptidase and that allows the cleaved protein product to attach by cysteinyl linkage to a glyceride-fatty acid lipid. Among the genes encoding membrane lipid proteins in strain H37Rv are two in tandem (Rv2543 and Rv2544). Nucleotide identity of >85% suggests that these two genes arose through duplication. The homologous genome region in strain CDC1551 contains the orthologs MT2618 and MT2620, respectively, as well as a third gene, MT2619, which by sequence similarity appears to represent an additional duplication (Figure 4). The increased induction of cytokines by CDC1551 is associated with the membrane lipid component (42). Modification of the lipid component by various protein components may contribute to differences in the immune response to M. tuberculosis infection in the host.
These examples illustrate how evolutionary information can benefit genome analysis. Complete genome sequences are also very useful. Gene loss, for example, cannot be readily identified without knowing the complete genome sequence of an organism. Since there are feedback loops between evolutionary and genome analyses, combining them into a single composite phylogenomic analysis may be advantageous (37, 44). As more and more genomes are completed, the benefits of combined evolutionary and genome analysis should become even more apparent.
Dr. Fraser is president and director of the Institute for Genomic Research. She is involved in genome analysis of microbial species and the use of comparative genome analysis in elucidating species diversity.
References
- Fleischmann RD, Adams MD, White O, Clayton RA, Kirkness EF, Kerlavage AR, Whole-genome random sequencing and assembly of Haemophilus influenzae Rd. Science. 1995;269:496–512. DOIPubMedGoogle Scholar
- Fraser CM, Gocayne JD, White O, Adams MD, Clayton RA, Fleischmann RD, The minimal gene complement of Mycoplasma genitalium. Science. 1995;270:397–403. DOIPubMedGoogle Scholar
- Bult CJ, White O, Olsen GJ, Zhou L, Fleischmann RD, Sutton GG, Complete genome sequence of the methanogenic archaeon, Methanococcus jannaschii. Science. 1996;273:1058–72. DOIPubMedGoogle Scholar
- Kaneko T, Sato S, Kotani H, Tanaka A, Asamizu E, Nakamure Y, Sequence analysis of the genome of the unicellular cyanobacterium Synechocystis sp. Strain PCC6803. II. Sequence determination of the entire genome and assignment of potential protein-coding regions. DNA Res. 1996;3:109–36. DOIPubMedGoogle Scholar
- Himmelreich R, Hilbert H, Plagens H, Pirki E, Li B-C, Hermann R. Complete genome sequence analysis of the genome of the bacterium Mycoplasma pneumoniae. Nucleic Acids Res. 1996;3:109–36.PubMedGoogle Scholar
- Tomb JF, White O, Kerlavage AR, Clayton RA, Sutton GG, Fleischmann RD, The complete genome sequence of the gastric pathogen Helicobacter pylori. Nature. 1997;388:539–47. DOIPubMedGoogle Scholar
- Blattner FR, Plunkett G III, Bloch CA, Perna NT, Burland V, Riley M, The complete genome sequence of Escherichia coli K-12. Science. 1997;277:1453–62. DOIPubMedGoogle Scholar
- Smith DR, Doucette-Stamm LA, Deloughery C, Lee H, Dubois J, Aldredge T, Complete genome sequence of Methanobacterium thermoautotrophicum deltaH: functional analysis and comparative genomics. J Bacteriol. 1997;179:7135–55.PubMedGoogle Scholar
- Kunst F, Ogasawara N, Moszer I, Albertini AM, Alloni G, Azevedo V, The complete genome sequence of the Gram-positive bacterium Bacillus subtilis. Nature. 1997;390:249–56. DOIPubMedGoogle Scholar
