Volume 10, Number 12—December 2004
Research
Identifying Rodent Hantavirus Reservoirs, Brazil
Cite This Article
Citation for Media
Abstract
This study describes the genetic analysis carried out on samples from hantavirus pulmonary syndrome (HPS) patients from southern and southeastern states of Brazil and rodents captured at the presumed site of infection of these patients. A total of 65 samples that were antibody-positive for Sin Nombre or Laguna Negra virus by enzyme-linked immunosorbent assay were processed by nested reverse transcription–polymerase chain reaction (RT-PCR) by using several primer combinations in the M and S genome segments. PCR products were amplified and sequenced from samples from 11 HPS patient and 7 rodent samples. Phylogenetic analysis of nucleotide sequence differences showed the cocirculation of Araraquara and Juquitiba-like viruses, previously characterized from humans. Our genetic data indicate that Araraquara virus is associated with Bolomys lasiurus (hairy-tailed Bolo mouse) and the Juquitiba-like virus is associated with Oligoryzomys nigripes (black-footed pigmy rice rat).
Hantaviruses are mainly rodentborne viruses of the family Bunyaviridae (1). Two clinical forms of infections by hantaviruses are known: hemorrhagic fever with renal syndrome (HFRS) in the Old World and hantavirus pulmonary syndrome (HPS) in the American continent (2–4). Hantaviruses are enveloped, single-stranded, negative-sense RNA viruses, with a genome with three segments, designated small (S), medium (M), and large (L). The S segment encodes the nucleocapsid protein N, the M segment encodes a glycoprotein precursor that is processed into the envelope glycoproteins G1 and G2, and the L segment encodes the RNA polymerase (5,6).
The hantaviruses that cause HPS are associated with wild rodents species of the subfamily Sigmodontinae. They are transmitted mainly by contact or through aerosols of excrete and secretions of infected rodents (7–9). Person-to-person transmission has been reported in the 1996 outbreak in Argentina, involving the Andes (AND) virus (10,11). In Chile, this kind of transmission is suggested by clusters of cases in household contacts (12).
In Brazil, during the 1980s and 1990s, virologic and serologic studies conducted in humans and urban rodents showed the circulation of a hantavirus related to Seoul virus (13–16). In 1993, cases of an acute respiratory illness were detected in a family cluster in Juquitiba County, approximately 80 km from São Paulo City, in southeastern Brazil. Three brothers were affected by the infection, and two of them died (17,18). Necropsy material from one of them allowed the genetic characterization of a new hantavirus, later named Juquitiba (JUQ) virus, by sequencing a fragment of 139 nucleotides (nt) of the M genomic segment G2 encoding region (19). During 1995 and 1996, three more cases of HPS were confirmed by enzyme-linked immunosorbent assay (ELISA); one patient was from the central western county of Vilarejo de Castelo dos Sonhos, in Mato Grosso State, and the remaining two patients were from Araraquara and Franca counties in São Paulo State. Molecular studies carried out on samples from those HPS patients identified two novel genetic lineages of hantaviruses, Castelo dos Sonhos (CAS) and Araraquara (ARA) viruses (20). In 1998, new cases of HPS were detected: two in Minas Gerais, four cases in Rio Grande do Sul, and five in São Paulo State. Since then, an increasing number of HPS cases have been diagnosed annually in many states of Brazil. By March 2004, 342 HPS cases had been diagnosed on the basis of characteristic clinical syndrome, epidemiologic data, and Ig (immunoglobulin) M and IgG serologic response against Sin Nombre (SN), Laguna Negra (LN), or AND virus antigens by ELISA (9,21). Some of these cases were also diagnosed by immunohistochemistry. Most of the HPS cases occurred in the southern and southeastern states of Brazil (177 and 113, respectively). Paraná reported the highest number of cases (n = 92), followed by São Paulo (n = 59), Minas Gerais (n = 54), Santa Catarina (n = 50), Rio Grande do Sul (n = 35), Mato Grosso (n = 33), Maranhão (n = 7), Pará (n = 4), Goiás (n = 3), Rio Grande do Norte (n = 1), and Bahia (n = 1) (M. Elkhoury, pers. comm.).
