Volume 9, Number 3—March 2003
Dispatch
Isolation of Escherichia coli O157:H7 from Intact Colon Fecal Samples of Swine1
Cite This Article
Citation for Media
Abstract
Escherichia coli O157:H7 was recovered from colon fecal samples of pigs. Polymerase chain reaction confirmed two genotypes: isolates harboring the eaeA, stx1, and stx2 genes and isolates harboring the eaeA, stx1, and hly933 genes. We demonstrate that swine in the United States can harbor potentially pathogenic E. coli O157:H7.
During the past two decades, disease caused by Escherichia coli O157:H7 has been increasing (1). Currently, the Centers for Disease Control and Prevention estimates that E. coli O157:H7 causes an average of 500 outbreaks that affect >73,000 persons and result in >61 deaths each year in the United States (2). The epidemiology of E. coli O157:H7 has become an important research topic as manure harboring E. coli O157:H7 is dispersed, and soil, food, and water are cross-contaminated with feces containing E. coli O157:H7 (1,3). Although cattle feces are the most important source of E. coli O157:H7, the need to evaluate the presence of E. coli O157:H7 in the feces of other animal species has been recognized (1). The presence of E. coli O157:H7 in swine feces has been reported in Japan (4), Norway (5), and Chile (6); however, to date, E. coli O157:H7 has not been reported in swine in the United States.
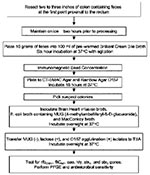
Colon samples were collected at a cooperating swine slaughter facility from 305 swine carcasses during evisceration. Two to three inches of distal colon that contained feces at the first point proximal to the rectum was resected and maintained on ice for approximately 2 hours before processing (Figure). Ten grams of feces from each colon was transferred to filter-lined sterile plastic bags. One hundred milliliters of brilliant green bile broth (Difco Laboratories, Detroit, MI), prewarmed to 37°C, was added to each filter stomacher bag containing feces and incubated at 37°C for 6 h with shaking (150 rpm) (7). After enrichment, 1.0-mL aliquots were processed by using Dynabeads anti–E. coli O157 (Dynal Biotech, Oslo, Norway), according to manufacturers’ instructions with modification. Bead/sample suspensions were incubated at room temperature for 30 min with continuous mixing on a Bellco roller drum (Bellco Glass, Inc., Vineland, NJ) before plating onto sorbitol MacConkey (SMAC; Difco Laboratories), cefixime/tellurite (CT; cefixime-tellurite supplement, Dynal Biotech)‑SMAC agars, and rainbow agar O157 (Biolog, Inc., Hayward, CA). Black colonies from rainbow agar O157 and sorbitol-negative colonies from CT-SMAC and SMAC agars were tested for the absence of ß-glucuronidase and the ability to ferment lactose by using E. coli broth containing 4-methlumbelliferyl-β-D-glucuronide (MUG) (EC medium with MUG; Difco Laboratories) and MacConkey broth (Difco Laboratories), respectively. Lactose-positive/MUG-negative isolates were serotyped by using the RIM E. coli O157:H7 Latex Test (Remel, Lenexa, KS). Up to 10 E. coli O157 latex agglutination–positive isolates per colon fecal sample were tested for the presence of the rfbO157 gene by using polymerase chain reaction (PCR) (8). Isolates positive for the rfbO157 gene were further characterized for the presence of genes encoding for the H7 flagellar protein (fliCH7), Shiga toxin 1 (stx1), Shiga toxin 2 (stx2), intimin protein (eaeA), and hemolysin (hly933) (9). We conducted further analysis using antimicrobial resistance patterns, pulsed-field gel electrophoresis (PFGE), and ribotyping on all E. coli O157 PCR–positive isolates containing fliCH7, stx1, stx2, eaeA, or hly933. However, for tabulation purposes, each sample ultimately contributed one isolate. When fliCH7, stx1, stx2, eaeA, or hly933 was not detected in PCR-confirmed E. coli O157 isolates, further analysis was performed on only one E. coli O157 isolate per colon sample.