- Klenk HP, Clayton RA, Tomb JF, White O, Nelson KE, Ketchum KA, The complete genome sequence of the hyperthermophilic, sulfate-reducing archaeon Archaoglobus fulgidus. Nature. 1997;390:364–70. DOIPubMedGoogle Scholar
- Fraser CM, Casjens S, Huang WM, Sutton GG, Clayton R, Lathigra R, Genomic sequence of a Lyme disease spirochete, Borrelia burgdorferi. Nature. 1997;390:580–6. DOIPubMedGoogle Scholar
- Deckert G, Warren PV, Gaasterland T, Young WG, Lenox AL, Graham DE, The complete genome of the hyperthermophilic bacterium Aquifex aeolicus. Nature. 1998;392:353–8. DOIPubMedGoogle Scholar
- Kawarabayasi Y, Sawada M, Horikawa H, Haikawa Y, Hino Y, Yamamoto S, Complete sequence and gene organization of the genome of a hyperthermophilic archaebacterium, Pyrococcus horikoshii OT3. DNA Res. 1998;5:55–76. DOIPubMedGoogle Scholar
- Cole ST, Brosch R, Parkhill J, Garnier T, Churcher C, Harris D, Deciphering the biology of Mycobacterium tuberculosis from the complete genome sequence. Nature. 1998;393:537–44. DOIPubMedGoogle Scholar
- Fraser CM, Norris SJ, Weinstock GM, White O, Sutton GG, Dodson RJ, Complete genome sequence of Treponema pallidum, the syphilis spirochete. Science. 1998;281:375–88. DOIPubMedGoogle Scholar
- Stephens RS, Kalman S, Lammel C, Fan J, Marathe R, Aravind L, Genome sequence of an obligate intracellular pathogen of humans: Chlamydia trachomatis. Science. 1998;282:754–9. DOIPubMedGoogle Scholar
- Andersson SGE, Zomorodipour A, Andersson JO, Sicheritz-Ponten T, Alsmark UCM, Podowski RM, The genome sequence of Rickettsia prowazekii and the origin of mitochondria. Nature. 1998;396:133–40. DOIPubMedGoogle Scholar
- Alm RA, Ling LS, Moir DT, King BL, Brown ED, Doig PC, Genomic sequence comparison of two unrelated isolates of the human gastric pathogen Helicobacter pylori. Nature. 1999;397:176–80. DOIPubMedGoogle Scholar
- Kalman S, Mitchell W, Marathe R, Lammel C, Fan J, Hyman RW, Comparative genomes of Chlamydia pneumoniae and C. trachomatis. Nat Genet. 1999;21:385–9. DOIPubMedGoogle Scholar
- Nelson KE, Clayton RA, Gill SR, Gwinn ML, Dodson RJ, Haft DH, Evidence for lateral gene transfer between Archaea and bacteria from genome sequence of Thermotoga maritima. Nature. 1999;399:323–9. DOIPubMedGoogle Scholar
- White O, Eisen J, Heidelberg J, Hickey E, Peterson J, Dodson RJ, Genome sequence of the radioresistant bacterium Deinococcus radiodurans R1. Science. 1999;286:1571–7. DOIPubMedGoogle Scholar
- Parkhill J, Wren B, Mungall K, Ketley JM, Churcher C, Basham T, The complete genome sequence of the food borne pathogen Campylobacter jejuni. Nature. 2000;403:665–8. DOIPubMedGoogle Scholar
- Tettelin H, Saunders NJ, Heidelberg J, Jeffries AC, Nelson KE, Eisen JA, Complete genome sequence of Neisseria meningitidis serogroup B strain MC58. Science. 2000;287:1809–15. DOIPubMedGoogle Scholar
- Parkhill J, Achtman M, James KD, Bentley SD, Churcher C, Klee SR, Complete DNA sequence of a serogroup A strain of Neisseria meningitidis Z2491. Nature. 2000;404:502–6. DOIPubMedGoogle Scholar
- Read TD, Brunham RC, Shen C, Gill SR, Heidelberg JF, White O, Genome sequences of Chlamydia trachomatis MoPn and Chlamydia pneumoniae AR39. Nucleic Acids Res. 2000;28:1397–406. DOIPubMedGoogle Scholar
