Volume 8, Number 8—August 2002
Research
DNA Vaccine Expressing Conserved Influenza Virus Proteins Protective Against H5N1 Challenge Infection in Mice
Abstract
Influenza vaccination practice, which is based on neutralizing antibodies, requires being able to predict which viral strains will be circulating. If an unexpected strain, as in the 1997 H5N1 Hong Kong outbreak, or even a pandemic emerges, appropriate vaccines may take too long to prepare. Therefore, strategies based on conserved influenza antigens should be explored. We studied DNA vaccination in mice with plasmids expressing conserved nucleoprotein (NP) and matrix (M) from an H1N1 virus. After vaccination, mice were challenged with A/H5N1 viruses of low, intermediate, and high lethality. A/NP+A/M DNA vaccination reduced replication of A/Hong Kong/486/97 (HK/486), a nonlethal H5N1 strain, and protected against lethal challenge with more virulent A/Hong Kong/156/97 (HK/156). After HK/156 exposure, mice survived rechallenge with A/Hong Kong/483/97 (HK/483), although the DNA vaccination alone protected poorly against this highly virulent strain. In the absence of antigenically matched hemagglutinin-based vaccines, DNA vaccination with conserved influenza genes may provide a useful first line of defense against a rapidly spreading pandemic virus.
The 1997 outbreak of H5N1 avian influenza in humans in Hong Kong (1,2) caused alarm because it involved highly pathogenic strains of an influenza subtype to which humans lack immunity. This outbreak led to fears about inability to control a pandemic if a new strain should spread efficiently from human to human. Although prevention by vaccination is more desirable than treatment after infection, conventional immunization strategies have major limitations.
Neutralizing antibodies are specific to subtype and often strain, so vaccination based on eliciting such antibodies requires accurate prediction of the viral strains that will circulate during the influenza season and leaves little time for vaccine preparation. Even with usual epidemic strains, difficulties and delays in the production of an adequate vaccine supply have occurred in some years (3). A rapidly developing pandemic would shorten the timeframe to identify the viral strain and prepare an antigenically matched vaccine, while the need to vaccinate an entirely naïve population would exacerbate vaccine production and supply issues. In addition, H5 vaccine candidates, either H5 recombinant protein or a conventional surface antigen vaccine prepared from apathogenic H5N3 virus, have shown suboptimal immunogenicity in human trials (4,5).
A recent report on the molecular basis for virulence of H5N1 viruses (6) was accompanied by an article that discussed related public health issues, in which Laver and Garman (7) addressed the problem of how to control pandemics and concluded that currently “the most promising first line of defense” is use of antiviral drugs. These drugs, however, reduce symptoms and duration of disease only partially (8), and their effectiveness during H5N1 infection is unknown. Laver and Garman further commented that various experimental vaccines, including DNA vaccines, may be more promising for pandemic control. These statements highlight the fact that additional approaches are needed to produce effective vaccines for H5N1 or other new subtypes (9).
Vaccines using conserved components of influenza A virus can induce protection against many influenza A strains, including those of divergent subtypes. Animal studies have demonstrated potent and long-lasting heterosubtypic immunity, that is, exposure to a virus of one subtype protects against challenge infection with another subtype (10–15). The mechanisms of heterosubtypic immunity are not completely understood but likely include both T-cell immunity, in particular CD8+ cytotoxic T-lymphocytes (CTL) (16,17) and CD4+ T cells (13), as well as antibodies to conserved epitopes (18). Heterosubtypic immunity has been reported in humans (19, 20), but its effectiveness and duration are unknown. Animal studies may show ways to optimize induction of heterosubtypic immunity, which could then be tested in humans. Heterosubtypic immunity induced by virus can protect against H5N1 infection in animals (21), and human T cells specific for antigens of an H1N1 virus, including nucleoprotein (NP) and matrix (M), can lyse target cells infected with H5N1 virus (22). In addition, exposure to H9N2 virus can induce heterosubtypic protection against H5N1 challenge in chickens (23) and mice (24).