E. coli O157 isolates were tested for susceptibility to 17 antimicrobial agents (amikacin, amoxicillin/clavulanic acid, ampicillin, apramycin, cefoxitin, ceftriaxone, cephalothin, chloramphenicol, ciprofoxacin, gentamicin, imipenem, kanamycin, nalidixic acid, streptomycin, sulfamethoxazole, tetracycline, and trimethoprim/sulfamethoxazole) as described (10) by using a custom-made semiautomated broth microdilution assay (Sensititre, Trek Diagnostics, Westlake, OH). Imipenem was used at concentrations of 0.25–8.0 μg with the following breakpoints: sensitive (<4) and resistant (≥16).
For PFGE, DNA was digested with 50 U XbaI (Invitrogen Corp., Carlsbad, CA) for 4 h at 37°C. PFGE was performed by using a CHEF Mapper XA system (Bio-Rad, Hercules, CA) at 14°C with pulses ramping from 2.16 s to 63.8 s over 18 h. PFGE patterns were evaluated visually, and isolates were assigned to the same pulsotype when exhibiting a difference of <3 bands from the index isolate. Ribotyping of the E. coli O157 isolates was done by using a RiboPrinter (Qualicon, Inc., Wilmington, DE) as described in the user’s manual. Restriction digests were performed on E. coli O157 isolates by using the EcoRI enzyme (Qualicon, Inc.).
A total of 305 colon samples were randomly collected on 8 different days over a 6-month period as follows: collection day 1 (February 16, 2001), n=5; collection day 2 (March 8, 2001), n=20; collection day 3 (March 22, 2001), n=40; collection day 4 (April 20, 2001), n=40; collection day 5 (May 4, 2001), n=50; collection day 6 (May 16, 2001), n=50; collection day 7 (June 20, 2001), n=50; and collection day 8 (July 10, 2001), n=50 (Table). Eighteen (5.9%) of the 305 colon samples had isolates positive for rfbO157. Isolates from 6 of these 18 colon samples also contained fliCH7. Two gene combinations based on the presence or absence of stx1, stx2, eae, and hly933 were detected in these E. coli O157:H7 PCR-confirmed isolates. The stx1, eaeA, and hly933 virulence pattern was detected in two isolates (isolates1 and 2) from two of the five colon samples collected on February 16, 2001, and the stx1, stx2, and eaeA virulence pattern was detected in 22 isolates (isolates 6–27) from 4 of the 50 colon samples collected on May 4, 2001. None of the E. coli O157:H7 isolates recovered contained all four of the virulence genes (stx1, stx2, eaeA, and hly933). None of the E. coli O157:non-H7 isolates (isolates 3–5, 28–36) in the present study contained stx1, stx2, eae, or hly genes. Non–Shiga toxin–producing E. coli O157:non-H7 isolates have been previously isolated from the feces of pigs (11,12). For slaughterhouse visits on March 8, March 22, June 20, and July 10, 2001, E. coli O157 and E. coli O157:H7 were not recovered from any of the colons sampled.
All E. coli O157:H7 isolates recovered in this study were sensitive to the antimicrobial agents tested, with the exception of one isolate (isolate 15) that was resistant to streptomycin. This isolate was recovered from a colon from which a pan-sensitive E. coli O157:H7 was also recovered. The antimicrobial sensitivity pattern of the E. coli O157:non-H7 isolates was more varied than that of the E. coli O157:H7 isolates with five different susceptibility patterns noted. Only one of the E. coli O157:non-H7 isolates was pan-sensitive. These data are similar to previous reports in which antimicrobial resistance among E. coli O157 non–Shiga toxin–producing isolates was higher than that of Shiga toxin–producing E. coli O157 isolates (11).
As previously shown, ribotyping did not discriminate among isolates within the E. coli O157:H7 serotype (13). Additionally, the E. coli O157:non-H7 isolates were indistinguishable from one another. Four PFGE profiles were noted. The E. coli O157:H7 isolates obtained from colon 1 and colon 2 on February 16, 2001, exhibited the PFGE type 1 pattern, whereas the E. coli O157:H7 isolates obtained from four colons on May 4, 2001 exhibited the PFGE type 2 pattern. The E. coli O157:non-H7 isolates obtained on April 20, 2001, and May 16, 2001, exhibited PFGE patterns 3 and 4, respectively.