- Casjens S, Palmer N, van Vugt R, Huang WM, Stevenson B, Rosa P, A bacterial genome in flux; the twelve linear and nine circular extrachromosomal DNAs in an infectious isolate of the Lyme disease spirochete Borrelia burgdorferi. Mol Microbiol. 2000;35:490–516. DOIPubMedGoogle Scholar
- Kawarabayasi Y, Hino Y, Horikawa H, Yamazaki S, Haikawa Y, Jin-no K, Complete genome sequence of an aerobic hyper-thermophilic crenarchaeon, Aeropyrun pernix K1. DNA Res. 1999;6:83–101. DOIPubMedGoogle Scholar
- Gardner MJ, Tettelin H, Carucci DJ, Cummings LM, Aravind L, Koonin EV, Chromosome 2 sequence of the human malaria parasite Plasmodium falciparum. Science. 1998;282:1126–32. DOIPubMedGoogle Scholar
- Myler PJ, Audleman L, deVos T, Hixson G, Kiser P, Lemley C, major Freidlin chromosome 1 has an unusual distribution of protein-coding genes. Proc Natl Acad Sci U S A. 1999;96:2902–6. DOIPubMedGoogle Scholar
- Bowman S, Lawson D, Basham D, Brown D, Chillingworth T, Churcher CM, The complete nucleotide sequence of chromosome 3 of Plasmodium falciparum. Nature. 1999;400:532–8. DOIPubMedGoogle Scholar
- Razin S. Molecular biology and genetics of mycoplasma (Mollicutes). Microbiol Rev. 1985;49:419–55.PubMedGoogle Scholar
- Peterson SN, Bailey CC, Jensen JS, Borre MB, King ES, Bott KF, Characterization of repetitive DNA in the Mycoplasma genitalium genome: possible role in the generation of antigenic variation. Proc Natl Acad Sci U S A. 1995;92:11829–33. DOIPubMedGoogle Scholar
- Hirt RP, Logsdon JM Jr, Healy B, Dorey MW, Doolittle WF, Embley TM. Microsporidia are related to fungi: evidence from the largest subunit of RNA polymerase II and other proteins. Proc Natl Acad Sci U S A. 1999;96:580–5. DOIPubMedGoogle Scholar
- Galtier N, Tourasse N, Gouy M. A nonhyperthermophilic common ancestor to extant life forms. Science. 1999;283:220–1. DOIPubMedGoogle Scholar
- Eisen JA. A phylogenomic study of the MutS family of proteins. Nucleic Acids Res. 1998;26:4291–300. DOIPubMedGoogle Scholar
- Tatusov RL, Koonin EV, Lipman DJ. A genomic perspective on protein families. Science. 1997;278:631–7. DOIPubMedGoogle Scholar
- Eisen JA. Phylogenomics: improving functional predictions for uncharacterized genes by evolutionary analysis. Genome Res. 1998;8:163–7.PubMedGoogle Scholar
- Lawrence JG, Ochman H. Molecular archaeology of the Escherichia coli genome. Proc Natl Acad Sci U S A. 1998;95:9413–7. DOIPubMedGoogle Scholar
- Pellegrini M, Marcotte EM, Thompson MJ, Eisenberg D, Yeates TO. Assigning protein functions by comparative genome analysis: protein phylogenetic profiles. Proc Natl Acad Sci U S A. 1999;96:4285–8. DOIPubMedGoogle Scholar
- Manca C, Tsenova L, Barry CE III, Bergtold A, Freeman S, Haslett PA, CDC1551 induces a more vigorous host response in vivo and in vitro, but is not more virulent than other clinical isolates. J Immunol. 1999;162:6740–6.PubMedGoogle Scholar
- Sreevatsan S, Pan X, Stockbauer KE, Connell ND, Kreiswirth BN, Whittam TS, Restricted structural gene polymorphism in the Mycobacterium tuberculosis complex indicates evolutionarily recent global dissemination. Proc Natl Acad Sci U S A. 1997;94:9869–74. DOIPubMedGoogle Scholar
- Eisen JA, Hanawalt PC. A phylogenomic study of DNA repair genes, proteins, and processes. Mutat Res. 1999;435:171–213.PubMedGoogle Scholar
Figures
Table
Cite This ArticleTable of Contents – Volume 6, Number 5—October 2000
| EID Search Options |
|---|
|
|
|
|
|
|