DNA vaccination can target immune responses to epitopes that are highly conserved in influenza A viruses, while avoiding the risks of live-virus vaccines. We and others have previously shown that DNA constructs expressing conserved influenza proteins induce antibody and T-cell responses and protect against H3N2 heterosubtypic challenge (25–27). Both CD4+ and CD8+ T cells play roles in this protective immunity. DNA vaccination has also been studied in the H5N1 system, although largely with constructs expressing HA. DNA constructs expressing H5 HA can protect against lethal H5N1 challenge in mice (28). In lethal challenge experiments with chickens, an H5 HA construct protected fully and a construct expressing NP of an H5N8 virus protected partially (29,30). However, DNA vaccines expressing heterosubtypic antigens have not been studied in the H5N1 system.
Studies of challenge with H5N1 viruses from the 1997 Hong Kong outbreak must take into account their phenotypic diversity. While all these viruses were highly pathogenic in chickens, two main pathogenicity phenotypes were observed in mice (31,32). Viruses of the two types were studied for histopathology, viral titers in various tissues, and lethality in mice. The H3N2 viruses A/Udorn or X31 were used for comparison in some cases. Some isolates, represented by A/Hong Kong/483/97 (HK/483), were lethal even at modest doses, replicating in multiple organs, including the brain, liver, spleen, and kidney, after intranasal administration (31), resulting in pathology of respiratory tissue and the heart, and producing immune effects (33). Other isolates, represented by A/Hong Kong/486/97 (HK/486), replicated only in the respiratory tract and were not lethal. One virus, A/Hong Kong/156/97 (HK/156), did not fit readily into either group, requiring higher doses to infect or kill mice in one of the studies (31) and showing some spread to nonrespiratory sites but more limited spread than was seen with HK/483 (31,32). HK/483 and HK/156, but not HK/486, were isolated from lethal infections in the original human cases.
In this study, we extended DNA vaccination based on conserved influenza components to heterosubtypic challenge with H5N1 virus. We investigated whether the broadly cross-reactive immunity induced by immunization of mice with DNA expressing NP and M from a mouse-adapted human H1N1 virus, A/Puerto Rico/8/34 (A/PR/8), could control infection with a range of H5N1 viruses.
Plasmid VR1012 was obtained from Vical Inc. (San Diego, CA) under a Materials Transfer Agreement. Full-length influenza genes for NP and M of A/PR/8 were prepared and inserted into VR1012 as described previously (27). The plasmid B/NP expresses the full-length NP gene from B/Ann Arbor/1/86 (B/AA), derived from a baculovirus vector generated by Rota et al. (34) and subcloned into VR1012. Plasmid DNA was prepared and tested as described (27). Endotoxin levels were <1 EU/100 μg dose.
H5N1 viruses used in this study were HK/156, HK/483, HK/485, and HK/486 (31). Other viruses used were H1N1 virus A/PR/8; reassortant virus X-31 with surface glycoproteins of A/Aichi/2/68 (H3N2) and internal proteins of A/PR/8 virus; and B/AA. The A/PR/8 and X-31 stocks were mouse adapted by passage through mouse lungs. Virus stocks were propagated in the allantoic cavity of embryonated hen eggs at 37°C for 24 hr (H5N1 viruses) or 34°C for 48–72 hr (other viruses). Fifty-percent egg infectious dose (EID50) titers and mouse infectious dose (MID50) titers were determined by serial titration in eggs or mouse lungs, respectively, and calculated by the method of Reed and Muench (35). All experiments with infectious H5N1 viruses were conducted under BSL-3+ containment, including work in animals.
BALB/c female mice were purchased from the Division of Cancer Treatment, National Cancer Institute, Frederick, Maryland, or from Jackson Laboratories, Bar Harbor, Maine. DNA was injected intramuscularly, 100 µg/mouse of each plasmid, three times at 2-week intervals, starting at 6–7 weeks of age. Approximately 1 week after the last immunization, mice were shipped from the Food and Drug Administration to Centers for Disease Control and Prevention or U.S. Department of Agriculture, allowed to rest for approximately a week, challenged under containment conditions with CO2 anesthesia, and monitored for weight loss and death. For viral titers, lung and brain tissues were collected 6 days postchallenge and frozen.
Enzyme-linked immunosorbent assay (ELISA) was performed as described previously (15) on plates coated with lysates of influenza virus–infected cells. Hemagglutination inhibition (HI) was performed by standard methods with sera pretreated with receptor-destroying enzyme (36).