Results from this study demonstrate that pigs in the United States can harbor E. coli O157:H7. The recovery rate of E. coli O157:H7 from colon fecal samples of pigs reported in this study was 2.0% (6/305). Previous attempts to isolate E. coli O157:H7 from swine feces in the United States have been unsuccessful (12,14). Use of more appropriate methods for sampling, processing, and culturing swine feces may have accounted for the ability to recover and isolate E. coli O157:H7 from swine feces in our study. For example, samples were obtained from the colon, transported on ice, and processed within 2 h of collection. The absence of antibiotics in our enrichment step may have also facilitated the recovery of E. coli O157:H7 from swine feces. Furthermore, although direct comparisons cannot be made between cattle studies, the recovery rate of Shiga toxin–producing E. coli O157 from cattle feces has improved over the past 10 years. This is most likely due to more conducive sampling procedures, culture practices, and detection methods than an increase in true carriers. The detection of E. coli O157 in swine feces has previously been based on the isolation techniques used for the recovery of E. coli O157 from cattle feces. The difficulty in detecting E. coli O157 from swine feces may in part be attributable to differences in the physiologic environment between swine and cattle feces. More appropriate isolation techniques may still be discovered for detecting E. coli O157 in swine.
Although our recovery rates of E. coli O157:H7 from swine are similar to recovery rates in Japan (4), we recovered a genotype in addition to the stx1, stx1, and eaeA genotype: the stx1, eaeA, and hly933 genotype. In Norway, the recovery rate (0.1%) of E. coli O157:H7 from pig feces was much lower (5). Isolates recovered from Norway possessed the stx2 and eae genes only; however, the presence of the hly gene was not determined (5).
The ability to produce one or more Shiga toxins is an important virulence characteristic of E. coli O157:H7 (1). However, production of Shiga toxins alone may not be sufficient for E. coli O157:H7 to be pathogenic (1). Other virulence factors such as the intimin protein (involved in the attachment of the E. coli O157 to enterocytes), the presence of a plasmid-encoded hemolysin, or both, are important in the pathophysiology of hemorrhagic disease (1). E. coli O157:H7 isolates recovered in this study possessed either two virulence factors, eaeA and hly933, in addition to stx1 or one virulence factor, eaeA, in addition to stx1 and stx2. These isolates can potentially cause disease and should be considered pathogenic to humans. Since human E. coli O157:H7 clinical isolates contain the stx1, stx2, eaeA, and hly genes, the human pathogenicity of E. coli O157:H7 isolates from pigs that lack the hly gene requires further study.
The clonal nature of the isolates that were isolated on a particular day suggests transmission of E. coli O157 between pigs. Unfortunately, we did not have access to information concerning the source of the pigs from which the samples were collected, the number of pigs slaughtered from a given farm, or the holding facilities and grouping of the pigs before slaughter. Therefore, we do not know whether E. coli O157 transmission between pigs occurred on the farm, in transit, or while the pigs were in a holding pen at the slaughterhouse.
This study did not permit inferences of E. coli O157:H7 isolation rates with respect to the season, nor can inferences of E. coli O157:H7 isolation rates be made with respect to swine or herd prevalence. The relatively low recovery rate of E. coli O157:H7 from swine feces compared to cattle feces warrants further study to determine the significance and prevalence of E. coli O157:H7 in swine and if different enrichment and isolation methods would have an impact on the recovery of E. coli O157:H7 from swine feces. In addition, future studies should be conducted to determine the occurrence of E. coli O157 on swine hides, in swine mouths, and in swine stomachs.
Dr. Feder is a research microbiologist with the U.S. Department of Agriculture. Her research interests include diagnostic microbiology, molecular epidemiology, and assay development of zoonotic pathogens, primarily Escherichia coli O157:H7 and Salmonella.
Acknowledgment
We thank Connie Briggs and. Lori Bagi for their technical support and assistance and Takiyah Ball and Jovita Hermosillo for their expertise in antimicrobial susceptibility testing.
