Volume 10, Number 1—January 2004
Research
Ecologic and Geographic Distribution of Filovirus Disease
Cite This Article
Citation for Media
Abstract
We used ecologic niche modeling of outbreaks and sporadic cases of filovirus-associated hemorrhagic fever (HF) to provide a large-scale perspective on the geographic and ecologic distributions of Ebola and Marburg viruses. We predicted that filovirus would occur across the Afrotropics: Ebola HF in the humid rain forests of central and western Africa, and Marburg HF in the drier and more open areas of central and eastern Africa. Most of the predicted geographic extent of Ebola HF has been observed; Marburg HF has the potential to occur farther south and east. Ecologic conditions appropriate for Ebola HF are also present in Southeast Asia and the Philippines, where Ebola Reston is hypothesized to be distributed. This first large-scale ecologic analysis provides a framework for a more informed search for taxa that could constitute the natural reservoir for this virus family.
The natural maintenance cycles of filoviruses (Order Mononegavirales, family Filoviridae) are unknown (1).1Although dynamics of filoviruses as causes of epidemic diseases among humans, great apes, and other primates have been described in detail (2–13), the natural reservoir, mode of transmission to hominids and pongids (humans, gorillas, and chimpanzees), and temporal dynamics remain unclear. Diverse taxa have been suggested as potential reservoirs, including bats, rodents, arthropods, and plants (14–18).
Two observations provide clues about the nature of the host-virus relationship. First, filovirus transmission to humans is not common, and most occurrences can be traced to a single index case (2,6,19) (exceptions occur—e.g., the Durba Marburg outbreak appears to have involved multiple independent infections of humans from a reservoir population presumably associated with a mine). We assume that introductions to nonhuman hominid populations also generally begin with single index cases, but this hypothesis is more difficult to investigate. This rarity argues against a common arthropod vector for transmission: if anthrophilic arthropod vectors were to carry filoviruses, multiple index cases would be more common, as many hominids in an area would have the opportunity for infection. In addition, filoviruses generally do not replicate in arthropods or arthropod cell lines, leading several authors to speculate on more incidental modes of transfer (e.g., direct contact) (20).
Second, filoviruses show clear geographically related phylogeographic structure. Viruses and subtypes from particular geographic areas cluster together phylogenetically, even when occurrences from different years are studied. This phylogeographic structure suggests a stable host-parasite relationship, in which viruses are maintained in permanent local-regional pools. This host would not experience high death rates, as primates do (7,9); evolution of avirulence in long-term host-parasite relationships is expected on theoretical grounds (21).
Searches for the natural reservoir of filoviruses have taken several paths. Epidemiologic studies designed to trace lineages of transmission in outbreaks have identified index cases but have not succeeded in specifying the mode of “jump” to hominids (2,6,19). Testing large numbers of organisms from the vicinity of outbreaks has failed to identify even a single nonhominid infection (14–16). Finally, laboratory tests of reservoir competence of species have documented the following: 1) no, or very limited, infection of plants or arthropods; 2) a single marginally successful infection of snakes but with very low levels of virus circulation; 3) successful infection of bats and possibly rodents; and 4) frequent successful, but fatal, infection in nonhuman primates (e.g., Chlorocebus, Macaca) (1,17,18). While these investigations have shed some light, they have not provided convincing evidence for a particular reservoir.
An unexplored approach to identifying the natural reservoir of filoviruses is large-scale ecologic and geographic comparisons to detect patterns of co-occurrence and codistribution of viruses with potential hosts. This approach has been applied successfully to identifying reservoir rodent species for Chagas disease (22). Our general approach is as follows: 1) to understand the large-scale ecology and geography of disease occurrences by using ecologic niche modeling (23), and 2) to compare these characteristics with ecologic and geographic patterns of potential reservoirs. Here, we address the first step and document broad-scale ecologic and geographic patterns in filovirus-associated HF occurrences.
Distributional data for filovirus-associated HF occurrences in hominids were accumulated from the literature (Table). Occurrences of unknown origin were excluded from analysis, but when reasonable guesses could be made as to point or general area of origin (e.g., 1995 outbreak of hemorrhagic fever due to Ebola Ivory Coast as originating at Plibo, Liberia), they were included. All occurrences were georeferenced (available from: URL: http://www.calle.com/world) to the nearest 0.001°. Although assigned geographic coordinates may not fix the exposure point precisely, they represent our best guess as to its position and are likely to be representative of the coarse-scale ecologic conditions. (The relatively crude spatial resolution at which analyses were conducted makes some error in georeferencing irrelevant.)