This study describes the genetic analysis carried out on samples from HPS-case patients from southern and southeastern states of Brazil and rodents captured at the presumed site of infection of the human case-patients. The primary aims were to identify the hantavirus lineages causing HPS in that area, because few reports were available on this topic, and to identify the potential rodent host reservoirs because genetic data were not available from hantavirus-positive rodents. Genetic analysis of the nucleotide sequences indicates that ARA and JUQ-like viruses are circulating in the studied area. We report the genetic identification of the putative primary rodent reservoirs for these viruses.
Area of Study
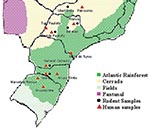
The studied areas included two kinds of natural ecosystems: the Atlantic rainforest and “cerrado” at the southern states of Paraná, Santa Catarina, and Rio Grande do Sul and at the southeastern states of Minas Gerais and São Paulo (Figure 1). Basically, the Atlantic rainforest extends along the Brazilian Atlantic Coast, and it is found as umbrofilous tropical forest of hillside or as its regional variation known as Araucaria forest. The cerrado occurs in the Brazilian central plateau, river basin, and part of northeastern region, and it is characterized by small trees, and grass vegetation, adapted to climates with long dry periods. Both kinds of ecosystems are found in São Paulo and Minas Gerais States.
Patient and Rodent Samples
We studied samples from HPS patients and from rodents captured at the potential sites where HPS exposures occurred. All samples included in the present study had tested positive to hantavirus by ELISA (9) by using SN virus and LN virus antigens (provided by T.G. Ksiazek, Centers for Disease Control and Prevention, Atlanta, GA).
Patients
A total of 40 blood and serum samples of HPS patients from five different states of Brazil were processed by nested RT-PCR: 6 samples from Minas Gerais (Patrocinio, Uberaba, Araxá, and Passos); 10 samples from São Paulo (Flórida Paulista, Batatais, Franca, São Carlos, Jaú, Cotia, Barra do Turvo, and Tupi Paulista); 7 samples from Paraná (General Carneiro, Bituruna, Ponta Grossa, Catanduvas, Curitiba, and Guarapuava); 7 samples from Santa Catarina (Seara, Arroio Trinta, and Lindóia do Sul); 10 samples from Rio Grande do Sul (Vacaria, Pelotas, Marcelino Ramos, São Lourenço do Sul, Capão Canoa, Santana do Livramento, Santa Cruz do Sul, Novo Hamburgo, and Arvorezinha). Of seven samples from HPS patients, five samples from Santa Catarina were from a family cluster reported by Seara (22).
Rodents
Rodents were captured by using Sherman live-capture traps (Sherman Traps Inc., Tallahassee, FL) set in rural or sylvan environments, around the presumed sites of HPS infection. The rodents were processed in the field; biologic samples (blood, liver, kidneys, spleen, heart, and lung) were obtained according to established biosafety guidelines (23) and stored in liquid nitrogen for further processing. The carcasses of the rodents were brought to the laboratory; the skins and craniums were used for further identification of the positive specimens. Samples of carcasses were deposited at Museu de Zoologia—Universidade de São Paulo, São Paulo, São Paulo State, and Museum of Southwestern Biology, University of New Mexico, Division of Mammals, Albuquerque, New Mexico, and most of the specimens were deposited in the Vertebrates Collection at Instituto Adolfo Lutz— Seção de Vírus Transmitidos por Artrópodos in São Paulo, SP.
A total of 25 rodent samples (subfamily Sigmodontinae) were studied by nested RT-PCR: 3 samples were from Uberlândia (2 Bolomys lasiurus) and Uberaba (1 B. lasiurus), in Minas Gerais; 13 samples were from Araraquara (1 B. lasiurus), Batatais (2 B. lasiurus), Franca (1 B. lasiurus and 1 Calomys tener), Cassia dos Coqueiros (1 Oximycterus rutilans), Cravinhos (1 B. lasiurus), Fartura (1 Akodon sp.), Mariápolis (2 B. lasiurus), Nuporanga (1 B. lasiurus and 1 Oligoryzomys nigripes), and São Carlos (1 B. lasiurus), in São Paulo; 3 samples from General Carneiro (2 O. nigripes and 1 Akodon sp.) in Paraná; 3 samples from Seara (2 O. nigripes and 1 Bolomys sp.) in Santa Catarina; 3 samples from Marcelino Ramos (1 O. nigripes and 1 Akodon sp.) and São Lourenço do Sul (1 Akodon sp.), in Rio Grande do Sul.