Thawed tissues were homogenized in 1 mL of sterile phosphate-buffered saline. Clarified lung, brain, kidney, and nose homogenates were titrated for virus infectivity in 10-day-old embryonated eggs (EID50) from initial dilutions of 1:10 (lungs and nose) or 1:2 (brain and kidney), with positive eggs identified by hemagglutination. Detection limits were 101.2 EID50/mL for lung and nose, and 100.8 EID50/mL for brain and kidney. Enzyme-linked immunospot assay (ELISPOT) for interferon-γ (IFN-γ) secreting cells were performed as described previously (37).
For CTL assays, splenocytes were restimulated in vitro and target cells prepared as described (38). CTL activity was measured by lactate dehydrogenase (LDH) release (CytoTox 96 Non-Radioactive Cytotoxicity Assay kit G170, Promega Corp., Madison, WI). Results were calculated as:
% Lysis = Experimental-Effector Spontaneous-Target Spontaneous X 100
Mice were immunized with a mixture of plasmids encoding A/NP+A/M, intended to provide greater protection than a single antigen (27,39). Plasmid DNA without an insert is often used as a control; although we used it initially, we later prepared a construct expressing the NP gene of influenza B/AA as a specificity control. The B/AA virus is only distantly related antigenically to influenza A, and no cross-protection is seen between influenza A and B viruses. The control plasmid expressing B/NP protects against challenge with influenza B, as shown by reduction in lung viral titers (40).
A/NP+A/M DNA induced antibodies against homologous A/PR/8 proteins (geometric mean ELISA titer 761), with no cross-reactivity to influenza B proteins (all titers <20). Mice immunized with B/NP DNA had comparable titers of antibody to influenza B proteins, with no cross-reactivity to A/PR/8 proteins.
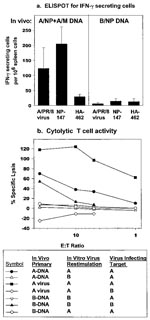
DNA immunization activated T cells in an antigen-specific manner by two measures, ELISPOT of IFN-γ secreting cells and CTL activity. Splenocytes from mice immunized with A/NP+A/M DNA generated an IFN-γ ELISPOT response when restimulated with NP147 peptide (the dominant CTL epitope in BALB/c mice), A/PR/8 virus, or concanavalin A (Con A), but not with control HA462 peptide, demonstrating antigen specificity (Figure 1a). Mice immunized with B/NP DNA did not respond to restimulation with either peptide or with A/PR/8 but did respond to Con A, indicating the cells were functional.
Antigen-specific CTL responses to DNA immunization were seen after in vitro restimulation (Figure 1b). Cells from mice immunized with A/NP+A/M DNA lysed A/PR/8-infected targets if they had been restimulated with A/PR/8 but not with B/AA. Controls immunized with B/NP DNA and restimulated with B/AA generated cytolytic activity detectable on influenza B-infected targets but not A/PR/8-infected targets.
A/NP+A/M DNA immunization was tested for protection against an H5N1 challenge virus of low virulence, HK/486. HK/486 is not lethal for mice, so control of virus replication was measured. A/NP+A/M vaccination reduced replication of HK/486 virus in the lungs approximately 17-fold, compared with viral titers in mice vaccinated with control DNA or unimmunized mice (Table, highly significant by Analysis of variance (ANOVA); see legend). As expected, infection of mice with X-31 virus induced substantial heterosubtypic immunity, reducing lung virus titers by approximately 3,000-fold compared with unvaccinated controls.
Next, we tested the ability of A/NP+A/M DNA vaccination to protect against HK/156, an H5N1 challenge virus of intermediate virulence. Mice vaccinated with A/NP+A/M DNA had only minor weight loss after challenge, while mice vaccinated with control DNA lost weight dramatically (Figure 2a). Four of mice per group were euthanized at day 6 after challenge to measure virus replication in lungs and brains. A/NP+A/M DNA immunization reduced lung titers by over two logs (approximately 500-fold, highly significant by ANOVA, Figure 2b). As expected (21), immunization with A/PR/8 virus also reduced lung titers substantially. Reductions in brain titers were not statistically significant because virus titers in the brain were low even in B/NP DNA-immunized controls.