Ecologic niches and potential geographic distributions were modeled by using the Genetic Algorithm for Rule-set Prediction (GARP) (30–32) (available from: URL: http://www.lifemapper.org/desktopgarp/). In general, GARP focuses on modeling ecologic niches (the conjunction of ecologic conditions wherein a species can maintain populations without immigration) (33). Specifically, GARP relates ecologic characteristics of occurrence points to those of points sampled randomly from the rest of the study region, developing a series of decision rules that best summarize factors associated with presence (23).
Occurrence points are divided evenly into training (for model building) and test (for model evaluation) datasets. GARP works in an iterative process of rule selection, evaluation, testing, and incorporation or rejection: a method is chosen from a set of possibilities (e.g., logistic regression, bioclimatic rules) and applied to the training data to develop or evolve a rule. Predictive accuracy is evaluated on the basis of the test data. Rules may evolve in ways that mimic DNA evolution (e.g., point mutations, deletions). Change in predictive accuracy between iterations is used to evaluate whether particular rules should be incorporated into the model; the algorithm runs 1,000 iterations or until convergence. Model quality was evaluated through independent test dataset reserved prior to modeling; a chi-square test was used to compare observed success in predicting the distribution of test points with that expected under a random model (proportional area predicted present provides an estimate of occurrence points correctly predicted, were the prediction random with respect to the distribution of the test points).
To characterize environments, we used 11 GIS coverages summarizing elevation, slope, aspect, flow direction, flow accumulation, and tendency to pool water (from the USGS Hydro-1K dataset [available from: URL: http://edcdaac.usgs.gov/gtopo30/hydro/), and climate characteristics, including daily temperature range; mean annual precipitation; maximum, minimum, and mean annual temperatures; solar radiation; frost days; wet days; and vapor pressure (1960–1990; Intergovernmental Panel on Climate Change [available from: URL: http://www.ipcc.ch/]). These coverages are worldwide and provide a consistent view of ecologic variation across regions studied. GARP’s predictive ability has been tested under diverse circumstances (22,23,34–47).
To optimize model performance, we developed 100 replicate models of ecologic niches based on independent random subsamples from available occurrences. We chose a “best subset” of these models on the basis of optimal error distributions for individual replicate models (34): median area predicted across all replicate modes was calculated, and the 20 models with predicted areas closest to the median were chosen for further consideration. These geographic predictions were combined to provide a summary of potential geographic distributions. Projection of the Africa-based rule-sets onto maps of Asia and the Pacific provided hypotheses of potential distributional areas in other regions (46).
To permit visualization of the ecologic dimensions of models, we combined best-subsets predictions with maps of the ecologic parameters used to build them in a GIS environment (COMBINE in ArcView 3.2). The resulting dataset represents unique combinations of environments and predictions; its attributes table provides the model prediction for all environmental combinations to permit visualization of ecologic variation. We also compared ecologic conditions inside and outside of the modeled Ebola HF distribution within 11 regularly spaced circular windows (radius 50 km); comparisons were summarized through Mann-Whitney U-statistics, permitting a nonparametric visualization of the strength of association of each ecologic dimension (temperature, precipitation, elevation) with the range limit.
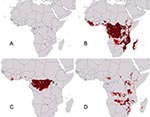
The geographic distribution of filovirus disease spreads generally across the humid Afrotropics (Figure 1A). Outlier occurrences lie at the eastern extreme of the distribution, consisting of occurrences associated with Ebola Sudan and Marburg viruses. Preliminary analyses of these geographic distributions, based on random subsets of the few data points available, indicated high statistical significance to model predictions: predictions of the geographic distribution of filovirus HFs correctly included random independent subsets much better than random model expectations (all p < 10-7). Although subsequent modeling was done without subsetting to maximize occurrence data, these preliminary results nonetheless indicated excellent predictivity of our distributional hypoptheses.