RNA Extraction and Nested-RT-PCR
Total RNA was extracted from human blood samples and rodent lung samples by using the RNaid (PLUS) Kit (BIO 101 Inc., La Jolla, CA) as described elsewhere (24). Briefly, approximately 100 mg of tissue was mixed with 300 μL of cell lysis solution containing guanidine thiocyanate extracted with phenol/chloroform and purified with RNA matrix beads. From some tissue samples, RNA was extracted with Trizol LS reagent (Invitrogen Co., Carlsbad, CA), following the manufacturer’s recommendations. When human serum specimens were used as source of viral RNA, the QIAmp Viral RNA Kit (Qiagen, Chastworth, CA) was used according to the manufacturer’s instructions. Amplification of virus RNA was performed by “RT-PCR-one step,” followed by a second PCR amplification as described previously (4). Numerous primer combinations in the M and S segments were used in nested RT-PCR reactions, including oligonucleotide sequences published (4,25,26) and unpublished that were designed to amplify conserved fragments of the S and M genome segments of South American hantaviruses.
Genetic and Phylogenetic Analysis
The DNA products of the nested PCR reactions were separated from an agarose gel, and bands of the correct predicted size were purified from gel slices with a GeneClean kit (BIO 101 Inc.) or GFXTM PCR DNA and Gel Band Purification Kit (Amersham Pharmacia Biotech, Inc., Piscataway, NJ). The nucleotide sequence of these products was determined on an ABI PRISM 377 Genetic Analyzer (Applied Biosystems, Foster City, CA.) using the dydeoxy cycle sequencing technique (4). Sequences were aligned with those of previously described hantaviruses by using BioEdit version 5.0.9 (North Carolina State University, Raleigh, NC) and the computer software package Clustal W 1.4 (27). Primer sequences were removed from sample sequences before being aligned. Phylogenetic analysis was carried out on the multiple nucleotide and amino acid sequence alignments by using maximum parsimony (PAUP* version 4.0b4a Macintosh computer software programs) (28) and the distance-based neighbor-joining method. Phylogenetic analysis by maximum parsimony was obtained by the heuristic search method. Pairwise genetic distances were computed by using the Kimura-2 parameter, as implemented in the computer program MEGA version 2.1 software (29). The bootstrap support for the results of the phylogenetic analysis was based on 500 replicates. GenBank accession numbers of the previously published sequences of the hantaviruses used in this study are listed in figure legends.
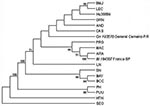
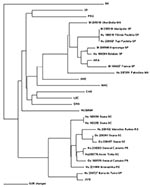
PCR products of the expected size were amplified from 11 of 40 HPS patient samples (Table 1) and 7 of 25 rodent samples studied (Table 2). A 303-nt fragment of the G2 gene was amplified and sequenced (bases corresponding to position 2807 to 3109 of Lechiguanas [LEC] virus). Phylogenetic analysis of nucleotide sequence differences showed the cocirculation of two genetic hantavirus lineages previously characterized from humans only: ARA virus and a genotype compatible with the previously identified JUQ virus. ARA virus sequences were derived from eight samples: from three HPS patients and three rodents (Bolomys lasiurus) from the state of São Paulo, and 1 HPS case-patient and one rodent (B. lasiurus) from Minas Gerais. Pairwise comparisons of the sequences of ARA virus strains from HPS patients and B. lasiurus showed an 85.1%-99.7% nt and 95%-99% amino acid (aa) identity. The viral sequence from the HPS patient Hu237251 from Patrocínio, Minas Gerais, the northernmost location included in this study, was most divergent from the other members of this group (16.7%). The second hantavirus genetic lineage identified was closely related to JUQ virus. Samples from seven HPS patients and three Oligoryzomys nigripes from the states of Rio Grande do Sul, Santa Catarina, Paraná, and São Paulo fall into this group (Figure 2).