References
- Mead PS, Slutsker L, Dietz V, McCaig LF, Bresee JS, Shapiro C, Food-related illness and death in the United States. Emerg Infect Dis. 1999;5:607–25. DOIPubMedGoogle Scholar
- Charatan F. New York outbreak of E. coli poisoning affects 1000 and kills two. BMJ. 1999;86:873.PubMedGoogle Scholar
- Nakazawa M, Akiba M. Swine as a potential reservoir of Shiga toxin-producing Escherichia coli O157:H7 in Japan. Emerg Infect Dis. 1999;5:833–4. DOIPubMedGoogle Scholar
- Johnsen G, Wasteson Y, Heir E, Berget OI, Herikstad H. Escherichia coli O157:H7 in faeces from cattle, sheep and pigs in the southwest part of Norway during 1998 and 1999. Int J Food Microbiol. 2001;65:193–200. DOIPubMedGoogle Scholar
- Rios M, Prado V, Trucksis M, Arellano C, Borie C, Alexandre M, Clonal diversity of Chilean isolates of enterohemorrhagic Escherichia coli from patients with hemolytic-uremic syndrome, asymptomatic subjects, animal reservoirs, and food products. J Clin Microbiol. 1999;37:778–81.PubMedGoogle Scholar
- Gray JT, Smith D, Moxley R, Rolfes C, Hungerford L, Younts S, Comparison of oral and rectal sampling to classify the Escherichia coli O157:H7 status of pens of feedlot cattle. Program and abstracts of American Society of Microbiologists annual meeting, May 21–22, Los Angeles, California. Washington: American Society for Microbiology; 2002:524–5.
- Paton AW, Paton JC. Detection and characterization of Shiga toxigenic Escherichia coli by using multiplex PCR assays for stx1, stx2, eaeA, enterohemorrhagic E. coli hlyA, rfbO111, and rfbO157. J Clin Microbiol. 1998;36:598–602.PubMedGoogle Scholar
- Fratamico PM, Bagi LK, Pepe T. A multiplex polymerase chain reaction assay for rapid detection and identification of Escherichia coli O157:H7 in foods and bovine feces. J Food Prot. 2000;63:1032–7.PubMedGoogle Scholar
- Fedorka-Cray P. National antimicrobial resistance monitoring system annual report. Available from: http://www.ars-grin.gov/ars/SoAtlantic/Athens/arru/narms_2000/2000_tables/2000_table01.pdf
- Schroeder CM, Zhao C, DebRoy C, Torcolini J, Zhao S, White DG, et al. Antimicrobial resistance of Escherichia coli O157 isolated from humans, cattle, swine, and food. Appl Environ Microbiol. 2002;68:576–81. DOIPubMedGoogle Scholar
- Feder I, Gray J, Pearce R, Fratamico P, Bush E, Wallace FM, et a. National animal health monitoring system swine 2000: a surveillance study of Escherichia coli O157 in swine. International Association for Food Protection annual meeting program and abstracts, June 29–July 3, 2001.
- Martin IE, Tyler SD, Tyler KD, Khakhria R, Johnsen WM. Evaluation of ribotyping as epidemiologic tool for typing Escherichia coli serogroup O157 isolates. J Clin Microbiol. 1996;34:720–3.PubMedGoogle Scholar
- Bush EUS. swine herd appears free of Escherichia coli O157:H7. Food Safety Digest 1997;4.
Figure
Table
Cite This Article1Presented in part at the 102nd Annual Meeting of the American Society of Microbiologists, Salt Lake City, Utah, May 2002.
Table of Contents – Volume 9, Number 3—March 2003
| EID Search Options |
|---|
|
|
|
|
|
|
Please use the form below to submit correspondence to the authors or contact them at the following address:
Ingrid E. Feder, Microbial Biophysics and Residue Chemistry and Core Technologies, USDA, Agricultural Research Service, Eastern Regional Research Center, 600 East Mermaid Lane, Wyndmoor, PA 19038, USA; fax: 215-836-3742
Top