Modeling the distribution of Filoviridae in general (all points in Figure 1A) produced a broad potential distribution across the Afrotropics, including areas from which filovirus HF occurrences have not been reported (Tanzania, Mozambique, Madagascar; Figure 1B). Predicted distributions of the two major Filoviridae clades—Ebola and Marburg viruses—showed different geographic patterns. When just the three African Ebola virus species were analyzed together, areas of overprediction in eastern Africa disappeared, and predicted distributional areas included only areas surrounding known occurrence points, except for a few small disjunct areas in West Africa (Figure 1C). The predicted distribution did not include all of the Afrotropics—coastal central Africa and most of West Africa appeared not to be included, although these models are based on very small samples of occurrences.
When we analyzed the relatively few Marburg HF occurrences for which distributional data exist (n = 4 occurrences), a complementary distributional area was predicted (Figure 1D). Marburg HF was predicted to be absent in the humid Afrotropics, rather appearing focused in drier areas in eastern and south-central Africa. In contrast to Ebola HF, Marburg virus appears to have the potential to occur in areas from which filovirus disease has not yet been described.
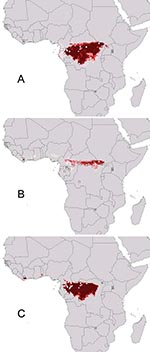
Sequential omission of Ebola virus species from analyses provided a view of ecologic similarity of species (45): if omission of a particular species causes little overall change, then its ecologic characteristics are not distinct from those of the remaining species. Omission of Ebola Ivory Coast had little effect on the prediction (Figure 2A; note predicted area in Ivory Coast); similarly, predictions omitting Ebola Zaire included at least part of the distribution of Ebola Zaire (southern portion omitted; Figure 2B). Eliminating Ebola Sudan, however, yielded a prediction completely excluding the distribution of Ebola Sudan (Figure 2C), which suggests that Ebola Sudan occurs under a distinct ecologic regime.
Inspection of niche models of Ebola HF occurrences (Marburg HF excluded) in ecologic space (Figure 3) provided insight into their ecologic distribution. Predicted Ebola HF occurrences were concentrated in regions presenting high precipitation combined with moderate-to-high temperatures (Figure 3A), coinciding with the ecologic distribution of evergreen broadleaf forest, although in specific cases that forest may be highly disturbed. In fact, >50% of African evergreen broadleaf forest is predicted to be within the niche of Ebola HF; no other land-cover type exceeded 5% within the Ebola HF niche (Figure 3B). In other dimensions, Ebola HF occurrences were distributed centrically in African environments and did not include extremes (Figure 3C–D).
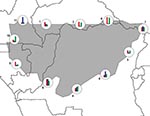
Distributional limits are complex results of multiple causal agents. A species is seldom limited on all sides by a single factor; rather, distributional limits are the combined result of many such factors. Inspection of the ecologic dimensions coincident with modeled geographic limits of Ebola HF occurrences (Figure 4) showed some of this complexity. At points around the distributional limit of Ebola HF distributional areas in central Africa, precipitation dominates the range limit at point 11, but temperature and elevation dominate at points 2, 3, and 6. Moreover, gradients are steeper in some areas than others (e.g., point 6 vs. 3). This preliminary analysis thus illustrates the complex relationships between ecologic dimensions and distributional limits.
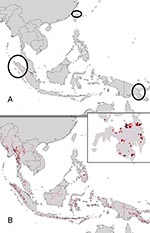
Given the mysterious origin of Ebola Reston virus (Ebola HF among macaques in a breeding facility on Luzon, Philippines) (9,13), a key question regarding Ebola HF distribution and ecology is whether similar ecologic conditions exist in Southeast Asia (e.g., Philippines). Projecting ecologic niche models for Marburg HF in Africa onto Asian environments identified few “appropriate” areas: only a few scattered areas in Papua New Guinea and Indonesia (Figure 5A). Projection of Ebola HF models, however, identified broader potential distributional areas in Southeast Asia (Figure 5B), including the lowlands of Mindanao (Figure 5, inset), a finding that suggests that similar ecologic conditions exist in the Philippines.