Sequence comparison with JUQ virus, limited to an overlapping piece of 139 nt of the G2 encoding region, the only available from this virus (19), showed 85.7%-97.7% nt identity (Figure 3). The maximum identity with the prototype JUQ virus was found to correspond to the viral RNA from the patient Hu239727 (97.7%) from Barra do Turvo, São Paulo, the closest location to the site of infection of the fatal HPS case from which JUQ virus was originally characterized in Juquitiba in 1993. Sequences in JUQ-like clade group in two subclades, one including JUQ prototype strain and Hu239727, and the other one comprising six human and the three O. nigripes sequences (distance 0.273). The distance observed between these two subclades is intermediate between in-group and between group distances (Table 3).
Comparison of the 303-nt G2-encoding region sequences derived from HPS-patient and O. nigripes samples of this group showed 86.6%-99.7% nt identity. Sequence comparison with the corresponding to ARA group showed 20.8% nt and 4.9% amino acid (aa) divergence. Genetic distances between the sequences studied are shown in Table 3. The general time reversible model (GTR) with a 0.2479 proportion of invariable sites and γ = 0.4121, was used in analysis. AND virus resulted in ARA closest sequence (mean distance = 0.516) and Hu39494, JUQ-like closest sequence (0.723). Mean distances between ARA and JUQ-like sequences further support that they are different viruses.
A longer fragment of 1,239 nt of the G2 encoding region of the M segment (Figure 4), as well as a fragment of 259 nt of the nucleoprotein encoding region of the S segment, was generated by one representative strain of each hantavirus group (Bl194307 for ARA virus and On193576 for JUQ-like virus). Comparison of viral RNA from B. lasiurus sequences with ARA virus showed a 96.8%-nt and 99.3%-aa identity for the 1,239-nt G2-M piece, and 98.8% nt and 100% aa identity for the S segment piece. When RNA viral sequences from B. lasiurus were compared with those from O. nigripes, 77.8% nt and 93.4% aa identity for the 1,239-nt G2-M piece, and 84.1% nt and 98.8% aa identity for the S segment piece were observed.
The phylogenetic relationship of ARA and JUQ-like genotypes to other hantaviruses of South America was determined on the nucleotide sequences of the 303-nt and 1,239-nt G2-encoding region of the M segment genomes by using maximum parsimony and neighbor-joining analysis. Both trees showed a similar topology, except for the altered placement of certain genotypes that displayed low bootstrap support. Phylogenetic analysis performed on the 303-nt sequence fragment (Figure 2) showed that all samples from B. lasiurus fell into the ARA virus group, whereas all samples from O. nigripes fell into the JUQ-like genetic group. The nodes separating these groups had high bootstraps values (72% and 91%, respectively); however, the exact branching order among the Brazilian and the other hantaviruses cannot be resolved by the present phylogenetic analysis. The phylogenetic tree, based on the analysis of the 1,239-nt M segment sequence differences, showed the same topology of the 303-nt sequence, data-based tree. Both maximum parsimony (Figure 2) and neighbor-joining (data not shown) analysis demonstrated that the divergent ARA and JUQ-like viruses form a unique monophyletic clade with the other South American hantaviruses (71% bootstrap support).
Within this clade the ARA virus forms a subclade with other two akodontine-borne Argentinean hantaviruses, Maciel (MAC) from Necromys benefactus and Pergamino (PRG) from Akodon azarae, although with a low bootstrap support (66%). The JUQ-like virus from O. nigripes 193576 is ambiguously placed within the South American viruses, together with CAS virus from Brazil and the rest of the Oligoryzomys-borne Argentinean hantaviruses (LEC, Orán, Hu39694, Bermejo, AND virus), as well as LN virus from Paraguay. A slight difference in the topology of this 1,239-nt database tree was observed when neighbor-joining analysis was used; the grouping of this Oligoryzomys-borne Argentinean hantavirus clustered in a subclade with a bootstrap support of 79% (data not shown).
Eleven (27.5%) of 40 human samples and 7 (28.0%) of 25 rodent samples studied tested positive for hantavirus by RT-PCR. Previous studies have characterized three different hantavirus genetic lineages associated with HPS cases in Brazil: CAS, ARA, and JUQ (20,30,31).
Previous serologic studies detected IgG antibodies in rodent biologic samples: in B. lasiurus and Akodon spp. captured in the state of São Paulo (32), as well as B. lasiurus from São Paulo and Minas Gerais States, and O. nigripes from São Paulo, Rio Grande do Sul, Santa Catarina, and Paraná States (33).