Mice vaccinated (six per group) with A/NP+A/M DNA all survived a HK/156 challenge dose lethal to controls, as did A/PR/8-primed mice (Figure 2c). Thus, DNA vaccination with conserved components is effective not only against strains of low virulence like HK/486 but also against a lethal strain. However, A/NP+A/M DNA vaccination was not protective against challenge with 100 MID50 of highly virulent A/HK/483 (none of six mice survived). An additional experiment used 100 MID50 and a lower challenge dose of HK/483 to determine whether A/NP+A/M DNA vaccination could protect against a less extreme challenge. Challenge with 100 MID50 of HK/483 again killed all the mice vaccinated with A/NP+A/M DNA (0/8 survived). With a challenge dose of 10 MID50, four of eight mice vaccinated with A/NP+A/M DNA survived, but zero of eight given B/NP DNA and zero of eight naïve controls survived. These results suggest some protective effect, though the numbers are not statistically significant. Preliminary testing of viral titers in lung, nose, kidney, and brain at day 6 showed significant differences in the lungs and noses between A/NP+A/M immunized mice and controls, suggesting some impact of the immunization (data not shown).
A/NP+A/M DNA-vaccinated mice that survived HK/156 infection (above) were rechallenged 14 weeks later with 100 MID50 of the virulent HK/483 strain. Since mice vaccinated with control DNA had all died after HK/156 challenge, a group of naïve animals was added to the HK/483 challenge to confirm lethality of the challenge dose. All mice primed with A/NP+A/M DNA and subsequently exposed to HK/156 survived this HK/483 challenge, whereas all naïve mice died by day 8 (Figure 2d).
Anti-HA (H5) antibodies induced by HK/156 exposure might account for the protection against HK/483 infection. To assess this possibility, we tested for HI reactivity in sera from the mice after HK/156 exposure but before HK/483 challenge. All mice immunized with A/NP+A/M DNA and then exposed to HK/156 had antibodies reactive with HK/156 and cross-reactive with HK/483 and HK/485 viruses in HI, while control naïve mice had no detectable antibody (data not shown).
The most straightforward approach to vaccination against a newly emerging influenza subtype is use of inactivated virus or recombinant HA. However, if antigenically matched vaccines were not available in time or in sufficient quantity, other options would be important. Our study examines one of these.
DNA vaccination using genes for conserved antigens could have several advantages. The constructs could be available at any time. Plasmid production in bacteria is more consistent than growth of different viruses in eggs, and a cold chain might not be necessary for storage. To explore the usefulness of this approach, we studied the ability of NP+M DNA vaccines derived from A/PR/8 (H1N1) to protect against H5N1 challenge.
Vaccination with A/NP+A/M DNA readily induced antigen-specific antibody and T-cell responses, as shown previously (26,27). We investigated the potential for A/NP+A/M DNA vaccination to control infection by H5N1 viruses of modest (HK/486), intermediate (HK/156), and very high (HK/483) virulence phenotypes. Upon challenge with HK/486, a strain that is not lethal in mice, lung titers were reduced approximately 17-fold. In previous work, even a 5- to 10-fold reduction in peak lung virus titers correlated with immunity protective against lethal challenge (14), so a 17-fold reduction and the accompanying difference in kinetics of viral clearance could alter biologic outcomes. In a test of its effectiveness, the vaccination provided benefit in the case of lethal challenge with HK/156, resulting in 100% survival and minimal morbidity as measured by weight loss, while unvaccinated controls demonstrated large weight losses and 100% death rates. After surviving HK/156 infection, the mice were resistant to lethal HK/483 challenge. Antibodies to HK/156 were demonstrated by HI to be present and cross-reactive with HK/483 virus before HK/483 challenge, which might account for the protection against HK/483. Of mice vaccinated only with A/NP+A/M DNA, half survived challenge with a dose of HK/483 lethal to all controls. While not statistically significant, the trend suggests some impact from immunization.
Regarding immune mechanisms of protection by A/NP+A/M DNA vaccination, candidates include CTL specific for NP (17) and antibodies to the N-terminal portion of M2 (18). Containing an infection with the kinetics of HK/483 may be difficult because it reaches near peak titers in as little as 24 hours. Only neutralizing antibody may be effective that early. Antigen presentation and reactivation of T-cell effectors take several days. However, when T cells specific for viral antigens are expanded substantially, they can reduce replication of highly lethal influenza viruses and clear infection more rapidly (41).-
Comparing amino acid sequences in GenBank from viruses of five subtypes, NPs were >90% identical, with considerable conservation of known dominant CTL epitopes. M1 sequences were >94% conserved, while M2 sequences varied somewhat more. However, not all protective epitopes are known, and even single mutations can alter protective epitopes. Therefore, studies like the present one are necessary for establishing the range of virus strains against which a vaccine can work.