Ecology and Geography of Filovirus Occurrences
The ecologic niche characteristics reconstructed for filovirus species disease outbreaks coincided closely with phylogenetic patterns in the group (1,48). That is, disease sites for Ebola Ivory Coast and Ebola Zaire coincided ecologically, and these viruses are phylogenetically sister taxa. Ebola Sudan is genetically and ecologically most distinct among Ebola virus species, and (with Ebola Reston) forms the sister clade to Ebola Ivory Coast + Ebola Zaire. Correspondence between phylogenetic and ecologic patterns suggests that ecologic distributions of Ebola Sudan and Ebola Reston may prove similar; hence, the ecologic characteristics of Ebola Sudan may provide clues about the origin of Ebola Reston.
Marburg HF occurrence sites are quite distinct, with minimal overlap with Ebola HF ecologic distributions, coinciding with Marburg virus’ distant position in the phylogeny of the Filoviridae. This pattern suggests that Marburg virus and the Ebola viruses may have host species with markedly different ecologic requirements.
Ebola Reston
The geographic origin of Ebola Reston virus has been subject of controversy (9,49). Although the Ebola virus-infected monkeys initially documented in Reston, Virginia, originated in the Philippines, whether Ebola Reston occurs naturally in the Philippines has been debated. Nevertheless, the virus is distinct, and its geographic distribution is otherwise unknown. Given the phylogeny-ecology correspondence documented above, the ecology of Ebola Sudan may prove key in predicting the distribution of Ebola Reston, but the scanty occurrence data make species-specific models difficult. Our results are relevant in that ecologic conditions under which Ebola HF occurs in Africa are also found in the Philippines.
In previous analyses of animals, the conservative nature of ecologic niches has been documented to lead to prediction into regions inhabited by congener species (45). To the extent that host-parasite codistribution and cospeciation may be involved in the virus-reservoir relationships of filoviruses, prediction of potential distributional areas in the Philippines may reflect conservative niche evolution in the host taxon. Of course, because of historical effects (e.g., limited dispersal) on species’ distributions, potential distributional areas are often predicted in areas not inhabited (44), so this evidence is not definitive.
Limitations of the Approach
Limitations of our approach should be recognized. First, small sample sizes become critical. Although predictive models can be developed with relatively small samples of occurrence points (39), sample sizes for filovirus HF disease outbreaks are so minimal that single data points can change overall results. Examples of this sensitivity include the Zimbabwe Marburg HF disease outbreak and the Booue, Gabon, Ebola Zaire HF outbreak; inclusion of these points causes geographic predictions to be expanded considerably.
Other limitations center on the ecologic dimensions in which the niche is modeled. If additional dimensions exist that limit species’ distributions (and they certainly do) , GARP predictions will be overly large. Jackknife manipulations (systematic omission of ecologic dimensions to assess sensitivity to coverage density) can, to some degree, help in assessing sensitivity to coverage completeness (42), but dimensions more important than the set actually used may exist. Particularly relevant is climate variability—extreme events such as droughts and heavy rainfall may prove particularly relevant to filovirus transmission but are not included herein; such more complex models are under development (A.T. Peterson et al., unpub. data). Spurious associations between occurrence points and ecologic dimensions, though usually detected through independent test datasets, can limit distributional predictions overmuch.
Natural Reservoirs for Filoviruses
Detailed understanding of the geography and ecology of filovirus HF outbreaks represents an underexplored avenue of investigation regarding natural transmission cycles of filoviruses. We assembled available information regarding filovirus HF outbreaks and used various analytical tools to arrive at a detailed understanding of geography and ecology of filovirus disease occurrences. Consequently, we can now assemble criteria by which potential reservoir taxa might be judged. If one assumes a fair degree of host specificity in this host-parasite system, patterns of codistribution and cophylogeny can be expected. Hence, criteria include the following: 1) African Ebola virus reservoirs would be distributed principally in evergreen broadleaf forest; 2) the main focus of the geographic distribution of the reservoir(s) would be in the Congo Basin; 3) a disjunct (allopatric) distributional area would be present in West Africa; 4) a related taxon in eastern Africa would range in more arid habitats; 5) the reservoir would belong to a clade more broadly distributed across Africa and Southeast Asia.
Assessment of potential reservoir taxa by using these criteria has begun (A.T. Peterson et al., unpub. data), with the idea of eventually testing hypotheses of host associations through ecologic niche comparison methods (22). The goal, to be explored in future publications, is to develop reduced lists of taxa of highest priority for virus survey.