The data we report on the phylogenetic analysis of viral M and S genome segment fragments, from HPS patients and rodent samples from different locations of southern and southeastern Brazil, showed the circulation of two distinct hantaviruses, closely related to ARA and JUQ viruses, previously characterized only from humans. Our data on the phylogenetic analysis of a 303-nt G2 encoding region of the M genome segment represents the first genetic evidence of the role of B. lasiurus as rodent host reservoir for ARA virus, as well as O. nigripes as rodent host reservoir for the JUQ-like virus in the region under study. The nodes separating each group of virus within the South American hantaviruses clade were highly supported (72% and 91% bootstrap for ARA and JUQ-like lineages, respectively). The two subclades observed in Figure 3 (i.e., 139-nt tree), and the intermediate distance between them suggested the possibility of some geographic isolation. More extensive sequencing of JUQ prototype and JUQ-like sequences is needed to clarify this point.
Comparison of the sequences of ARA virus strains obtained from B. lasiurus–infected patient samples and HPS patient samples showed an identity of up to 99.7% at the nucleotide level, while within the JUQ-like virus group, the comparison between O. nigripes and HPS patient virus strains showed an identity of up to 97.7%. However, the phylogenetic relationship of JUQ-like hantavirus to other members of the South American hantavirus lineages, determined on the nucleotide sequences of the 303-nt sequence, as well as on a longer fragment of a 1,239-nt piece of the M segment performed for one representative strain derived from one O. nigripes, systematically failed to resolve the branching order.
In South America, human illnesses associated with hantaviruses have been linked to viruses from the Oryzomyini and Phyllotini tribes. Known akodontine-borne hantaviruses have not been associated with human illnesses. Thus, the data reported here on ARA rodent reservoir constitute the first evidence that a hantavirus associated with an akodontine rodent can cause HPS. Phylogenetic tree based on a 1,239-nt G2 sequence fragment places together the akodontine-borne ARA virus from B. lasiurus in Brazil, MAC from N. benefactus, and PRG from A. azarae in Argentina (34). Although the bootstrap support displayed was low (66%), this finding is in accordance with previous observations based on the S genome phylogeny of Argentinean hantaviruses (35). This would support the hypothesis of cospeciation of hantaviruses with their specific rodent hosts. As it has been described with other hantaviruses, biogeographic factors are also involved in the evolution of hantavirus lineages (36,37). The human- and rodent-derived ARA strains analyzed in the current study were distributed at a distance of approximately 650 km. As expected, ARA virus strains originated from the more distant localities displayed the highest genetic divergence, as shown between samples from Patrocínio (Hu237251) and Uberlândia (Bl235018) in Minas Gerais State, and Flórida Paulista (Hu196618) in São Paulo (16.7%-nt difference). Similarly, the divergent JUQ-like virus sample Hu239727 originated from Barra do Turvo, São Paulo, in relation to the rest of the human and rodent JUQ-like virus samples included in this group from locations in Santa Catarina, Paraná, and Rio Grande do Sul may be associated with the geographic distance between them (500 km on average).
Although the data from serologic testing by ELISA indicated four positive Akodon spp., one C. tener, and one Ox. rutilans, specific viral sequences could not be amplified from those specimens. Thus, additional studies are needed to determine the possible role of these species in the epidemiology of hantavirus in Brazil.
The habitats and behavior of the rodents are important aspects to consider in elucidating the reservoirs of etiologic agents. ARA virus was recovered mostly from HPS patients as well as B. lasiurus samples from the ecosystem called cerrado, while JUQ-like virus was recovered mostly from human and O. nigripes samples from the ecosystem called Atlantic rainforest. Geographic distribution of B. lasiurus in Brazil includes the original areas of cerrado and “caatinga.” This environment is typical in northeastern Brazil, characterized by deciduous trees and cactus and an extremely prolonged dry season. The B. lasiurus distribution shows its ability to adapt to anthropic environments, especially grasses (Brachiaria) and sugar cane cultures. These rodents are aggressive and usually dominate the areas they infest (38); they do not colonize human dwellings, although occasionally they can invade houses.