H5 viruses differ in virulence, and one cannot predict which strain might emerge in a future pandemic. With the threat of a pandemic and suboptimal existing vaccine candidates, new approaches to influenza vaccination should be considered. Our results suggest that DNA vaccination with conserved components has the potential to ameliorate disease caused by H5N1 viruses. The immunity induced by this mode of DNA vaccination does not completely prevent infection but passed the stringent test of protecting against lethal H5N1 challenge. Vaccines inducing neutralizing antibody could be administered subsequently to confer immunity against even the most virulent strains. In the absence of an antigenically matched HA-based vaccine, this approach might be useful as a first line of defense against a rapidly spreading influenza pandemic and should be further explored.
Dr. Epstein is chief of the Laboratory of Immunology and Developmental Biology, Division of Cellular and Gene Therapies, Office of Therapeutics Research and Review, Center for Biologics Evaluation and Research, and the Food and Drug Administration. Her research expertise is in immunology, and she also has expertise in regulatory review and policy work in the area of gene therapy. Her research program studies mechanisms of protective immunity against influenza virus infection in mouse models, with emphasis on heterosubtypic immunity.
Acknowledgments
We also appreciate the following contributions: Zhihong Wang of the USDA/ARS/Southeast Poultry Research Laboratory participated in the challenge experiment using two different doses of HK/483. Thomas Rowe performed the serum HI testing; Judy Beeler, Steven Bauer, and Karen Elkins provided critical reviews of the manuscript; Anthony Ferrine and other animal care staff provided excellent animal care; Yumiko Matsuoka assisted in subcloning the influenza B/NP gene; Direct Services Trucking transported mice safely from Bethesda to Atlanta during a snowstorm; and Terrence M. Tumpey conducted portions of the experimental work reported in this paper.
This work was supported in part by a grant from the National Vaccine Program to S.L.E.
References
- Subbarao K, Klimov A, Katz J, Regnery H, Lim W, Hall H, Characterization of an avian influenza A (H5N1) virus isolated from a child with a fatal respiratory illness. Science. 1998;279:393–6. DOIPubMedGoogle Scholar
- Yuen KY, Chan PKS, Peiris M, Tsang DNC, Que TL, Shortridge KF, Clinical features and rapid viral diagnosis of human disease associated with avian influenza A H5N1 virus. Lancet. 1998;351:467–71. DOIPubMedGoogle Scholar
- Centers for Disease Control and Prevention. Updated recommendations from the Advisory Committee on Immunization Practices in response to delays in supply of influenza vaccine for the 2000–01 season. MMWR Morb Mortal Wkly Rep. 2001;49:888–92.
- Treanor JJ, Wilkinson BE, Masseoud F, Hu-Primmer J, Battaglia R, O'Brien D, Safety and immunogenicity of a recombinant hemagglutinin vaccine for H5 influenza in humans. Vaccine. 2001;19:1732–7. DOIPubMedGoogle Scholar
- Nicholson KG, Colegate AE, Podda A, Stephenson I, Wood J, Ypma E, Safety and antigenicity of non-adjuvanted and MF59-adjuvanted influenza A/Duck/Singapore/97 (H5N3) vaccine: a randomized trial of two potential vaccines against H5N1 influenza. Lancet. 2001;357:1937–43. DOIPubMedGoogle Scholar
- Hatta M, Gao P, Halfmann P, Kawaoka Y. Molecular basis for high virulence of Hong Kong H5N1 influenza A viruses. Science. 2001;293:1840–2. DOIPubMedGoogle Scholar
- Laver G, Garman E. The origin and control of pandemic influenza. Science. 2001;293:1776–7. DOIPubMedGoogle Scholar