Dr. Peterson is associate professor in the Department of Ecology and Evolutionary Biology and Curator in the Natural History Museum and Biodiversity Research Center, University of Kansas. His research interests focus on the geography of species’ distributions
Acknowledgments
We thank David Vieglais, Ricardo Scachetti-Pereira, and David Stockwell for technical innovations; Rigoberto Esono Anvene for assistance with georeferencing localities; Emily Jentes for literature research; Tom Ksiazek, Pierre Rollin, Dan Bausch, and Darin Carroll for valuable comments on versions of the manuscript; and Ellen Salehi for the spark of interest in infectious diseases.
Much of the writing of this contribution was carried out at the Centro de Referência em Informação Ambiental, funded by the Fundação de Amparo à Pesquisa do Estado de São Paulo, Brazil, and the U.S. National Science Foundation.
References
- Murphy FA, Peters CJ. Ebola virus: where does it come from and where is it going? In: Krause RM, editor. Emerging infections: biomedical research reports. San Diego: Academic Press; 1998. p. 375–410.
- Ebola haemorrhagic fever in Zaire, 1976. Bull World Health Organ. 1978;56:271–93.PubMedGoogle Scholar
- Ebola haemorrhagic fever in Sudan, 1976. Bull World Health Organ. 1978;56:247–70.PubMedGoogle Scholar
- Baron RC, McCormick JB, Zubeir OA. Ebola virus disease in southern Sudan: hospital dissemination and intrafamilial spread. Bull World Health Organ. 1983;61:997–1003.PubMedGoogle Scholar
- Bertherat EJ, Renaut AA, Nabias RJ, Dubreuil G, Georges-Courbot M-C. Leptospirosis and Ebola virus infection in five gold-panning villages in northeastern Gabon. Am J Trop Med Hyg. 1999;60:610–5.PubMedGoogle Scholar
- Conrad JL, Isaacson M, Smith EB, Wulff H, Crees M, Geldenhuys P, Epidemiologic investigation of Marburg virus disease, southern Africa, 1975. Am J Trop Med Hyg. 1978;27:1210–5.PubMedGoogle Scholar
- Formentry P, Boesch C, Wyers M, Steiner C, Donati F, Dind F, Ebola virus outbreak among wild chimpanzees living in a rain forest of Cote d'Ivoire. J Infect Dis. 1999;179:S120–6. DOIPubMedGoogle Scholar
- Georges AJ, Leroy E, Renaut AA, Benissan CT, Nabias RJ, Trinh Ngoc M, Ebola hemorrhagic fever outbreaks in Gabon, 1994–1997: epidemiologic and health control issues. J Infect Dis. 1999;179:S65–75. DOIPubMedGoogle Scholar
- Hayes CG, Burans JP, Ksiazek TG, del Rosario RA, Miranda ME, Manaloto CR, Outbreak of fatal illness among captive macaques in the Philippines caused by an Ebola-related filovirus. Am J Trop Med Hyg. 1992;46:664–71.PubMedGoogle Scholar
- Heymann DL, Weisfeld JS, Webb PA, Johnson KM, Cairns T, Berquist H. Ebola hemorrhagic fever: Tandala, Zaire, 1977–1978. J Infect Dis. 1980;142:372–6.PubMedGoogle Scholar
- Khan AS, Kweteminga T, Heymann DL, Le Guenno B, Nabeth P, Kerstiens B, The reemergence of Ebola hemorrhagic fever, Democratic Republic of the Congo, 1995. J Infect Dis. 1999;179:S76–86. DOIPubMedGoogle Scholar
- Leroy E, Souquiere S, Rouquet P, Drevet D. Re-emergence of Ebola haemorrhagic fever in Gabon. Lancet. 2002;359:712. DOIGoogle Scholar
- Miranda ME, Ksiazek TG, Retuya TJ, Khan AS, Sanchez A, Fulhorst CF, Epidemiology of Ebola (subtype Reston) virus in the Philippines, 1996. J Infect Dis. 1999;179:S115–9. DOIPubMedGoogle Scholar
- Arata AA, Johnson B. Approaches toward studies on potential reservoirs of viral haemorrhagic fever in southern Sudan (1977). In: Pattyn SRS, editor. Ebola virus haemorrhagic fever. New York: Elsevier; 1978. p. 191–200.