O. nigripes is adapted to live in the primary and secondary forests, especially in the Atlantic rainforest and Araucaria forest. It is primarily found in anthropic environment, such as the lineal natural habitats bordering cultivated areas, especially those with corn, where it is the most abundant species. These rodents can easily invade dwelling houses and barns to search for food, and they can nest in the domestic habitats.
Among the rodents captured in the cerrado, B. lasiurus was the most abundant species (44%) and showed the highest prevalence of antibodies to hantavirus (11%). Akodon spp. and O. nigripes were the two most abundant among those rodents captured in the transition area between cerrado and the Atlantic rainforest, but the highest prevalence of antibodies to hantavirus was found in O. nigripes specimens (8%) (33). These data help incriminate B. lasiurus and O. nigripes in the transmission of ARA and JUQ-like viruses, respectively, to humans.
Dr. Suzuki is chief of the Seção de Vírus Transmitidos por Artrópodos at Instituto Adolfo Lutz, São Paulo State Health Secretary. Her research interests include the epidemiology and ecology of arboviruses, hantaviruses, and arenaviruses, and the laboratory diagnosis of infections by these agents.
Acknowledgments
We thank the staff of Seção de Vírus Transmitidos por Artrópodos, São Paulo State Health Secretary and A. Degleue, S. Bourda, and V. Fasciani for their laboratory help. We also thank the personnel of Brazilian Ministry of Health, State and Municipal Health Secretaries in the States of São Paulo, Minas Gerais, Paraná, Santa Catarina, and Rio Grande do Sul for their efforts in the surveillance of notifiable diseases.
This work was supported in part by VIGISUS/FUNASA/MS–Project no. 914/BRA/02. The work was done at Instituto Adolfo Lutz–Secretaria de Estado da Saúde de São Paulo, São Paulo/SP, Brazil, and Instituto Nacional de Enfermedades Virales Humanas Dr. Julio I. Maiztegui, Pergamino, Argentina.
References
- Schmaljohn CS, Hasty SE, Dalrymple JM, LeDuc JW, Lee HW, von Bonsdorff CH, Antigenic and genetic properties of viruses linked to haemorrhagic fever with renal syndrome into a newly defined genus of Bunyaviridae. Science. 1985;227:1041–4. DOIPubMedGoogle Scholar
- Schmaljohn C, Hjelle B. Hantavirus: a global disease problem. Emerg Infect Dis. 1997;3:95–104. DOIPubMedGoogle Scholar
- Peters CJ. Hantavirus pulmonary syndrome in the Americas. In: Scheld WM, Craig WA, Hughes JM, editors. Emerging infections 2. Washington: ASM Press, 1998. p. 17–64.
- Nichol ST, Spiropoulou CF, Morzunov S, Rollin PE, Ksiazek TG, Feldmann H, Genetic identification of a hantavirus associated with an outbreak of acute respiratory illness. Science. 1993;262:914–7. DOIPubMedGoogle Scholar
- Plyusnin A, Vapalahti O, Vaheri A. Hantaviruses: genome structure, expression and evolution. J Gen Virol. 1996;77:2677–87. DOIPubMedGoogle Scholar
- Elliott RM. Molecular biology of the Bunyaviridae. J Gen Virol. 1990;71:501–2. DOIPubMedGoogle Scholar
- Childs JE, Ksiazek TG, Spiropoulou CF, Krebs JW, Morzunov S, Maupin GO, Serologic and genetic identification of Peromyscus maniculatus as the primary rodent reservoir for a new hantavirus in the southwestern United States. J Infect Dis. 1994;169:1271–80. DOIPubMedGoogle Scholar
- LeDuc J. Epidemiology of Hantaan and related viruses. Lab Anim Sci. 1987;37:413–8.PubMedGoogle Scholar