- Nicholson KG, Aoki FY, Osterhaus AE, Trottier S, Carewicz O, Mercier CH, Efficacy and safety of oseltamivir in treatment of acute influenza: a randomized controlled trial. Lancet. 2000;355:1845–50. DOIPubMedGoogle Scholar
- Peiris M, Yuen KY, Leung CW, Chan KH, Ip PS, Lai RM, Human infection with influenza H9N2. Lancet. 1999;354:916–7. DOIPubMedGoogle Scholar
- Schulman JL, Kilbourne ED. Induction of partial specific heterotypic immunity in mice by a single infection with influenza A virus. J Bacteriol. 1965;89:170–4.PubMedGoogle Scholar
- Yetter RA, Barber WH, Small PA Jr. Heterotypic immunity to influenza in ferrets. Infect Immun. 1980;29:650–3.PubMedGoogle Scholar
- Mbawuike IN, Six HR, Cate TR, Couch RB. Vaccination with inactivated influenza A virus during pregnancy protects neonatal mice against lethal challenge by influenza A viruses representing three subtypes. J Virol. 1990;64:1370–4.PubMedGoogle Scholar
- Liang S, Mozdzanowska K, Palladino G, Gerhard W. Heterosubtypic immunity to influenza type A virus in mice: Effector mechanisms and their longevity. J Immunol. 1994;152:1653–61.PubMedGoogle Scholar
- Epstein SL, Lo C-Y, Misplon JA, Lawson CM, Hendrickson BA, Max EE, Mechanisms of heterosubtypic immunity to lethal influenza A virus infection in immunocompetent, T cell-depleted, β2-microglobulin-deficient, and J chain-deficient mice. J Immunol. 1997;158:1222–30.PubMedGoogle Scholar
- Benton KA, Misplon JA, Lo C-Y, Brutkiewicz RR, Prasad SA, Epstein SL. Heterosubtypic immunity to influenza A virus in mice lacking either IgA, all Ig, NKT cells, or γδ T cells. J Immunol. 2001;166:7437–45.PubMedGoogle Scholar
- Lukacher AE, Braciale VL, Braciale TJ. In vivo effector function of influenza virus-specific cytotoxic T lymphocyte clones is highly specific. J Exp Med. 1984;160:814–26. DOIPubMedGoogle Scholar
- Taylor PM, Askonas BA. Influenza nucleoprotein-specific cytotoxic T-cell clones are protective in vivo. Immunol. 1986;58:417–20.
- Neirynck S, Deroo T, Saelens X, Vanlandschoot P, Jou WM, Fiers W. A universal influenza A vaccine based on the extracellular domain of the M2 protein. Nat Med. 1999;5:1157–63. DOIPubMedGoogle Scholar
- Slepushkin AN. The effect of a previous attack of A1 influenza on susceptibility to A2 virus during the 1957 outbreak. Bull World Health Organ. 1959;20:297–301.PubMedGoogle Scholar
- Sonoguchi T, Naito H, Hara M, Takeuchi Y, Fukumi H. Cross-subtype protection in humans during sequential overlapping and/or concurrent epidemics caused by H3N2 and H1N1 influenza viruses. J Infect Dis. 1985;151:81–8.PubMedGoogle Scholar
- Tumpey TM, Renshaw M, Clements JD, Katz JM. Mucosal delivery of inactivated influenza vaccine induces B-cell-dependent heterosubtypic cross-protection against lethal influenza a H5N1 virus infection. J Virol. 2001;75:5141–50. DOIPubMedGoogle Scholar
- Jameson J, Cruz J, Terajima M, Ennis FA. Human CD8+ and CD4+ T lymphocyte memory to influenza a viruses of swine and avian species. J Immunol. 1999;162:7578–83.PubMedGoogle Scholar
- Seo SH, Webster RG. Cross-reactive, cell-mediated immunity and protection of chickens from lethal H5N1 influenza virus infection in Hong Kong poultry markets. J Virol. 2001;75:2516–25. DOIPubMedGoogle Scholar
- O'Neill E, Krauss SL, Riberdy JM, Webster RG, Woodland DL. Heterologous protection against lethal A/HongKong/156/97 (H5N1) influenza virus infection in C57BL/6 mice. J Gen Virol. 2000;81:2689–96.PubMedGoogle Scholar
- Rhodes GH, Dwarki VJ, Abai AM, Felgner J, Felgner PL, Gromkowski SH, Injection of expression vectors containing viral genes induces cellular, humoral, and protective immunity. In: Chanock RM, Brown F, Ginsberg HS, Norrby E, editors. Vaccines 93. Cold Spring Harbor (NY): Cold Spring Harbor Laboratory Press; 1993: p. 137–41.