- Germain M. Collection of mammals and arthropods during the epidemic of haemorrhagic fever in Zaire. In: Pattyn SRS, editor. Ebola virus haemorrhagic fever. New York: Elsevier; 1978. p. 185–9.
- Leirs H, Mills JN, Krebs JW, Childs JE, Akaibe D, Woollen N, Search for the Ebola virus reservoir in Kikwit, Democratic Republic of the Congo: Reflections on a vertebrate collection. J Infect Dis. 1999;179:S155–63. DOIPubMedGoogle Scholar
- Swanepoel R, Leman PA, Burt FJ, Zachariades NA, Braack LEO, Ksiazek TG, Experimental inoculation of plants and animals with Ebola virus. Emerg Infect Dis. 1996;2:321–5.PubMedGoogle Scholar
- Turell MJ, Bressler DS, Rossi CA. Lack of virus replication in arthropods after intrathoracic inoculation of Ebola Reston virus. Am J Trop Med Hyg. 1996;55:89–90.PubMedGoogle Scholar
- Smith DH, Johnson BK, Isaacson M, Swanepoel R, Johnson KM, Killey M, Marburg-virus disease in Kenya. Lancet. 1982;1:816–20. DOIPubMedGoogle Scholar
- Murphy FA, Kiley MP, Fisher-Hoch SP. Filoviridae: Marburg and Ebola viruses. In: Fields BN, Knipe DM, editors. Virology. New York: Raven Press, Ltd.; 1990. p. 933–42.
- Clayton DH, Moore J. Host-parasite co-evolution. General principles and avian models. Oxford: Oxford University Press; 1997.
- Peterson AT, Sanchez-Cordero V, Beard CB, Ramsey JM. Ecologic niche modeling and potential reservoirs for Chagas disease, Mexico. Emerg Infect Dis. 2002;8:662–7.PubMedGoogle Scholar
- Peterson AT, Stockwell DRB, Kluza DA. Distributional prediction based on ecological niche modeling of primary occurrence data. In: Scott JM, editor. Predicting species occurrences: issues of scale and accuracy. Washington: Island Press; 2002. p. 617–23.
- Le Guenno B, Formentry P, Wyers M, Guonon P, Walker F, Boesch C. Isolation and partial characterisation of a new strain of Ebola virus. Lancet. 1995;345:1271–4. DOIPubMedGoogle Scholar
- World Health Organization. Suspect cases of Ebola in Liberia. Press Release 11 December 1995. Available from: URL: http://www.who.int/archives/inf-pr-1995/r95-90.html
- Centers for Disease Control and Prevention. Outbreak of Ebola hemorrhagic fever—Uganda, August 2000–January 2001. MMWR Morb Mortal Wkly Rep. 2001;50:73–7.PubMedGoogle Scholar
- Bertherat EJ, Talarmin A, Zeller H. [Democratic Republic of the Congo: between civil war and the Marburg virus]. Med Trop. 1999;59:201–4.
- Zeller H. [Lessons from the Marburg virus epidemic in Durba, Democratic Republic of the Congo (1998–2000)]. Med Trop. 2000;60:23–6.
- World Health Organization. Marburg disease in Democratic Republic of Congo—update 2. Communicable Disease Surveillance and Response Disease Outbreak News. 6 May 1999.
- Stockwell DRB, Peters DP. The GARP modelling system: problems and solutions to automated spatial prediction. International Journal of Geographic Information Systems. 1999;13:143–58. DOIGoogle Scholar
- Stockwell DRB, Noble IR. Induction of sets of rules from animal distribution data: A robust and informative method of analysis. Math Comput Simul. 1992;33:385–90. DOIGoogle Scholar
- Stockwell DRB. Genetic algorithms II. In: Fielding AH, editor. Machine learning methods for ecological applications. Boston: Kluwer Academic Publishers; 1999. p. 123–44.
- MacArthur R. Geographical ecology. Princeton (NJ): Princeton University Press; 1972.