- Ksiazek TG, Peters CJ, Rollin PE, Zaki S, Nichol S, Spiropoulou C, Identification of a new North American hantavirus that causes acute pulmonary insufficiency. Am J Trop Med Hyg. 1995;52:117–23.PubMedGoogle Scholar
- Wells RM, Sosa Estani S, Yadon ZE, Enria D, Padula P, Pini N, An unusual Hantavirus outbreak in Southern Argentina: person-to-person transmission? Emerg Infect Dis. 1997;3:1–5. DOIPubMedGoogle Scholar
- Padula PJ, Edelstein A, Miguel SD, Lopez NM, Rossi CM, Rabinovich RD. Hantavirus pulmonary syndrome outbreak in Argentina: molecular evidence for person-to-person transmission of Andes virus. Virology. 1998;241:323–30. DOIPubMedGoogle Scholar
- Galeno H, Mora J, Villagra E, Fernandez J, Hernandez J, Mertiz GJ, First human isolate of hantavirus (Andes virus) in the Americas. Emerg Infect Dis. 2002;8:657–61.PubMedGoogle Scholar
- Le Duc JW, Smith GA, Pinheiro FP, Vasconcelos PFC, Travassos da Rosa ES, Maiztegui JI. Isolation of Hantaan-related virus from Brazilian rats and serologic evidence of its widespread distribution in South America. Am J Trop Med Hyg. 1985;34:810–5.PubMedGoogle Scholar
- Hindrichsen S, Medeiros de Andrade A, Clement J, Leirs H, McKenna P, Matthy P, Hantavirus infection in Brazilian patients from Recife with suspected leptospirosis. Lancet. 1993;341:50. DOIPubMedGoogle Scholar
- Iversson LB, Travassos da Rosa APA, Rosa MDB, Lomar AV, Saski M, LeDuc JW. Infecção humana por hantavírus nas regiões Sul e Sudeste do Brasil. Rev Assoc Med Bras. 1994;40:85–92.PubMedGoogle Scholar
- Vasconcelos PFC, Travassos da Rosa ES, Travassos da Rosa APA, Travassos da Rosa JFS. Evidence of circulating hantaviruses in Brazilian Amazonia through high prevalence of antibodies in residents of Manaus, Brazil. Cienc Cult. 1992;44:162–3.
- Da Silva MV, Vasconcelos MJ, Hidalgo NTR, Veiga APR, Canzian M, Marotto PC, Hantavirus pulmonary syndrome. Report of the first three cases in São Paulo, Brazil. Rev Inst Med Trop Sao Paulo. 1997;39:231–4.PubMedGoogle Scholar
- Vasconcelos MJ, Lima V, Iversson L, Rosa M, Travassos da Rosa E, Travassos da Rosa A, Hantavirus pulmonary syndrome in the rural area of Juquitiba, São Paulo, Metropolitan Area, Brasil. Rev Inst Med Trop Sao Paulo. 1997;39:237–8.PubMedGoogle Scholar
- Monroe MC, Morzunov SP, Johnson AM, Bowen MD, Artsob H, Yates T, Genetic diversity and distribution of Peromyscus-borne hantaviruses in North America and comparison with other hantaviruses. Emerg Infect Dis. 1999;5:75–86. DOIPubMedGoogle Scholar
- Johnson AM, Souza LTM, Ferreira IB, Pereira LE, Ksiazek TG, Rollin PE, Genetic investigation of novel hantaviruses causing fatal HPS in Brazil. J Med Virol. 1999;59:527–35. DOIPubMedGoogle Scholar
- Feldmann H, Sanchez A, Morzunov S, Spiropoulou CF, Rollin PE, Ksiazek TG, Utilization of autopsy tissue RNA for the synthesis of nucleocapsid antigen of a newly recognized virus associated with Hantavirus pulmonary syndrome. Virus Res. 1993;30:351–67. DOIPubMedGoogle Scholar
- Tavares LMS, Elkhouri MR, Lanzieri TN, Dias RF, Hilleshein N, Caldas AC, Investigation of a cluster of hantavirus infection among family members in a household in Southern Brazil. Abstracts of the Fifth International Conference on Hemorrhagic Fever with Renal Syndrome (HFRS), Hantavirus Pulmonary Syndrome (HPS), and Hantaviruses, Annecy, France, 2001. Lyon, France: Fondation Mérieux; 2001. p. 189.