- Ulmer JB, Donnelly JJ, Parker SE, Rhodes GH, Felgner PL, Dwarki VJ, Heterologous protection against influenza by injection of DNA encoding a viral protein. Science. 1993;259:1745–9. DOIPubMedGoogle Scholar
- Epstein SL, Stack A, Misplon JA, Lo C-Y, Mostowski H, Bennink J, Vaccination with DNA encoding internal proteins of influenza virus does not require CD8+ CTL: either CD4+ or CD8+ T cells can promote survival and recovery after challenge. Int Immunol. 2000;12:91–101. DOIPubMedGoogle Scholar
- Kodihalli S, Goto H, Kobasa DL, Krauss S, Kawaoka Y, Webster RG. DNA vaccine encoding hemagglutinin provides protective immunity against H5N1 influenza virus infection in mice. J Virol. 1999;73:2094–8.PubMedGoogle Scholar
- Kodihalli S, Haynes JR, Robinson HL, Webster RG. Cross-protection among lethal H5N2 influenza viruses induced by DNA vaccine to the hemagglutinin. J Virol. 1997;71:3391–6.PubMedGoogle Scholar
- Kodihalli S, Kobasa DL, Webster RG. Strategies for inducing protection against avian influenza A virus subtypes with DNA vaccines. Vaccine. 2000;18:2592–9. DOIPubMedGoogle Scholar
- Lu XH, Tumpey TM, Morken T, Zaki SR, Cox NJ, Katz JM. A mouse model for the evaluation of pathogenesis and immunity to influenza A (H5N1) viruses isolated from humans. J Virol. 1999;73:5903–11.PubMedGoogle Scholar
- Gao P, Watanabe S, Ito T, Goto H, Wells K, McGregor M, Biological heterogeneity, including systemic replication in mice, of H5N1 influenza A virus isolates from humans in Hong Kong. J Virol. 1999;73:3184–9.PubMedGoogle Scholar
- Tumpey TM, Lu XH, Morken T, Zaki SR, Katz JM. Depletion of lymphocytes and diminished cytokine production in mice infected with a highly virulent influenza A (H5N1) virus isolated from humans. J Virol. 2000;74:6105–16. DOIPubMedGoogle Scholar
- Rota PA, Black RA, De BK, Harmon MW, Kendal AP. Expression of influenza A and B virus nucleoprotein antigens in baculovirus. J Gen Virol. 1990;71:1545–54. DOIPubMedGoogle Scholar
- Hawkes RA. General principles underlying laboratory diagnosis of viral infections. In: Lennette EH, Schmidt NJ, editors. Diagnostic procedures for viral, rickettsial and chlamydial infections. Washington: American Public Health Association; 1979. p. 3–48.
- Kendal AP, Skehel JJ, Pereira MS. Concepts and procedures for laboratory-based influenza surveillance. B17– B35. Atlanta: Centers for Disease Control; 1982.
- Sambhara S, Switzer I, Kurichh A, Miranda R, Urbanczyk L, James O, Enhanced antibody and cytokine responses to influenza viral antigens in perforin-deficient mice. Cell Immunol. 1998;187:13–8. DOIPubMedGoogle Scholar
- Deng YP, Yewdell JW, Eisenlohr LC, Bennink JR. MHC affinity, peptide liberation, T cell repertoire, and immunodominance all contribute to the paucity of MHC class I- restricted peptides recognized by antiviral CTL. J Immunol. 1997;158:1507–15.PubMedGoogle Scholar
- Donnelly JJ, Friedman A, Martinez D, Montgomery DL, Shiver JW, Motzel SL, Preclinical efficacy of a prototype DNA vaccine: enhanced protection against antigenic drift in influenza virus. Nat Med. 1995;1:583–7. DOIPubMedGoogle Scholar
- Epstein SL, Stack A, Misplon JA, Lo C-Y, Mostowski H, Bennink J, Vaccination with DNA encoding conserved influenza viral proteins. Proceedings of the meeting, Options for the control of influenza IV, Crete, Greece 23-28 Sep 2000. Amsterdam: Elsevier Science; 2001.
- Christensen JP, Doherty PC, Branum KC, Riberdy JM. Profound protection against respiratory challenge with a lethal H7N7 influenza A virus by increasing the magnitude of CD8+ T-cell memory. J Virol. 2000;74:11690–6. DOIPubMedGoogle Scholar
Figures
Table
Cite This ArticleTable of Contents – Volume 8, Number 8—August 2002
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
Suzanne Epstein, DCGT, OTRR, CBER, FDA, 1401 Rockville Pike, HFM-521, Rockville, MD 20852-1448, USA; fax: 301-827-0449;
Top