- Anderson RP, Lew D, Peterson AT. Using intermodel variation in error components to select best subsets of ecological niche models. Ecol Modell. 2003;162:211–32. DOIGoogle Scholar
- Anderson RP, Laverde M, Peterson AT. Using niche-based GIS modeling to test geographic predictions of competitive exclusion and competitive release in South American pocket mice. Oikos. 2002;93:3–16. DOIGoogle Scholar
- Anderson RP, Laverde M, Peterson AT. Geographical distributions of spiny pocket mice in South America: insights from predictive models. Glob Ecol Biogeogr. 2002;11:131–41. DOIGoogle Scholar
- Feria TP, Peterson AT. Using point occurrence data and inferential algorithms to predict local communities of birds. Divers Distrib. 2002;8:49–56. DOIGoogle Scholar
- Stockwell DRB, Peterson AT. Controlling bias in biodiversity data. In: Scott JM, editor. Predicting species occurrences: issues of scale and accuracy. Washington: Island Press; 2002. p. 537–46.
- Stockwell DRB, Peterson AT. Effects of sample size on accuracy of species distribution models. Ecol Modell. 2002;148:1–13. DOIGoogle Scholar
- Peterson AT. Predicting species’ geographic distributions based on ecological niche modeling. Condor. 2001;103:599–605. DOIGoogle Scholar
- Peterson AT, Ball LG, Cohoon KC. Predicting distributions of tropical birds. Ibis. 2002;144:e27–32. DOIGoogle Scholar
- Peterson AT, Cohoon KC. Sensitivity of distributional prediction algorithms to geographic data completeness. Ecol Modell. 1999;117:159–64. DOIGoogle Scholar
- Peterson AT, Papes M, Kluza DA. Predicting the potential invasive distributions of four alien plant species in North America. Weed Sci. In press.
- Peterson AT, Sanchez-Cordero V, Soberon J, Bartley J, Buddemeier RH, Navarro-Siguenza AG. Effects of global climate change on geographic distributions of Mexican Cracidae. Ecol Modell. 2001;144:21–30. DOIGoogle Scholar
- Peterson AT, Soberon J, Sanchez-Cordero V. Conservatism of ecological niches in evolutionary time. Science. 1999;285:1265–7. DOIPubMedGoogle Scholar
- Peterson AT, Vieglais DA. Predicting species invasions using ecological niche modeling. Bioscience. 2001;51:363–71. DOIGoogle Scholar
- Peterson AT, Scachetti-Pereira R, Hargrove WW. Potential distribution of Asian longhorned beetles (Anoplophora glabripennis) in North America. Am Midl Nat. 2004. In press.
- Georges-Courbot M-C, Sanchez A, Lu C-Y, Baize S, Leroy E, Lansout-Soukate J, Isolation and phylogenetic characterization of Ebola viruses causing different outbreaks in Gabon. Emerg Infect Dis. 1997;3:59–62.PubMedGoogle Scholar
- Peters CJ, Johnson ED, Jahrling PB, Ksiazek TG, Rollin PE, White J, In: Morse SS, editor. Emerging viruses. Oxford: Oxford University Press; 1993. p. 159–75.
Figures
Table
Cite This Article1Although filovirus taxonomy undergoes frequent revision, we follow nomenclature most recently established by the International Committee of Taxonomy of Viruses in 2002. Filoviruses consist of two genera. The genus Marburgvirus contains one species, Lake Victoria marburgvirus, with several recognized strains, and Ebolavirus contains four species: Ivory Coast ebolavirus, Reston ebolavirus, Sudan ebolavirus, and Zaire ebolavirus. For simplicity, we refer to the viruses using the unitalicized vernacular (e.g., Ebola Zaire). We use “Ebola viruses” to refer in general to members of Ebolavirus and “Marburg viruses” to refer in general to members of Marburgvirus. The diseases caused by filoviruses are termed Ebola hemorrhagic fever (HF; diseases caused by Ebola viruses) and Marburg HF (diseases caused by Marburg viruses).
Table of Contents – Volume 10, Number 1—January 2004
| EID Search Options |
|---|
|
|
|
|
|
|

Please use the form below to submit correspondence to the authors or contact them at the following address:
A. Townsend Peterson, Natural History Museum and Biodiversity Research Center, University of Kansas, Lawrence, Kansas 66045, USA; fax: 785-864-5335
Top