- Mills JN, Yates TL, Childs JE, Parmenter RR, Ksiazek TG, Rollin PE, Guidelines for working with rodents potentially infected with hantavirus. J Mammal. 1995;76:716–22. DOIGoogle Scholar
- Spiropoulou CF, Morzunov S, Feldmann H, Sanchez A, Peters CJ, Nichol ST. Genome structure and variability of a virus causing hantavirus pulmonary syndrome. Virology. 1994;200:715–23. DOIPubMedGoogle Scholar
- Levis SC, Morzunov SP, Rowe JE, Enría DA, Pini N, Calderón G, Genetic diversity and epidemiology of hantaviruses in Argentina. J Infect Dis. 1998;177:529–38. DOIPubMedGoogle Scholar
- Johnson AM, Bowen MD, Ksiazek TG, Williams RJ, Bryan RT, Mills JN, Laguna Negra Virus associated with HPS in Western Paraguay and Bolivia. Virology. 1997;238:115–27. DOIPubMedGoogle Scholar
- Thompson JD, Higgins DG, Gibson TJ. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choices. Nucleic Acids Res. 1994;22:4673–80. DOIPubMedGoogle Scholar
- Swofford DL. PAUP*: Phylogenetic Analysis Using Parsimony (and other methods), v. 4.0b4a. Sunderland (MA): Sinauer Associates, Inc.; 2000.
- Kumar S, Tamura K, Jakobsen IB, Nei M. MEGA2: Molecular evolutionary genetics analysis software. Tempe (AZ): Arizona State University; 2001.
- Morelli ML, Souza RLM, Badra SJ, Pinto AA, Figueiredo LTM. Phylogenetic study of the M segment of hantavirus from patients having cardiopulmonary syndrome. In: XIII National Meeting of Virology. Virus Rev Res. 2002;7:68.
- Souza RLM, Morelli ML, Badra SJ, Pinto AA, Figueiredo LTM. Phylogenetic study of the S segment of hantavirus from patients having cardiopulmonary syndrome. In: XIII National Meeting of Virology. Virus Rev Res. 2002;7:91–2.
- Katz G, Williams RJ, Burt MS, Souza LTM, Pereira LE, Mills JN, Hantavirus pulmonary syndrome in the State of São Paulo, Brazil, 1993–1998. Vector Borne Zoonotic Dis. 2001;1:181–9. DOIPubMedGoogle Scholar
- Souza LTM, Suzuki A, Pereira LE, Ferreira IB, Souza RP, Cruz AS, Identification of hantavirus rodent reservoirs species in south and southeastern Brazil. Short communication. Informe Epidemiológico do SUS/Centro Nacional de Epidemiologia, coord. Brasília: Ministério da Saúde. Fundação Nacional de Saúde. 1999;11:249–51.
- Levis S, Rowe J, Morzunov S, Enria DA, St Jeor S. New hantaviruses causing hantavirus pulmonary syndrome in central Argentina. Lancet. 1997;349:998–9. DOIPubMedGoogle Scholar
- Bohlman MC, Morzunov SP, Meissner J, Taylor MB, Ishibashi K, Rowe J, Analysis of hantavirus genetic diversity in Argentina: S segment-derived phylogeny. J Virol. 2002;76:3765–73. DOIPubMedGoogle Scholar
- Plyusnin A, Morzunov SP. Virus evolution and genetic diversity of hantaviruses and their rodent hosts. In: Schmaljohn CS, Nichol ST, editors. Hantaviruses (Curr Top Microbiol Immunol 256). Berlin: Springer-Verlag; 2001. p. 47–75.
- Morzunov SP, Rowe JE, Monroe MC, Ksiazek TG, Peters CJ, St. Jeor SC, Genetic analysis of the diversity and origin of hantaviruses in Peromyscus leucopus mice in North America. J Virol. 1998;72:57–64.PubMedGoogle Scholar
- Streilen KE. Ecology of small mammals in the Semiarid Brazilian Caatinga I. Climate and faunal composition. Andrew Carnegie Museum. 1982;51:79–106.
Figures
Tables
Cite This ArticleTable of Contents – Volume 10, Number 12—December 2004
| EID Search Options |
|---|
|
|
|
|
|
|


Please use the form below to submit correspondence to the authors or contact them at the following address:
Akemi Suzuki, Seção de Vírus Transmitidos por Artrópodos - Instituto Adolfo Lutz, Avenida Dr. Arnaldo, 355 – CEP 01246-902 – São Paulo/SP, Brazil; fax: 5511-3088-3753
Top